SDB-ISD 2016 Meeting Report
Posted by Aidan Maartens, on 18 August 2016
The joint meeting of the Society for Developmental Biology and the International Society for Differentiation was held in Boston between August 4th and 8th. It was a meeting of firsts for me: first meeting representing the Node, first time at the SDB (having regularly enjoyed the Brit version, I was looking forward to hearing the US angle on what’s exciting at the moment), and first time in Boston. The venue was the gargantuan Copley Place Marriott, which offered a stunning view of the city (from my hotel room at least) as well as a central base to explore. Boston turned out to be beautiful and walkable. Great local beer too.
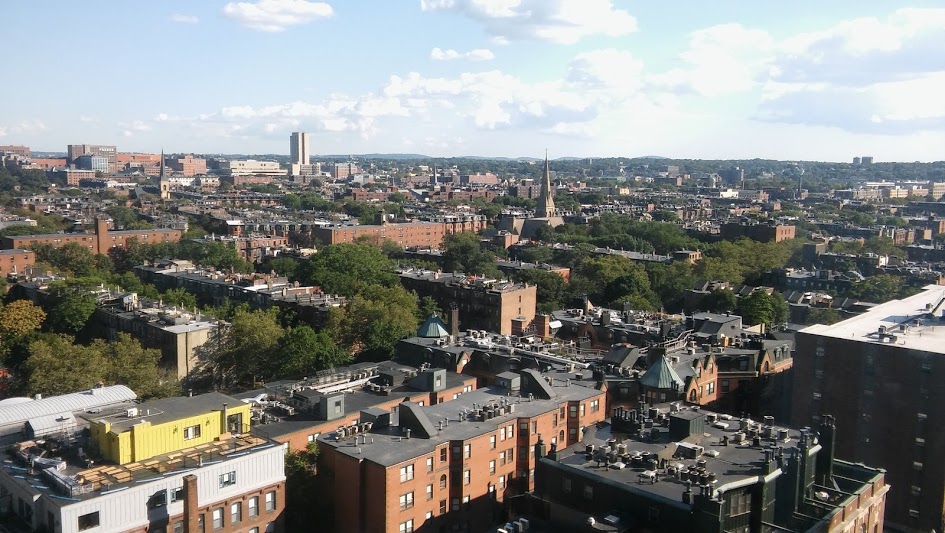

True to our diverse field, the meeting covered a lot of bases, from cell mechanics to genome editing, patterning to computational modelling, as well as life away from the bench (education, publishing, grant-writing) and the planet (designing experiments for NASA’s Genelab). What follows is a selection of my personal highlights from the talks I saw. Let me know if I missed anything or have mangled things beyond comprehension.
Regenerating parts and patterns
I began the meeting in a satellite symposium, Evolution of Regenerative Capacities: recapitulation of development or novel mechanisms? The symposium set the tone for the rest of the meeting in terms of speaker composition: a mix of postdocs, young PIs and established leaders in the field, plus the odd grad student, and an even gender balance.
Most of the speakers started with the same observation: some animals are great at regeneration, some (like us) not so great, and even within the same animals, some tissues regenerate better than others (though axolotls apparently can regenerate everything!). However, we really lack a mechanistic explanation for this spectrum of regenerative capacity; such an explanation might inform our efforts to coax currently intransigent human tissues to regenerate. The evolution part came from the range of species under investigation, which allowed for a fairly broad phylogenetic sample.
For someone fairly naïve to this field, I was struck by the diversity of mechanisms used in the different contexts, as illustrated by a rough list the proteins discussed: piwi piRNA pathway members, various transcription factors, second messengers, growth factors, cytokines, cell adhesion molecules, glycosylation enzymes, and matrix proteins. (I guess this shouldn’t really have been surprising given the tasks at hand for the wounded tissues.) The extent to which regeneration redeploys developmental pathways also seems to depend on the tissue: for instance, Ken Muneoka described how digit regeneration in the mouse shared some features with normal bone development, but not others.
One of the more remarkable stories involved the moon jellyfish, described by Lea Goentoro. Ephyra (juvenile medusa) stage animals typically have 8 evenly-spaced arms which pulsate synchronously to drive propulsion and feeding. Cut off some of the arms, and the ephyra proceeds not to regenerate them, but to reorganise its remaining limbs to regain radial symmetry, in a manner requiring muscle contractility but not cell division. The ephyra regenerates not body parts but a feature of the body pattern, perhaps because repaired animals have still got a reasonable chance of surviving. Got to love nature sometimes.
