A day in the life of a Spider Lab
Posted by Daniel_Leite, on 17 February 2016
We (Anna Schönauer, Daniel Leite and Christian Bonatto) are PhD students in Alistair McGregor’s group (http://mcgregor-evo-devo-lab.net) at Oxford Brookes University, and it is a pleasure to briefly present our research on spiders. The university is located up on Headington Hill, from where we can look out across the beautiful spires of the great academic city of Oxford. Our research focuses on animal development and evolution using the common house spider Parasteatoda tepidariorum (previously known as Achaearanea tepidariorum). Currently, the topics we are interested in include the regulation of segmentation of the opisthosoma (the posterior region of the spider body) and the evolution and function of microRNAs. Parasteatoda is now emerging as an excellent model to study these biological mechanisms and evolutionary processes.
The Parasteatoda Culture
Our spider culture was originally founded from individuals collected from a basement in Göttingen, Germany. We keep the bulk of our culture in a 25°C room with a multitude of other arthropods and even the departments pet corn snake (Figure 1). To maximise their health and productivity, we feed the spiders twice a week, on Monday and Friday, and the mated female spiders get an extra feed on Wednesday. Depending on their size, both the mated and unmated adult females are fed with crickets, while males and juvenile spiders are given flies.

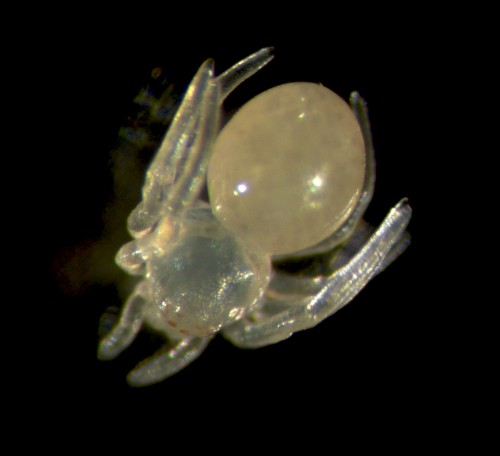
Handling the spiders is safe and easy because they don’t bite humans and in any case their chelicerae can’t penetrate our skin. Adults are small in size (~2 cm leg span) so they can be kept individually in small vials (otherwise they tend to eat each other!). While the females are fairly sedentary, the males are more active as they search for females or subsequently try to escape from them. Once males and females (Figure 2) have reached their final moults they are brought together to mate (Video). From then on we generally keep the male in the same vial as the female, to ensure successful mating but also for the females to have a little snack if she runs out of crickets! We always maintain around 50 mated females to provide enough embryos for experiments and ensure enough cocoons are available to sustain the culture.

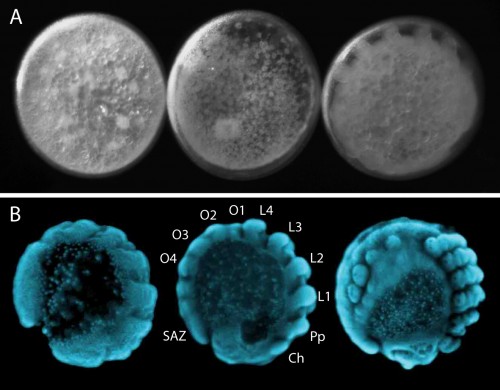
Our regular day starts with collecting cocoons from females, which they produce approximately every four days. There are usually hundreds of embryos per cocoon, all at similar developmental stages (Figure 3). Developing embryos can be kept in halocarbon oil, which makes the otherwise opaque chorion transparent and allows us to observe the developmental stages (Figure 4A). This helps us identify the exact stage of embryogenesis, which is key for the different approaches that we use to study their development. It takes approximately 10 days for the embryos to develop into translucent, hairless, immotile hatchling spiders (Figure 5). Once the juveniles have progressed through several molts and eaten some flies (as well as a few of their brothers and sisters), they are separated into individual vials. In total, after hatching, it takes about another month for Parasteatoda to reach reproductive maturity.


