Two Neuro-PhD Positions in Manchester: Mathematical Modelling & Neurodegeneration
Posted by Andreas Prokop, on 21 November 2015
Closing Date: 15 March 2021
 Two positions are available as part of two Research Council-funded doctoral training programmes at The University of Manchester, the first one supported by the BBSRC and the second one by the MRC. Both projects involve work on the fruit fly Drosophila as a highly efficient and relevant model organism to study fundamental mechanisms of neuronal ageing and degeneration with unique detail and depth and delivering understanding of high biomedical applicability. Further information about fruit flies as a model organism is available here. Note that Andreas Prokop, who supervises on both projects, drives active programmes of science outreach and public engagement (see here), and students will have unique opportunities to develop transferable skills in science communication, which is of increasing relevance in modern science and an important category on a researcher’s CV. Full details on how to apply for these positions can be found on the UoM BBSRC DTP website.
Two positions are available as part of two Research Council-funded doctoral training programmes at The University of Manchester, the first one supported by the BBSRC and the second one by the MRC. Both projects involve work on the fruit fly Drosophila as a highly efficient and relevant model organism to study fundamental mechanisms of neuronal ageing and degeneration with unique detail and depth and delivering understanding of high biomedical applicability. Further information about fruit flies as a model organism is available here. Note that Andreas Prokop, who supervises on both projects, drives active programmes of science outreach and public engagement (see here), and students will have unique opportunities to develop transferable skills in science communication, which is of increasing relevance in modern science and an important category on a researcher’s CV. Full details on how to apply for these positions can be found on the UoM BBSRC DTP website.
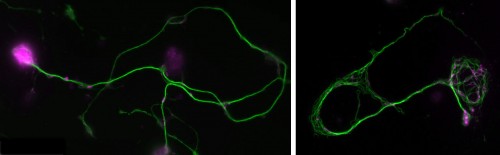
Position 1: Advanced imaging and mathematical modelling of ageing and neurodegeneration in the nervous system
Enquiries: Andreas.Prokop@manchester.ac.uk
Project: Do you have mathematics skills, are keen to combine them with biological research and to work on a novel, highly interdisciplinary project investigating processes of ageing and degeneration in the nervous system? To carry out this exciting research you will be trained in and apply advanced live imaging, electron microscopy (EM), tomography, genetics and mathematical/computational modelling.
You will study axons, which are the cable-like (~1mm in diameter, metres-long!) extensions of neurons wiring the nervous system. These delicate structures are maintained for many decades in humans. They are key lesion sites in spinal cord injury, trauma and many neurodegenerative diseases (e.g. spastic paraplegia), and we lose 50% of our axons during healthy ageing. The essential structural backbone of axons is formed by continuous bundles of filamentous protein-polymers called microtubules. Disorganisation of these microtubule bundles leads to axon swellings correlating with axon decay, but the underlying mechanisms preventing/causing disorganisation are not at all understood.
To gain this understanding, you will genetically induce MT disorganisation, and use imaging to capture volume and space relationships (EM) as well as the dynamics of MT disorganisation processes (live imaging). Based on your collected data and existing algorithms for MTs, you will develop computational/mathematical models describing MT disorganisation and deduce the underlying rules.
Supervision:
Outstanding supervision and training will be provided through an interdisciplinary consortium of specialists with longstanding expertise in the research areas and technologies/strategies involved in this project:
- Andreas Prokop (developmental and cellular neuroscience, Drosophila genetics, live imaging, electron microscopy)
- Simon Pearce (mathematical modelling of biological systems)
- Matthias Heil (mathematics)
- Karl Kadler (electron microscopy, tomography, high pressure freezing techniques)
Note, that Andreas Prokop and Matthias Heil are also members of the Wellcome Trust-funded PhD programme “Quantitative & Biophysical Biology” providing further opportunities in this area of research.
Literature:
- axons: Prokop, 2013, Neur Dev 8, 17ff.
- Drosophila neuro-genetics: Prokop, 2013, J Cell Sci. 126, 2331ff.
- mathematics: Ziebert, 2015, Phys Rev Lett 114, 148101ff.
- live imaging: Alves-Silva, 2012, J Neurosci 32, 9143ff.
- EM: Starborg, 2013, Nat Protoc 8, 1433ff.
Position 2: An interdisciplinary approach to unravel mechanistic understanding of Frontotemporal lobar degeneration
Enquiries: SPB@manchester.ac.uk
Project:
Dementia causes enormous personal hardship and costs the UK ~£23 billion every year. The second most common form is Frontotemporal lobar degeneration (FTLD). About 40% of FTLD cases have genetic causes, with >8% involving abnormal intronic GGGGCC hexanucleotide repeat expansions in the C9orf72 gene which can additionally cause motor neuron disease (OMIM #105550). These pathological expansions are actively transcribed and, via bidirectional repeat-associated non-ATG (RAN) translation, generate 5 different aggregate-forming GA, GR, PR, GP and AP dipeptide repeat proteins (DPRs).This project will gain new understanding of this type of FTLD by unravelling neurodegenerative pathomechanisms of DRPs through using interdisciplinary approaches. We will focus on the hypothesis that toxicity is caused by DRP structure, comparable to amyloid plaques in Alzheimer’s disease. The project will capitalise on the complementary expertises of the three supervisors, and a readily available, unique set of 4 GFP-tagged constructs with high, pathologically relevant repeat numbers. The detailed aims and outcomes are:
- To generate purified DRPs and perform biochemical and biophysical analyses, in order to understand the reasons for their toxicity and identify useful therapeutic strategies which will benefit patients and their families.
- To generate transgenic Drosophila fly stocks to obtain primary neurons expressing the four DRPs. We will use powerful fly genetics and well established cell biological approaches to identify the neuronal death pathway and of the DRP-induced pathomechanisms upstream.
- There is substantial proof-of-principle for the use and translational potential of Drosophila To validate identified DRP pathomechanisms in mammalian contexts, we will carry out complementary experiments using well established DRP models in SH-SY5Y cells and inducible neuronal cell lines.
This project is highly interdisciplinary, with the analysis of the dipeptide repeat proteins spanning systems from in vitro solvents to human and fly cell lines. Students will receive a broad training in characterisation of polypeptide structure and aggregation, cell culture, fly and human genetics, and cell biology, all in the context of genetic disease and neurodegeneration.
Supervision:
- Stuart Pickering-Brown (human geneticist specialised on causes of FTLD)
- Andreas Prokop (developmental and cellular neurobiologist; pioneered primary neurons of the fruit fly Drosophila as a powerful model to decipher the fundamental cell biology of neuropathologies)
- Andrew Doig (biochemist specialised on the structural and mechanistic analysis of protein aggregation in the context of neurodegeneration)
Literature
- dipeptide repeat proteins: Mizielinska et al., 2014, Science 345, 1192ff.
- Drosophila neuro-genetics: Prokop, 2013, J Cell Sci. 126, 2331ff.
- structural biology: Doig & Derreumaux, 2015, Curr Opin Struct Biol 30, 50ff.


 (No Ratings Yet)
(No Ratings Yet)