A day in the life of a ctenophore lab
Posted by Burkhardt Group, on 1 September 2016
Who are we?
Hi, my name is Ruairi Kavanagh and I’m a Master’s student at Plymouth University. For my dissertation I am based in The Marine Biological Association (MBA). I am carrying out my research in the recently established Burkhardt Lab. Our lab’s research is focused on tracing the origin and evolution of synaptic proteins, to better understand how synapses and neurons evolved in animals. Strikingly, many of the building blocks of animal synapses originated before the first synapses and neurons evolved. We work on a number of model organisms, including choanoflagellates, sponges, cnidarians, and of course ctenophores (Figure 1). We use choanoflagellates, the closest unicellular relatives of animals, to better understand which synaptic signalling machineries for neuronal functions have been co-opted. Sponges, basal animals with no synapses and neurons, are used to elucidate the origin of synapses and neurons. Ctenophores and cnidarians, basal animals with synapses and neurons, are helping us to answer the question, did synapses and neurons originate once or multiple times independently? Typical analyses are centred on identifying the function of key “neuronal” proteins in these organisms. I work on ctenophores and have identified homologs of synaptic proteins from a transcriptome of the ctenophore Pleurobrachia pileus, a local species that is found in Plymouth sound.
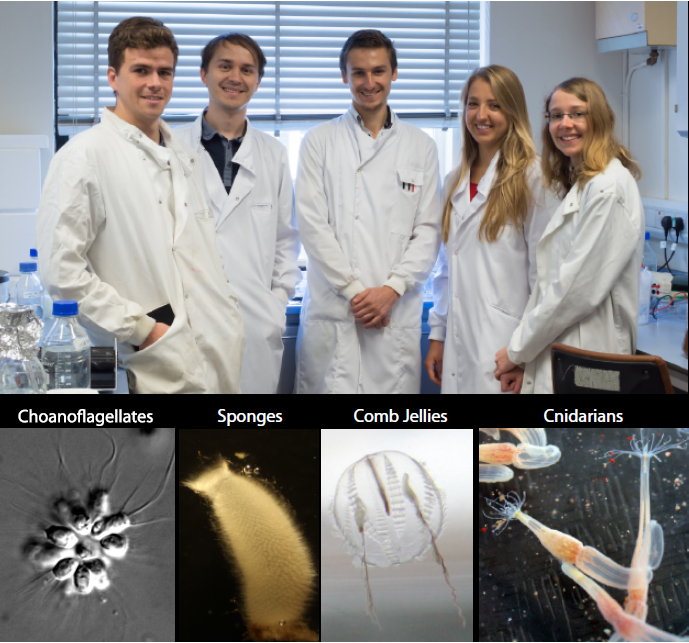

Ctenophore characteristics
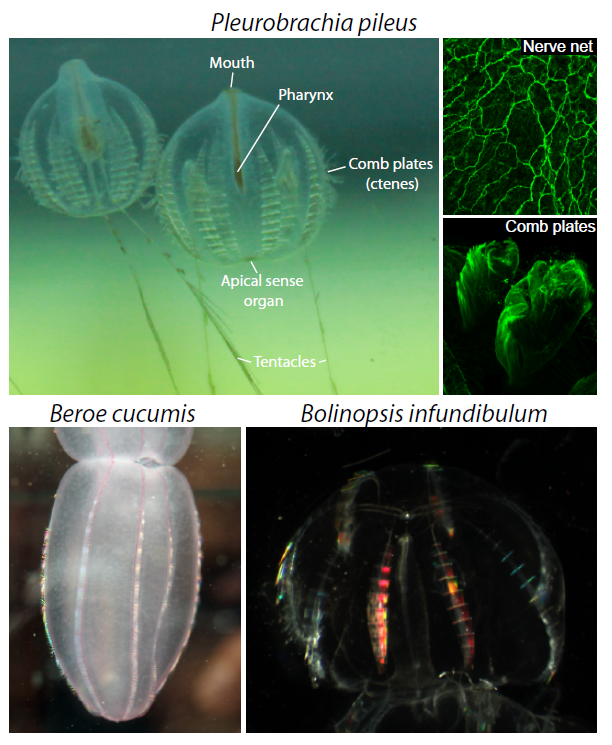
Ctenophores or “comb jellies” are fantastic creatures who have recently come into the evolutionary limelight. There are about 150 known species, almost all of which are pelagic marine predators. Comb jellies contain eight longitudinal rows of ciliated “combs” (or ctenes – ctenophore means “comb-bearing”), which are a distinct apomorphy of the phylum (Figure 2). A nerve net envelops the animal in a kind of polygonal mesh, which is most densely concentrated at the apical sense organ (also called the statocyst). The apical sense organ is thought to detect stimuli such as light, gravity and pressure, to coordinate beating of the ciliated combs, and to control the animal’s orientation. At the oral end, the gastro-vascular tract is connected to the mouth, via a pharynx. At the aboral end there are usually two anal pores, on either side of the apical organ. Interior canals distribute nutrients around the body. Ctenophores are generally found with two tentacles, but some species have secondarily lost the need for them. If tentacles are present, they are lined with sticky cells known as colloblasts, which are used in prey capture (and are another apomorphy). The tentacles of the sea gooseberry, Pleurobrachia (pileus), can be 10-15 times the length