BSDB Gurdon/The Company of Biologists Summer Studentship Report #22 – Annabel Adams
Posted by BSDB, on 17 December 2018
Established by the British Society for Developmental Biology in 2014, The Gurdon/The Company of Biologists Summer Studentship scheme provides financial support to allow highly motivated undergraduate students an opportunity to engage in practical research during their summer vacation. Each year, ten successful applicants spend eight weeks in the research laboratories of their choices, and the feedback we receive is outstanding. You can read accounts from previous years here.
Our first report from the 2018 group of student awardees comes from Annabel Adams (student at King’s College London), who undertook her project with Andrea Streit, also at King’s.
As a second-year neuroscience undergraduate, I wanted specialist involvement in a working lab to progress my knowledge beyond the practical classes offered on my course. The Gurdon Studentship provided the support which allowed me to translate my interest in research into tangible experience. I was therefore very grateful to be able to join Andrea Streit’s lab at King’s College London for eight weeks. During my studies, my embryology module caught most of my attention. In particular, part of my coursework focused on a paper that used the chick model to observe the development of craniofacial features. When reading around the subject I encountered various techniques used in chick and wanted to see their implementation first hand. Therefore, the Streit lab was ideal because not only do they use some of these methods, but they were also able to take me on as an intern.
One of the objectives of the Streit lab is to investigate the mechanisms behind how progenitor cells become committed to a certain lineage, specifically focusing on sense organs like the ear. They have previously characterised a circuit of eight transcription factors at the top of the gene network that governs the way in which cells progressively commit to ear identity. (Chen et al., 2017)
To build upon this discovery, the question my project aimed to address is whether or not this transcription factor module is active elsewhere in the chick during development, and if so how its architecture changes. In humans mutations in some of these factors not only result in deafness, but also in kidney and limb abnormalities. This suggests these regions could share common features with the ear module. To begin to establish if this is the case, I first performed in situ hybridisation (ISH) on a range of chick stages in order to analyse the expression patterns of these factors. I used antisense probes complimentary to the mRNA of each of six transcription factors present in the ear module: Lmx1a, Prdm1, Sox8, Sox13, Pax2 and Zbtb16. The probes were labelled with DIG (digoxigenin) allowing the use of anti-DIG antibodies followed by a colorimetric reaction to detect where each gene is expressed.
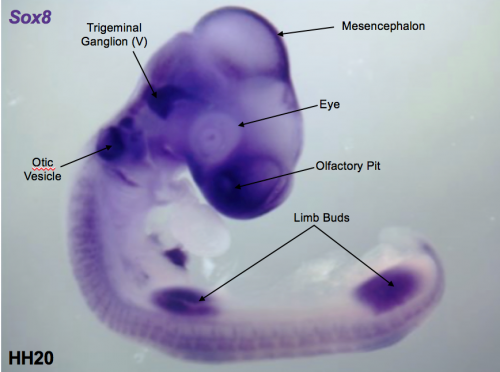
The procedure itself is relatively straight forward. However, the process is quite slow since several steps require a long time to complete. Therefore, planning and managing time pressure is one of the main challenges I came across. This was apparent when juggling several experiments at once. At first, it was a difficult to balance the fragility of the samples with furiously pipetting washes to get them in the incubator before the day’s end. The ability to multitask was a skill I had not previously considered to be important as a scientist. However, my experience has helped me see how I need to improve my organisation in order to get the most out of lab hours. Learning this lesson early stands me in good stead for my research project in third year. Eventually, I became more efficient at managing my time which lead to me feeling able to attempt a more complicated procedure. Having established expression patterns, the next challenge was to assess whether some of the regulatory relationships in the ear circuit are maintained in other organs. For example, the Pax 2 enhancer that is active in the ear, but does it also show activity in the kidney or limb? This was accomplished through the electroporation of a reporter plasmid, where the enhancer drives GFP, followed by fluorescence imaging. This involved successful embryo culture, plasmid injection and transfer of the plasmid using a current – many steps involving numerous opportunities for things to go wrong. The samples were a challenge to handle owing to their small size. Therefore, it took several attempts to build up enough dexterity and confidence to execute each step properly. Initially, it was frustrating when most of the cultures didn’t survive or the plasmid injections missed their target. Yet, this was outweighed by the satisfaction of when it all came together and I saw the fluorescence through microscope signifying my first successful electroporation.

Overall, an internship was a big time commitment but one that was invaluable in helping me validate my desire to pursue a career in research myself. My time in a supportive working lab has taught me not only how to obtain results but also how to interpret them. Each day I was becoming more accustomed to techniques and equipment simply through practice. However, through engaging with the lab and their work as a whole, both in formal meetings and in conversation, I became more familiar with what a scientist looks for in their experiments, and crucially, assessing how what they found could apply to a wider context. I have learnt the importance of being aware of the work outside one’s own. As a student, going beyond a surface level understanding of what you are doing begins with immersing yourself in the field. An internship in a lab working on a project you have interest in is a great way to start.
References
1. Chen, J., Tambalo, M., Barembaum, M., Ranganathan, R., Simões-Costa, M., Bronner, M. and Streit, A. (2017). A systems-level approach reveals new gene regulatory modules in the developing ear. Development, 144(8), pp.1531-1543.



 (No Ratings Yet)
(No Ratings Yet)