This month in preLights – August
Posted by preLights CoB, on 5 September 2018
Welcome to our monthly summary of developmental biology (and related) preLights.

preLighters are early-career researchers who select and highlight preprints which they feel are interesting for the life-science community. While writing highlight posts is mostly an individual effort, plenty of interactions between the preLights team members take place on our Slack channel. This is where last month several preLighters decided to respond to a controversial World View article in Nature about the danger of preprints, and they shared their well-argued (and highly read) commentary here on the Node. Apart from that, the month of August was not short of exciting developmental biology (and related) preLights, we hope you enjoy this selection!
Gene expression and cell fate
There was a good mix of preLights dealing with gene expression in various models, such as fish, flies and living neuronal tissue. Idoia Quintana-Urzainqui discussed a preprint showing that in zebrafish embryos, this first wave of transcription is restricted to a special nuclear compartment. Later on in development, transcription factors, which are often tissue-specific, regulate cell fate. Amanda Haage highlighted how the progression from pluripotent blastula cells to neural crest cells is regulated by a transition from SoxB1 to SoxE transcription factors in zebrafish. Moving to flies, Clarice Hong’s and Natalie Dye’s preLight both dealt with transcription factor function. Clarice discussed how binding site orientation and spacing determines the activity and specificity of Hox transcription factors, while Natalie reported on how the transcription factor Doublesex is involved in the decision making of male vs. female gonad stem cell niches. Finally, Theresa Rayon illustrated how cells make transitions during neurogenesis. The study she preLighted imaged the Hes5 transcription factor in real-time and showed that its fluctuating expression in neuronal progenitors turns oscillatory as cells enter differentiation.
Neurodevelopment and morphogenesis
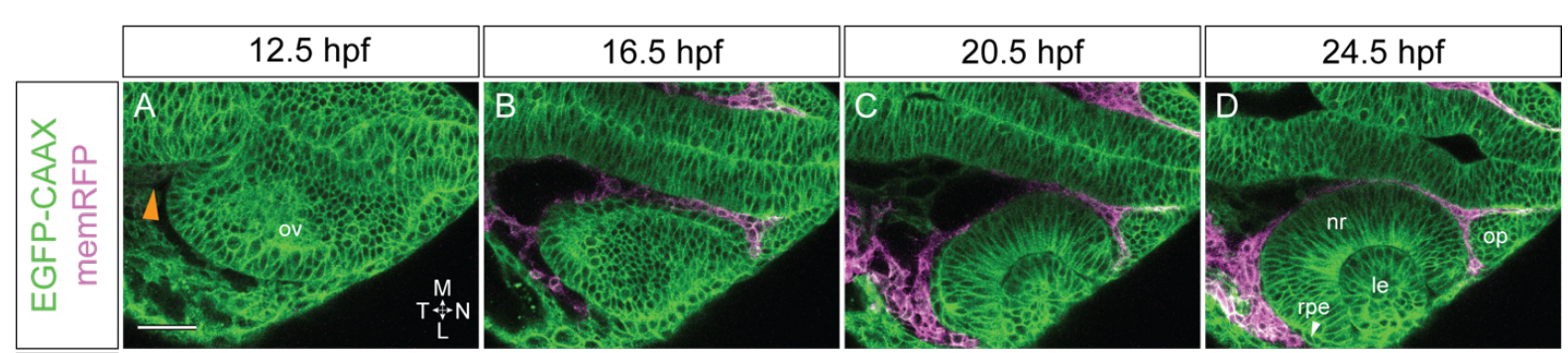
Neurogenesis during development of the cortex was the focus of Boyan Bonev’s preLight. By using a new labelling approach, the study uncovered the temporal order in the production of diverse neuronal subtypes and found that many of them emerge simultaneously. Ashrifia Adomako-Ankomah highlighted the spectacular morphogenetic events occurring during development of the eye. The research identified an important role for the protein nidogen, secreted by the surrounding neurocrest cells, in shaping the optic cup in zebrafish. Nidogen also featured in the preLight of Nargess Khalilgharibi, and it turned out that this protein is not essential for Drosophila morphogenesis, but still has tissue-specific functions. Lastly, Andreas van Impel reviewed how the local translation of a transcript at the leading edge of migrating endothelial cells regulates blood vessel formation.
Tools & Technologies
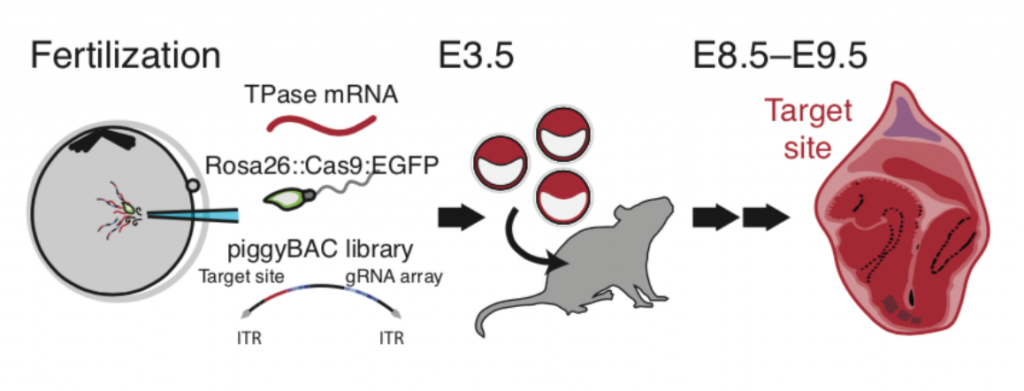
Undoubtedly, new methodologies are key in paving the way for discoveries in developmental biology. Hannah Brunsdon covered a technique with enormous potential, in which the authors combined CRISPR lineage tracing with single-cell RNA sequencing to decode early mammalian development. Tools to regulate gene expression are highly desired not only for basic research, but also for potential future gene therapies. Tim Fessenden highlighted a clever technique to control the level of transgene expression by integrating synthetic miRNA target sites into the transgene. Tuning gene expression by changing the regulatory 3D interactions was the focus of Ivan Candido-Ferreira’s highlight, which showcased the use of CRISPR technology in combination with optogenetics. While optogenetics is broadly used in many experimental systems, it seems tricky to get it to work in some organisms, such as the C. elegans embryo. Angika Basant discussed an approach in her preLight that showed how such a limitation (in this case germline silencing) could be overcome. Lastly, Samantha Seah preLighted the development of a microfluidic device that allows imaging of sea anemone larvae, which are models for coral symbiosis. Make sure to read the authors’ comments who tell us about how the project was created.
Do also visit the preLights website to discover further interesting preprint highlights, such as Gautam Dey’s post on a statistical tool that provides an alternative to the p-value , or Nicola Stevenson’s highlight on the horizontal transfer of RNAs in honeybees!


 (3 votes)
(3 votes)