Gene expression and chromosome topology
After the regeneration fest, Ken Zaret kicked off the Presidential Symposium by celebrating developmental biology, and insisting on the continued need for basic research in model systems (his reaction, perhaps, to recent events in the US and Canada). He then stated that, for him, changes in gene expression are the real heart of development (the symposium did have an epigenetics theme, after all). Ali Shilatifard, in his exploration of histone methylation and childhood leukaemia, agreed. Coming from a bit more of a cell mechanics background, I wanted to come up with a counter argument, but by that point didn’t have the brain power for it.
One of the most remarkable epigenetic feats is the inactivation of an entire X chromosome in female mammals, providing almost complete gene silencing from early development onwards. The long non-coding RNA Xist is key to inactivation, and Jeannie Lee described how RNA proteomics had identified the proteins that interact with Xist (it turns out there are a surprisingly large number of them), and how super-resolution imaging had visualised individual Xist molecules decorating the inactive X (it turns out there are a surprisingly small (~100) number per X). An analysis of Xist mutants then implicated other silencing mechanisms acting synergistically with Xist, and showed that even modest changes in X gene dosage can have profound effects on viability.
Revelations about the X chromosome resulting from a combination of cutting-edge technology and genetics…history repeated itself the next morning, in Job Dekker’s talk, part of a fascinating Chromosome Topology and Gene Regulation session. Revealing the organisation of the inactive X is complicated by the fact that the two copies of the chromosome are too similar to be distinguishable by HiC. The solution: cross two mice lines, creating a (frankly rather shabby looking) F1 hybrid, which contains enough SNPs to distinguish maternal and paternal chromosomes, and then derive ES cells from these mice and induce X inactivation in them. Dekker showed how these cells revealed the fine-grained structure of the inactive X, with its two highly compact mega-domains, mysterious boundary region between them, and islands of topologically associating domains around the few genes that remain active.
Elsewhere in the session, Alistair Boettiger gave us another example of how super resolution imaging is transforming our view of biology: he was able to distinguish chromatin domains, and their features, by ‘painting’ chromosomes with fluorescent oligos, and imaging with 3D-STORM. It was so impressive to actually see some of the different chromatin domains Ken Zaret and Ali Shilatifard had talked about the previous night.
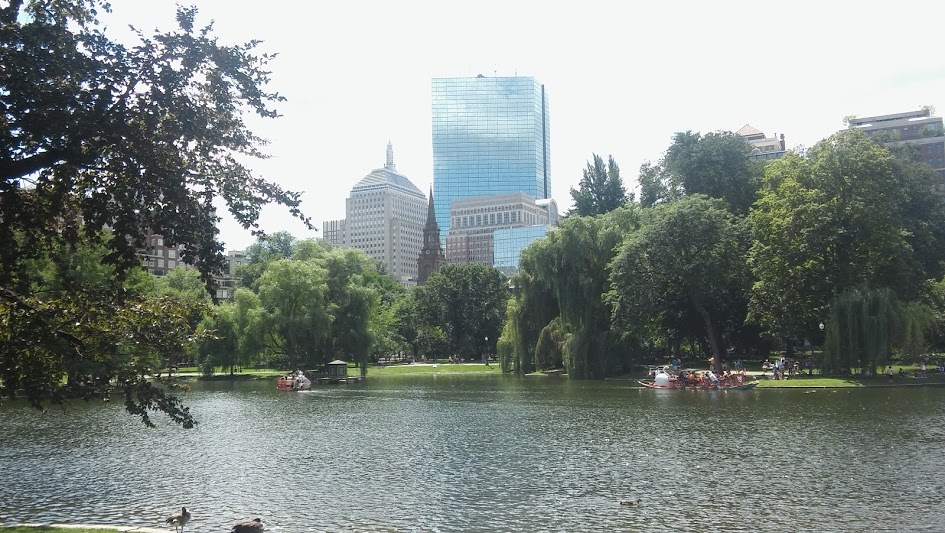

Postdocs and single cells
After a delightful interview with Dave McClay (look out for it in an upcoming Development issue and on the Node), and an hour jaywalking around Boston for lunch, I was back for the Hilde Mangold Postdoctoral Symposium.
