Addgene Resources to Grow Your Developmental Biology Toolkit
Posted by Mary Gearing (Addgene), on 8 November 2017
Addgene is a global, nonprofit repository that was created to help scientists share plasmids. Before we go over the developmental biology resources available at Addgene, here’s a little background on our organization. Our mission is to accelerate research and discovery by improving access to useful research materials and information. Labs deposit plasmids with Addgene at no cost, and we handle storage, distribution, and record keeping. Researchers can request plasmids and, in some cases, ready-made lentivirus and AAV, from our collection, and we coordinate Material Transfer Agreements to facilitate easy sharing.
We also provide numerous educational resources on our blog and website. Our Plasmids 101 and CRISPR 101 blog series were designed to help scientists of all levels learn more about molecular biology, cloning, genome engineering, and other popular technologies. To make it easy to find plasmids for a given purpose, we provide curated pages on our website for various plasmid collections like CRISPR, stem cells, cancer, and fluorescent proteins. Addgene’s technical support team also provides scientists with real-time plasmid troubleshooting. By providing these services, Addgene’s goal is to create a lasting resource for research and discovery around the world.
Developmental biologists have deposited many of Addgene’s most popular plasmids. I’ll cover the general collections most relevant to developmental biology and also highlight a few standouts from our collection.
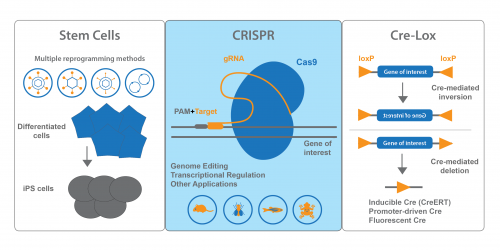
Stem Cell Tools
Our stem cell collection is a true Addgene strength. We have many plasmids that cover methods for iPS cell generation – retroviral, lentiviral, adenoviral, episomal and non-viral methods. We also distribute plasmids for direct differentiation and transdifferentiation. These plasmids are conveniently organized on our Stem Cell Collection page. Keep an eye out for the blue flame plasmids – these each have been requested over 100 times and have proven useful for many researchers!
 Plasmid spotlight: Andras Nagy Piggybac reprogramming plasmids
Plasmid spotlight: Andras Nagy Piggybac reprogramming plasmids
Lentiviral and retroviral reprogramming methods carry the caveat of insertional mutagenesis, while plasmid and adenoviral reprogramming methods suffer from lower efficiency. Woltjen et al. created a Tet-inducible reprogramming system using the Piggybac transposase. This system allows you to easily reprogram fibroblasts into iPS cells using only plasmids – there’s no need to prepare or deliver virus to your cells. Instead, you co-express PB-TET-MKOS containing the four Yamanaka factors, PB-CA-rtTA Adv containing the rtTA Tet transactivator, and a plasmid containing the Piggybac transposase (available from Transposagen and others.) Doxycycline treatment then turns on MKOS expression, promoting reprogramming. Once reprogrammed lines have been isolated, transient expression of Piggybac transposase will remove the MKOS transgene from the genome.
CRISPR Plasmids for Use With Developmental Models
Addgene’s CRISPR collection also has resources for scientists interested in developmental biology. On our CRISPR landing page, you can find plasmids sorted by model organism or system. In addition to our many mammalian expression plasmids, we have plasmids available for common developmental models like Drosophila, C. elegans, Xenopus and zebrafish. Addgene also maintains a list of Pre-designed gRNAs that have been used in previous publications, and you can easily search the table for your favorite gene. We also encourage you to deposit your own gRNA plasmids.
 Plasmid spotlight: Leonard Zon Zebrafish CRISPR plasmids
Plasmid spotlight: Leonard Zon Zebrafish CRISPR plasmids
Zebrafish have proven very amenable to CRISPR genome engineering, and the Zon lab Gateway constructs can help you to knock out your gene of interest in a tissue-specific manner. Their Tol2 based system enables you to examine mosaic disruption of your gene in F0 embryos or to generate stable tissue-specific knockout lines. All four plasmids from this publication have earned blue flames for 100+ requests! To use this system, you’ll combine Gateway destination vector pDestTol2pA2-U6:gRNA or pDestTol2CG2-U6:gRNA with a middle entry vector containing Cas9 (e.g. pME-Cas9). You’ll also need a 5’ entry vector containing your tissue-specific promoter and a 3’ entry vector with a polyA sequence, both of which can be obtained from the Tol2kit. A Gateway reaction will bring the pieces together, allowing you to examine the effects of gene knockout in a specific tissue. A detailed protocol is available from the Zon lab on the plasmid pages of each Addgene plasmid.