of its body. Watching them fish for prey really is an amazing sight. They will extend out their tentacles like big nets and sit in the water column waiting for something to swim into their trap. If they feel the slightest contact, they instantly retract the tentacles towards the mouth, which also triggers them to do a sort of death roll, as they suck down whatever it is they have caught. The two other species that occur here in Plymouth are Beroe cucumis and Bolinopsis infundibulum (Figure 2).
Why study ctenophores?
For a long time, ctenophores were considered to be closely related to cnidarians, since they look superficially quite similar, are both gelatinous, contain comparable tissue-level organization, and perform similar ecological roles. However on closer inspection it is clear that there are significant differences between the two phyla. The most obvious difference between them has to do with locomotion. If we look at jellyfish, they move by contractions of the “bell”, and can only move in one direction. Whereas I have already mentioned that ctenophores use their ciliated combs. The coordinated beating of these combs enables the animal to swim forwards or backwards, and allows for a very efficient way of rapidly changing their orientation.
The coordinated beating of combs from the ctenophore Beroe cucumis. Video taken by Ruairi Kavanagh and Pawel Burkhardt (MBA).
In addition, recent phylogenetic studies proposed that ctenophores might after all not be as closely related to cnidarians as previously thought. And here is why: the genomes of two different ctenophores have recently been sequenced. The first ctenophore genome sequenced was the one from Mnemiopsis leidyi, and the second from Pleurobrachia bachei. These studies, using whole genome sequences, showed that ctenophores and cnidarians are actually not closely related. But even more strikingly, these studies showed that ctenophores, and not sponges, are the sister-group to the rest of animals. If true, this would have important implications: for example, as all ctenophores possess synapses & neurons, in contrast to sponges, an independent origin of synapses and neurons in this phylum is possible. Indeed, neuronal components that are found widely in other animal groups appear to be missing from the genomes of P. pileus and M. leidyi, and many neuronal components do in fact not localize to neurons, but to other, non-neuronal cells. Other researchers have called for more detailed phylogenetic analyses and the need to sequence more ctenophore genomes before such contrary conclusions can be drawn. Ctenophores have, after all, been in the spotlight for only a short period of time. Our work will provide valuable insights into ctenophore biology, and will hopefully help to resolve the current dispute on the origin of synapses and neurons.