From a very strong field (the eight talks were picked from over a hundred applicants), two really stuck with me. Yaniv Elkouby, from Mary Mullins’ lab at UPenn, documented the intricate relationship between the centrosome, telomeres, a chromosomal ‘bouquet’, microtubules, and the Balbiani Body in the polarisation of the fish oocyte. Beautiful in vivo cell biology. Jacqueline Tabler (John Wallingford lab at UTexas), pointed out that given how critical vocalisation is for vertebrates, it’s striking how little we know about the development of the organ that creates vocalisations, the larynx. Aided by her own whistling and beautiful images, she told us how interfering with Hedgehog signalling led to an expansion of neural crest-derived tissues in the larynx, and defective vocalisations in newborn pups. (Her previous work has looked at various aspects of craniofacial development.)
Friday afternoon brought another plenary: Biology at the level of single cells. Single cell sequencing is a remarkable advance, but sequencing for the sake of sequencing does not necessarily advance the field; you need a strong biological question. It was therefore wonderful to hear Mark Krasnow reconstruct the program of alveolar development by sequencing individual cells of various stages from a dissociated lung. With exquisite sensitivity (‘a single RNA in a single cell’), the technique revealed new players in the developmental program by correlating expression with differentiation markers, providing a quick, non-genetic screen.
Tony Hyman then took us back to basics and asked: how does the cell organise chemical reactions without membranes? Part of the answer appears to be liquid-liquid phase separations. Unstructured protein regions (for instance in prion-like domains) promote phase transitions, and changing the sequence of these regions could alter the properties of the resulting agglomeration, from liquids that have fast turnover and promote chemical reactions, to gel-like, stable, dense matrices prohibitive of reactions (such as the Balbiani body we encountered in Yaniv’s talk earlier in the day). The talk really made me think differently about the way cells work.
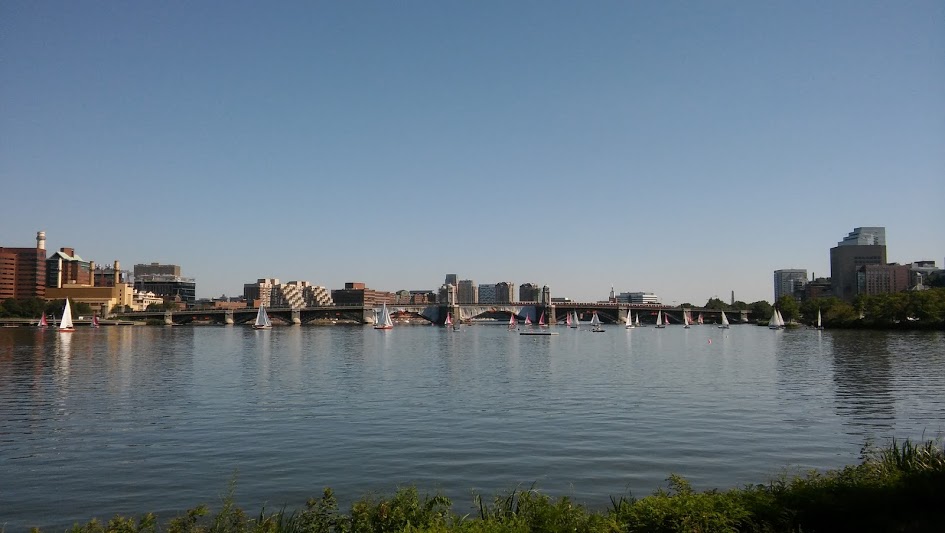

Fat fish and organoids
I started Saturday in the Emerging model organisms and evo-devo session. The general feeling was that techniques like CRISPR were opening up many organisms to developmental analysis; your question might now dictate your model, rather than the other way round. (Lo and behold: a recent piece in Nature.)
Nicolas Rohner introduced us to Astyanax mexicanus, the blind cave fish, which has emerged as a model for metabolic diseases. These fish are always hungry, get really fat (without any clear health disadvantages), and are remarkably resistant to starvation. They are also an evolutionary windfall: not only do river forms persist (providing an ‘ancestral’ population that blind forms can still breed with), but the cave forms themselves have evolved multiple times independently, so they are a great model for convergent evolution.
During the coffee break, I interviewed Doug Melton – another wonderful half hour, look out for the write-up in the future – prior to his Jean Brachet lecture, in which he promoted the idea of applied developmental biology in harnessing stem cells to treat diabetes. Applied developmental biology is also being advanced by organoids, which were celebrated in a special plenary session in honour of Yoshiki Sasai. One recurrent problem with organoids is the variation you get between samples, and Jürgen Knoblich described how growing brain organoids on strands of a synthetic polymer reduces variability, and promotes development of a cortical plate-like structure . Hans Clevers discussed modelling cancer with tumour organoids (somewhat counterintuitively, tumour-derived organoids always grow slowly), and promoted the idea that, in many cases, stem cells are not a rigid cell fate, but a labile cell state.

