Postdoc: RNAi biology in intragenomic conflict and speciation (Lai lab, NYC)
Posted by Eric Lai, on 7 April 2018
Closing Date: 15 March 2021
Eric Lai Lab, Sloan-Kettering, NYC
We have an opening for a postdoctoral position that will address fundamental questions in small RNA biology, genomic conflict, and speciation.
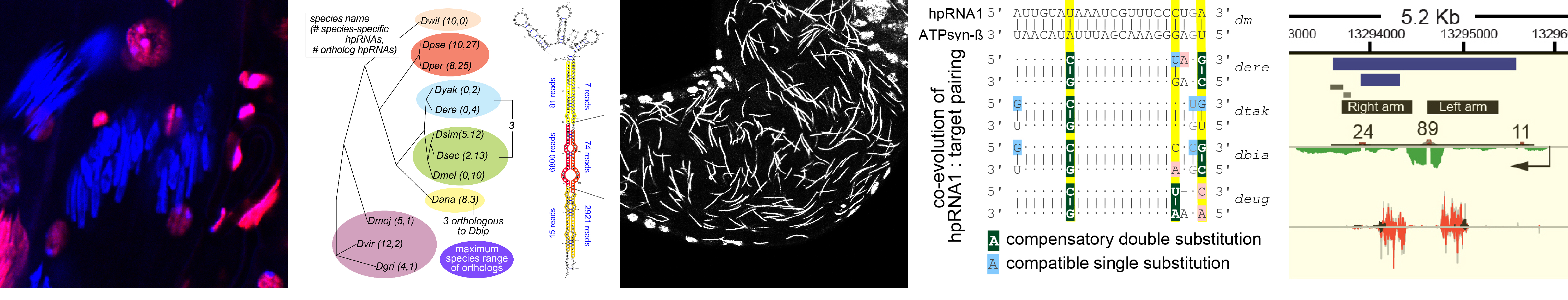
What is endogenous RNAi utilized for? We earlier described a mysterious class of endo-RNAi substrates termed hpRNAs (Okamura Nature 2008), and recently recognized that these mediate adaptive gene regulation in testis (Wen Molecular Cell 2015). These data open a window on a vital biology of RNAi, and now lead us to explore the evolution and function of RNAi systems across the Drosophilid phylogeny.
We discovered networks of rapidly evolving RNAi substrates we hypothesize resolve intragenomic conflicts, and successfully started to use CRISPR/Cas9 in non-model fly species to test some of these ideas (Lin 2018, in submission). Amazingly, while endogenous RNAi was not historically thought to have much phenotypic impact, we find that rapidly evolving genomic battles are being waged and critically depend upon the powerful weapon of RNAi silencing to propagate the species. Reciprocally, we are fascinated to understand by what novel molecular mechanisms de novo selfish meiotic factors can paradoxically drive population extinction.
We seek a motivated postdoctoral fellow with strong Drosophila molecular genetics experience and interest in integrating genome engineering, biochemistry, and bioinformatics to analyze the evolution and function of RNAi silencing systems in resolving deleterious intragenomic conflicts in testis. Although our entry point is RNAi biology, we anticipate that selfish factors yet to be discovered may mediate their effects through chromatin mechanisms. The successful candidate will integrate into a team that is actively engaged in diverse topics in gene regulation in Drosophila and mammalian models, and the Sloan-Kettering Institute provides a vibrant research community.
Funded position with housing and medical benefits are available immediately. Please provide CV, motivation letter and references to Eric Lai, laie@mskcc.org.
Relevant recent papers on hpRNAs and testis post-transcriptional regulation.
see also https://www.mskcc.org/research-areas/labs/eric-lai
Lin, C.-J., F. Hu, R. Dubruille, J. Wen, J. Vedanayagam, P. Smibert, B. Loppin and E. C. Lai (2018). The hpRNA/RNAi pathway is essential to resolve intragenomic conflict to preserve balanced sex ratio. Submitted.
Mohammed, J., A. Flynt, A. Panzarino, M. Mondal, M. DeCruz, A. Siepel and E. C. Lai (2018). Deep experimental profiling of miRNA diversity, deployment, and evolution across the Drosophila genus. Genome Research 28: 52-65.
Kondo S., J. Vedanayagam, J. Mohammed, S. Eizadshenass, L. Kan, N. Pang, R. Aradhya, A. Siepel, J. Steinhauer and E. C. Lai (2017). New genes often acquire male-specific functions but rarely become essential in Drosophila. Genes and Development 31: 1841–1846. (Highlighted in Genes and Dev 31: 1825-1826.)
Lin, C.-J., J. Wen, F. Bejarano, F. Hu, D. Bortolamiol-Becet, L. Kan, P. Sanfilippo, S. Kondo and E. C. Lai (2017). Characterization of a TUTase/RNase complex required for Drosophila gametogenesis. RNA 23: 284-296.
Wen, J., H. Duan, F. Bejarano, K. Okamura, L. Fabian, J. A. Brill, D. Bortolamiol-Becet, R. Martin, J. G. Ruby and E. C. Lai (2015). Adaptive regulation of testis gene expression and control of male fertility by the Drosophila hairpin RNA pathway. Molecular Cell 57: 165-78.


 (1 votes)
(1 votes)