A day in the lab
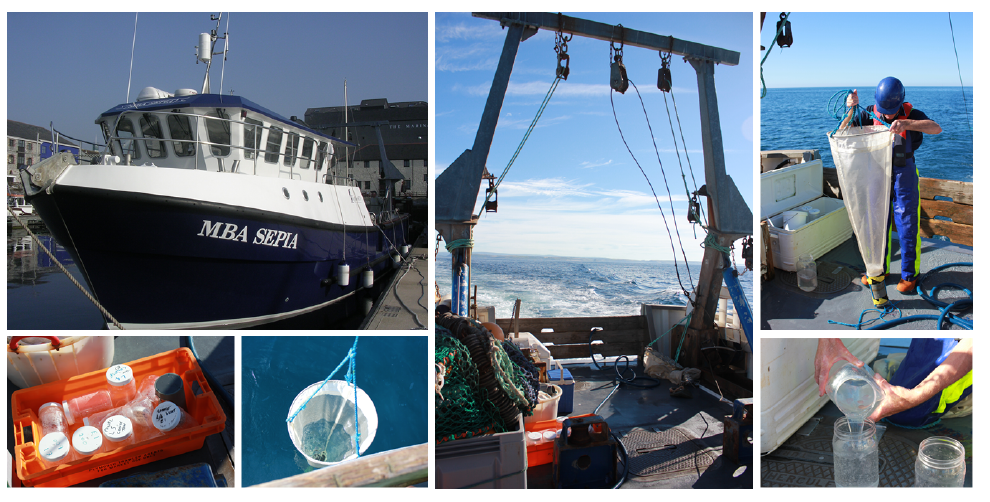
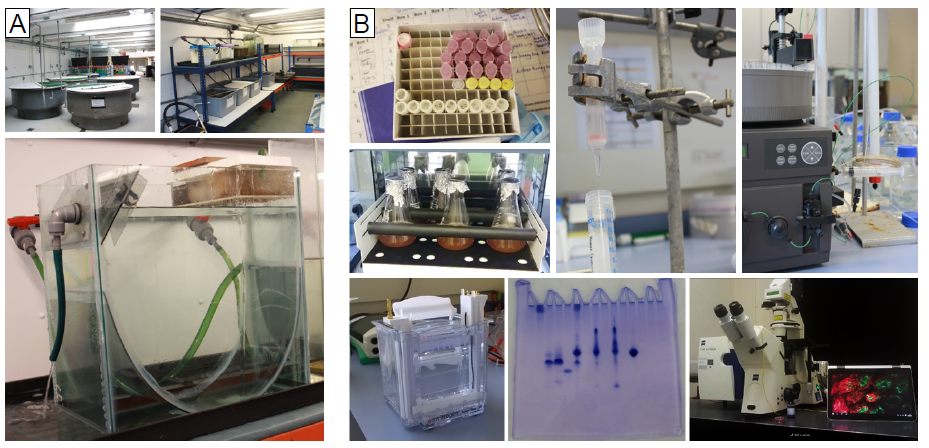
A typical day for us, as you might have guessed, begins at the crack of dawn. The first job of the day is to go ctenophore hunting. For this we use our research vessel named MBA Sepia (Figure 3). The MBA Sepia can accommodate 12 passengers and offers a spacious laboratory. Ctenophore sampling is done using a standard plankton net, and we do both vertical and horizontal trawls (Figure 3). Ctenophores are made up of 99% water and 0.4% salt so one can imagine how difficult it is to take them from the sea, and get them into culture intact back at the lab. In fact, it is difficult to do anything with these buggers! But we persevere. The MBA houses a state-of-the-art seawater facility for marine biological research with a wide range of tanks and we culture ctenophores in so called pseudo-kreisels (Figure 4A). Basically this is a tank that has fresh sea water pumped in from one end and a filter at the other. A cyclical current naturally occurs, which the ctenophores love. One important lesson (that has been learned the hard way) is that you don’t put Pleurobrachia and Beroe together in the same tank, as Beroe is one of Pleurobrachia’s main predators. Ideally I like to leave Pleurobrachia in the tank for 24 hours or so, as this allows them time to recover from the trauma of being handled. They are then ready to use for my experiments, for example for live imaging microscopy and immunostaining assays (Figure 4B). We have generated a couple of antibodies against ctenophore synaptic proteins and are in the process of studying their intracellular localization. In addition, we are using many different biochemical methods, for example immunoprecipitation techniques, to isolate synaptic proteins from ctenophores to determine their in vivo binding partners (Figure 4B).

