BSDB Gurdon/The Company of Biologists Summer Studentship Report #29 – Rachel Wong
Posted by BSDB, on 19 December 2018
Established by the British Society for Developmental Biology in 2014, The Gurdon/The Company of Biologists Summer Studentship scheme provides financial support to allow highly motivated undergraduate students an opportunity to engage in practical research during their summer vacation. Each year, ten successful applicants spend eight weeks in the research laboratories of their choices, and the feedback we receive is outstanding. You can read accounts from previous years here.
Our final report from the 2018 group of student awardees comes from Rachel Wong (student at Queen’s University Belfast) who undertook her research with Karen Liu (King’s College, London).
Analysis of Rapgerf5 and canonical Wnt signalling in embryonic mouse development
During the summer of 2018, I worked with Dr John Griffin in Dr Karen Liu’s lab at King’s College London. My focus was on the gene RAPGEF5, which was previously identified as a candidate gene for heterotaxy, a congenital disease affecting heart development and the spatial arrangement of organs. It is estimated that 1 in 10,000 people are diagnosed with heterotaxy, and is the cause of 3% of all congenital heart cases [1]. However, the genetics of heterotaxy are still unclear. Thus research is necessary to understand the disease mechanism in more detail.
Not much is known about RAPGEF5 protein, but we know it is involved in the canonical Wnt pathway, in the transportation of beta-catenin into the nucleus [2]. When Wnt is active, a cascade of chemical reactions prevent the degradation of beta-catenin in the cytoplasm, allowing it to bind to a transporter protein to enter the nucleus. Our current model suggests that in response to Rap-GDP conversion to Rap-GTP by RAPGEF5, beta-catenin can dissociate from the transporter protein. This frees beta-catenin, allowing it to interact with DNA-binding proteins to alter gene expression. Therefore, my project aimed to answer three key questions:
- Where is canonical Wnt signalling active during embryonic development?
- Where is RAPGEF5 expressed during embryonic development?
- Does loss of RAPGEF5 lead to any developmental abnormalities such as heterotaxy?
To answer my first question, I used genetically modified mouse embryos carrying a TCF/Lef-dependent reporter to visualize the areas with active Wnt signalling. When Wnt is active, the transcription factors TCF/Lef are active and bind to specific binding sites on DNA. This activates a promoter that causes the GFP reporter gene to be expressed and produce proteins that fluoresce under a specific wavelength.
I dissected embryos from their sacs at weeks E11.5, E12.5, E13.5 and E14.5. They were then photographed using a fluorescence microscope. In general, fluorescence can be seen in the ears, edges of limbs, spine, branchial arches, whiskers and brain. The heart was only visible in the E11.5, as the skin over the heart was too thick at later stages (Figure 1.a). At week E14.5, fluorescence was only faintly visible at the ear, due to the thicker skin.
In response to question 2, I fixed and dehydrated the E11.5 embryos for whole mount mRNA in situ hybridization. A Rapgef5-specific probe was used to stain the embryo, and the result was shown in figure 1.b. RAPGEF5 mRNA was expressed in the heart, brain, spine and the tip of the hind limb.
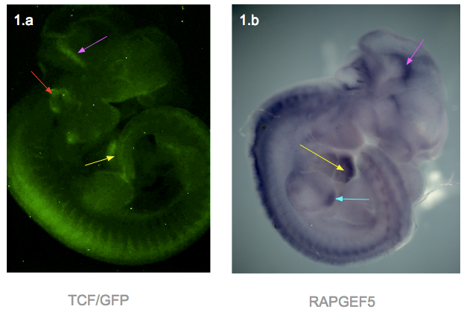
Figure 1 presents two photographs of an E11.5 embryo, one showing the distribution of RAPGEF5 specifically, and the other for active Wnt signalling. When compared, the distribution appears similar, however there are discrepancies such as the ear, and tip of the hind limb.
Finally, for my third question, we bred RAPGEF5 mutant mice and inspected them at stages E9.5, E10.5, E14.5 and 6 weeks after birth. Tail clippings were taken from the embryos and ear clippings from the pups for DNA extraction and PCR to confirm their genotype, as they could either be wild-type, heterozygous or homozygous. Unfortunately the genotyping was still in the stages of trial-and-error, as the bands in the gel electrophoresis did not match the reference DNA ladder, and further tweaking with the PCR temperature and primers is necessary.
However, there were some phenotypic changes found. Out of the total of 9 E9.5 amniotic sacs, 4 were had healthy embryos, 4 were empty and 1 was malformed and underdeveloped. It is possible that this embryo was in the process of being reabsorbed to match the 4 other empty sacs. For the E10.5, there were 8 embryos in total, and 2 had underdeveloped heads and lacked proper surface morphology. All 8 of the E14.5 embryos were phenotypically normal.
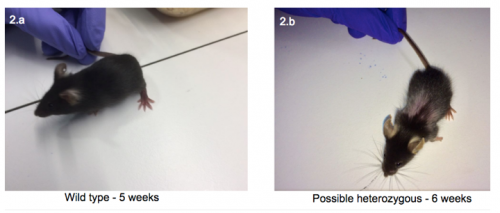

Interestingly, as seen in figure 2, when compared with the wild-type, the 6 week old pup had bald patches in a ‘Christmas tree’ pattern and a possible front limb deformity. Both were similar in size and behaviour.
I thoroughly enjoyed my summer studentship at King’s, and learnt many new techniques such as wax sectioning and mounting, in situ hybridisation and using fluorescence microscopy. I would like to thank Mr John Griffin, Dr Karen Liu and the Liu lab for taking time out of their schedule for their help and guidance.
References
[1] Genetics Home Reference. (2018). Heterotaxy syndrome. [online] Available at: https://ghr.nlm.nih.gov/condition/heterotaxy-syndrome [Accessed 6 Aug. 2018].
[2] John N. Griffin et al. RAPGEF5 Regulates Nuclear Translocation of β-Catenin. Developmental Cell 2018; 44(2)



 (No Ratings Yet)
(No Ratings Yet)