Circumventing Mendel and celebrating scientists
Sunday: the final day; bleary-eyed and near-saturated, but not dead yet. Before the SDB award lectures, I heard from Ethan Bier on the ‘mutagenic chain reaction’, a CRISPR/Cas9 technique which gives a rapid form of non-Mendelian inheritance. Not only does this drastically cut the time it takes to make mutants (hypothetically, to make a mouse quadruple mutant: 1/64 mice in the 4th generation of a standard cross; 64/64 in the 2nd generation using MCR), and provide a tool for organisms without established genetic techniques, it also allows you to rapidly spread a genetic element through a population (a gene drive). Stunning, revolutionary genetics.
The afternoon Education Symposium was a change of gears, as educators discussed how to provide biological training in institutions with shrinking budgets. The general consensus was that inquiry-driven learning could provide a more engaging route for biology students than your standard rote lab work (this report sums up a lot of the speakers’ ideas). Getting undergrads to feel a sense of ownership over the projects also helps: Sarah Elgin described how hundreds of undergrads had contributed to the annotation of the oft-forgotten dot chromosome of Drosophila melanogaster (leading to a paper with over a thousand authors!).
And finally, the SDB awards. Ida Chow (Viktor Hamburger Outstanding Educator Prize) encouraged all of us to fight for science and science education, and got a raucous standing ovation (from a show of hands, many in the audience had directly benefited from one of her many initiatives). We got to see old pictures of Dave McClay (Lifetime Achievement Award) as a high school football player and an underwater diver, before he described how each of the EMT events in the early sea urchin is driven by a separate gene regulatory network. And finally, we heard from Kathryn Anderson (Edwin G. Conklin medal; I also had the pleasure of interviewing Kathryn, keep an eye out for it) who described the beauty and mystery of gastrulation in the mouse.
The evening ended with drinks, poster awards, dinner (including waiters impatient to get you on to your next course: “coffee with your chicken, Sir?”), drinks, dancing, watching-of-the-dancing, and then final drinks in a dive bar (Cambridge really lacks places like these).

I left the meeting with the feeling that developmental biology is in a good state. This was not just down to the general positivity of Americans: new technologies are unlocking doors everywhere, and the increasing applicability of developmental research to human health (with stem cells, organoids) gives a public persona that I hope will draw in more money without side-lining all of the strong non-applied research that sits under the DB umbrella. Coming from England, you also forget how vast the US is and how much wonderful research is being carried out there.
I would also like to commend all of the speakers who suffered technical difficulties and kept their poise to finish their talks, rather than melt down and destroy the equipment (in one of the meeting’s funniest moments, Mina Bissell walked the length of the hall to fix her frozen computer at the back AV desk, all the while giving running commentary).
And finally: the posters. More than 500, each up for one night only, they were excellent too, and included many with added extras (3D printed models, interactive whiteboard sections, and built-in videos!).
Quotes of the meeting (paraphrased, potentially misremembered)
“Three cheers for developmental biology!”
Ken Zaret opening the meeting, cue cheers and applause.
“This is for you, Sarah Palin…”
Ali Shilatifard on how research in flies helped to inform children’s cancer treatment
“Like Republicans, Aedes aegypti ‘repeal and replace’”
Marc Halfon on mosquito GRN evolution (not that I immediately got the reference)
“It is not birth, marriage, or death, or gastrulation, but epithelium formation that is truly the most important time in your life”
Matthew Gibson takes Wolpert back a step
“If your grandfather was a fish, he’d have no problem hearing you”
Shawn Burger on the amazing regenerating ear hair cells of zebrafish
“I just love the spiny mouse!”
Overheard at the Regeneration symposium (before I found out that they were a species, not a Pokemon)
“Blind cavefish are fat”
Nicolas Rohner pulls no punches in describing his beloved model organism
“We must build the embryo to understand it”
Eric Siggia goes a bit Feynman
“If you’re not arrogant, you can look at your data and see what it’s trying to tell you”
Mina Bissel warning against over-confidence
“There are no ‘higher’ or ‘lower’ organisms”
Richard Berhinger on Twitter
“Other hashtag is … different”
James Gagnon on Twitter, after confusion over #2016SDB (the correct one) versus #SDB2016 (Seventh Day Baptists)


 (4 votes)
(4 votes)