Working in the Burkhardt Lab has been a brilliant experience. Probably the most valuable thing I have learned from my time here has been how to properly manage my time in the lab. The MBA molecular lab corridor can be a very busy place. Different lab groups are constantly bustling about, and access to resources has to be negotiated amongst the various research groups. If I want to get everything finished by the end of the day then I have to be very efficient, which means I will usually have three or four things going on at any one time. This takes a lot of concentration, and by the end of the day I am absolutely shattered! My favourite thing about the MBA is that it’s right on the coast, so I like to finish the day with a swim to clear my head. My other favourite thing about the MBA is that there are loads of Italians working here. Tip of the day: in my experience Italian people equals food. After my swim I usually head to one of their houses for some home-made pasta, a glass of vino, and a chat. By the time I get home I am asleep before my head hits the pillow.
The bigger picture
What really attracted me to this project was one simple question. How could such a complex and specialised cell as a neuron possibly have evolved twice in animals? Until very recently, most of us took it for granted that synapses and neurons emerged only once during animal evolution. The two newly available ctenophore genomes have caused us to question our current understanding of the course of neuron evolution. It is exciting to know that the questions surrounding ctenophores and the origin of neurons are only a few years old. As such, these strange and beautiful creatures are likely to remain an important source of information, as we strive to unravel the mysteries of past evolutionary events. For me it will be rewarding to trace the evolutionary origin of synapses and neurons, and find out more about our own evolutionary history. Furthermore, I get to study one of the coolest animals I have ever seen. If this does not excite a biologist, then I don’t know what does!
References
Achim, K. & Arendt, D. Structural evolution of cell types by step-wise assembly of
cellular modules. Curr. Opin. Genet. Dev. 27, 102–108 (2014).
Brusca, R. C., Wendy, M. & Shuster, S. M. Invertebrates. Sinauer Associates, (2016).
Jager, M. & Manuel, M. Ctenophores: an evolutionary-developmental perspective
Curr. Opin. Genet. Dev. 39, 85–92 (2016).
Jekely, G., Paps, J. & Nielsen, C. The phylogenetic position of ctenophores and the
origin(s) of nervous systems. Evodevo 6:1 (2015).
Moroz, L. L. et al. The ctenophore genome and the evolutionary origins of neural
systems. Nature 510, 109–14 (2014).
Moroz, L. L. & Kohn, A. B. Unbiased View of Synaptic and Neuronal Gene
Complement in Ctenophores: Are There Pan-neuronal and Pan-synaptic Genes across
Metazoa? Integrative and Comparative Biology 55, 1028–1049 (2015).
Pang, K. & Martindale, M. Q. Comb jellies (Ctenophora): A model for basal
metazoan evolution and development. Cold Spring Harb. Protoc. 3, (2008).
Philippe, H. et al. Phylogenomics Revives Traditional Views on Deep Animal
Relationships. Curr. Biol. 19, 706–712 (2009).
Pisani, D. et al. Genomic data do not support comb jellies as the sister group to all
other animals. Proc. Natl. Acad. Sci. 112, 201518127 (2015).
Ryan, J. F. et al. The genome of the ctenophore Mnemiopsis leidyi and its
implications for cell type evolution. Science 342, 1336–1344 (2013).
Ryan, J. F. Did the ctenophore nervous system evolve independently? Zoology 117,
225–226 (2014).
Whelan, N. V, Kocot, K. M., Moroz, L. L. & Halanych, K. M. Error, signal, and the
placement of Ctenophora sister to all other animals. Proc. Natl. Acad. Sci. U. S. A.
112, 5773–8 (2015).


 (3 votes)
(3 votes)