Cre-Lox Systems for Spatio-Temporal Control of Gene Expression
If you’re using Cre-lox recombination to turn gene expression ON or OFF, chances are Addgene has plasmids you can use. In addition to standard Cre recombinase, we also provide inducible, promoter-regulated, and optimized Cres, to name a few. Cre-dependent vectors expressing fluorescent proteins or luciferase are also available. FLEx (FLip-Excision) vectors, which conditionally turn off one gene and activate another, are available from many Addgene depositors. PhiC31 recombinase and other non-Cre recombinases are available for use in Drosophila.
 Plasmid spotlight: Connie Cepko pCAG-ERT2CreERT2 plasmid
Plasmid spotlight: Connie Cepko pCAG-ERT2CreERT2 plasmid
This plasmid expresses CreERT, or tamoxifen-inducible Cre, in a mammalian expression system. In cell lines transfected with this plasmid, tamoxifen will activate Cre, allowing removal or inversion of a floxed gene of interest. This plasmid has been requested over 1100 times, showing its versatility and applicability to many kinds of studies.
Lineage tracing plasmids make up another exciting mini-collection at Addgene. Brainbow is a combinatorial Cre-lox based system that colors single cells with as many as 90-160 different colors. Many of Addgene’s Brainbow plasmids have blue flames. Although originally developed for neuronal mapping, these plasmids have been adapted for lineage tracing. Multibow plasmids adapted from the Brainbow system for use in zebrafish create unique “barcodes” allowing researchers to track individual cells and their clonal progeny.
It’s impossible to cover all of the resources available at Addgene in one blog post, so we encourage you to take a look at our website. If you’re looking for plasmids from a specific publication, start by searching for the paper’s corresponding author to see if she or he has deposited with us. You can also sign up for Addgene Alerts for your favorite PIs and you’ll get emails whenever new plasmids are available from their labs. If you have trouble finding what you need, or you’d like us to reach out to a potential depositor, feel free to email us at help@addgene.org. We also encourage you to deposit your own plasmids thereby making them easily accessible to other members of the research community.
Mary Gearing is a Scientist at Addgene. She enjoys developing educational content to help scientists learn more about molecular biology. Follow her on Twitter @megearing.
Addgene Resources
Addgene Blog
Cre-ating New Methods for Site-Specific Recombination in Drosophila
References
piggyBac transposition reprograms fibroblasts to induced pluripotent stem cells. Woltjen K, Michael IP, Mohseni P, Desai R, Mileikovsky M, Hamalainen R, Cowling R, Wang W, Liu P, Gertsenstein M, Kaji K, Sung HK, Nagy A. Nature. 2009 Apr 9;458(7239):766-70. doi: 10.1038/nature07863. PMID: 19252478
A CRISPR/Cas9 Vector System for Tissue-Specific Gene Disruption in Zebrafish. Ablain J, Durand EM, Yang S, Zhou Y, Zon LI. Dev Cell. 2015 Mar 4. pii: S1534-5807(15)00075-1. doi: 10.1016/j.devcel.2015.01.032. PMID: 25752963
High-Efficiency FLP and PhiC31 Site-Specific Recombination in Mammalian Cells. Raymond CS, Soriano P. PLoS ONE. 2007 Jan 17;2(1):e162. PMID: 17225864
Multiple new site-specific recombinases for use in manipulating animal genomes. Nern A, Pfeiffer BD, Svoboda K, Rubin GM. Proc Natl Acad Sci USA. 2011 Aug 23;108(34):14198-203. doi: 10.1073/pnas.1111704108. PMID: 21831835
Controlled expression of transgenes introduced by in vivo electroporation. Matsuda T, Cepko CL. Proc Natl Acad Sci U S A. 2007 Jan 16;104(3):1027-32. PMID: 17209010
Improved tools for the Brainbow toolbox. Cai D, Cohen KB, Luo T, Lichtman JW, Sanes JR. Nat Methods. 2013 May 5;10(6):540-7. doi: 10.1038/nmeth.2450. PMID: 23817127
Multibow: digital spectral barcodes for cell tracing. Xiong F, Obholzer ND, Noche RR, Megason SG. PLoS One. 2015 May 26;10(5):e0127822. doi: 10.1371/journal.pone.0127822. eCollection 2015. PMID: 26010570


 (1 votes)
(1 votes)