An embryo-maternal dialogue in Arabidopsis
Posted by Yujuan Du, on 20 July 2022
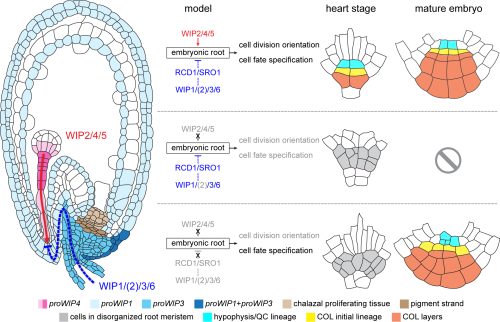
In our recent paper, we uncovered a WIP-mediated embryo-maternal communication in Arabidopsis, which specifically regulates embryonic root development (Fig. 1). WIP2, WIP4 and WIP5 expression in the embryonic root is necessary for the oriented cell division and distal stem cell fates within the root meristem. WIP1, WIP3 and WIP6 are expressed in the maternal tissues surrounding the embryo and suspensor, where they act non-cell autonomously to repress root cell fate specification through SIMILAR TO RADICAL-INDUCED CELL DEATH ONE (SRO) gene family members. The embryonic WIPs functionally oppose those maternal WIPs to orchestrate cell division orientation and cell fate specification in the embryonic root, thereby promoting regular root formation.
Nature Plants: Spatially expressed WIP genes control Arabidopsis embryonic root development
Nature Plants News & Views: A dialogue between generations
How did you get started on this project?
It has been shown that loss-of-function wip245 triple mutants are rootless (Crawford et al., 2015). WIP2, WIP4 and WIP5 are expressed in the hypophysis and its derivatives, where they act redundantly to specify distal root stem cell fates (Crawford et al., 2015). We observed that roots were able to form and grow in wip123456 sextuple mutants, this observation prompted us to explore how WIP genes are coordinated to control embryonic root development.
What was already known about the regulation of tissue organization during root development?
Tissue organization during root development requires precise coordination of root cell division, fate specification and differentiation in a spatiotemporal manner. Components in auxin signaling cascade are crucial for root formation in Arabidopsis: the BODENLOS (BDL)/IAA12-MONOPTEROS (MP)/AUXIN RESPONSIVE FACTOR5 (ARF5) and SOLITARY-ROOT (SLR)/IAA14-ARF7/ARF19 modules are required for the initiation of embryonic and lateral roots respectively (Fukaki et al., 2002; Hamann et al., 2002; Okushima et al., 2005; Przemeck et al., 1996). In terms of embryonic roots, BDL and MP expressed in the proembryo cells non-cell autonomously regulate hypophysis specification via two direct target genes of MP, TARGET OF MONOPTEROS5 (TMO5) and TMO7 (Schlereth et al., 2010). It has been shownWIP2, WIP4 and WIP5 act downstream of MP to promote embryonic root formation (Crawford et al., 2015), but hypophysis specification remains normal in wip245 mutants. Auxin and auxin signaling is also considered important in initiating formative cell divisions during both embryonic and lateral root formation (Marhavy et al., 2016; Yoshida et al., 2014). Moreover, PLETHORA (PLT) genes are key effectors for establishment of the root stem cell niche during embryonic pattern formation (Aida et al., 2004; Galinha et al., 2007). Loss-of-function of plt1-/-plt2+/-plt3-/-bbm-2-/- mutants show embryonic root defects that are morphologically similar to the ones displayed in wip245 mutants.
What was the key experiment?
Our results from three experiments made the story outline: the rescued root development in wip123456 sextuple mutants; the identification of SRO gene family members that are responsible for the overexpression of WIP1 induced plant growth arrest from the EMS mutagenesis; the rescued root development in rcd1-4wip245 quadruple mutants.
When doing the research, did you have any particular result or eureka moment that has stuck with you?
When we found that WIP1, WIP3 and WIP6 act maternally in a non-cell autonomous manner to repress root formation. This result points to a WIP-mediated embryo-maternal dialogue.
And what about the flipside: any moments of frustration or despair?
Generation of the multiple mutant combinations was time consuming and demanding.
Where will this story take the lab?
Our lab focuses on the sex determination process in cucurbits. The gynoecious gene WIP1 represses carpel development, causing the formation of male flowers, and its loss-of-function leads to purely female plants (Martin et al., 2009). In Arabidopsis, melon WIP1 and AtWIPs share common functions in inhibiting plant growth (Roldan et al., 2020), suggesting that the molecular network regulated by AtWIPs is likely conserved in plants. Therefore, we aim to transfer the knowledge gained from Arabidopsis back to the sex determination process in melon.
What next for you/your lab after this paper – let us know if you are continuing this research, starting/looking for a new position?
Currently, I am looking for a position and would like to continue the study of maternal-embryo communication in plants. Abdel’s lab will continue to characterize WIP functions in Arabidopsis and in cucurbits.
References:


 (No Ratings Yet)
(No Ratings Yet)