EvoDevo in the spider
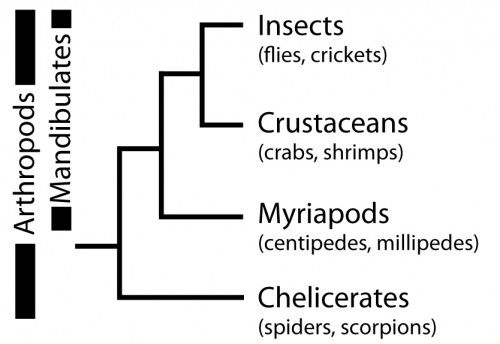
So, why are we using Parasteatoda to ask both developmental and evolutionary questions? Spiders, along with scorpions, mites, ticks and others, belong to the Chelicerata subphylum, which branches at the base of the Arthropoda (Figure 6). Arthropods are a hugely successful and diverse group of animals, however, many evolutionary and developmental studies have focused on insects such as Drosophila (flies), Tribolium (beetles) and Apis (bees). Therefore, investigating chelicerates species like Parasteatoda offers an important perspective to arthropod evolution and development, and to metazoans more broadly.
With the establishment of experimental tools, mainly thanks to pioneering work by Hiroki Oda and Yasuko Akiyama-Oda, Parasteatoda has become a powerful model chelicerate species for EvoDevo research. We can study gene expression in the developing spider with in situ hybridisation (ISH) and can knockdown expression with both parental and embryonic RNA interference (RNAi). A detailed description of embryonic development helps those who work with this spider to make standardised comparisons of developmental time points to interpret expression patterns and RNAi phenotypes.
In addition to the tools used to study gene expression and function during development in Parasteatoda, genomic resources have been developed, including a comprehensive embryonic transcriptome and genome (through the i5k initiative (http://www.arthropodgenomes.org/wiki/i5K). These are indispensable resources for quickly designing probes and RNAi constructs for our research on segmentation, and for mapping RNA sequence data for microRNA discovery.
Segmentation in Parasteatoda
The segmented body plan of arthropods is an important feature that has likely contributed to their evolutionary success and morphological diversity. However, despite conservation of segmentation genes, different mechanisms of segmentation are used among arthropods. In contrast to the somewhat simultaneous development of all segments in Drosophila, Parasteatoda employs a short germ mode of development, which means that some anterior segments are specified first and subsequently the posterior segments are sequentially added from the growth zone or segment addition zone (SAZ) (Figure 4B). In the spider, aspects of vertebrate somitogenesis (Delta/Notch, Wnt signaling), as well as components of the well-studied Drosophila segmentation gene cascade have been found to play a crucial role in posterior segmentation.
In order to functionally test interactions between those components in Parasteatoda, we utilise embryonic RNAi, where dsRNA is injected into a single blastomeres using a protocol developed in the Oda lab. For the injections, embryos have to be lined up one by one on double-sided sticky tape, which can be fairly fiddly and tricky. We use a fluorescent dye in the injection mix so that we can mark the clone of cells that is affected by the RNAi knockdown. After injections we have a peek down the microscope to see where the population of affected cells is located within the developing embryo, but we can also visualize the clone in fixed tissue by staining for Biotin.
In spiders, RNAi knockdown embryos can also be generated by injecting adult females with dsRNA, whereby the knockdown effect is transmitted to their offspring. The injection requires careful positioning of the needle to avoid damaging the heart or other vital organs. One injected female will produce a series of cocoons, with each cocoon exhibiting a different phenotypic severity. The fact that each cocoon consists of many embryos, all at roughly similar stages of development, is really useful for trying to capture this dynamic process of segment addition. When we knock down genes with parental or embryonic RNAi, the results show a disruption of the gene expression in the posterior and the truncation of embryonic tissue and help to decipher the role of particular genes during posterior development.
MicroRNAs in Parasteatoda
MicroRNAs have been shown to be important fine tuners of gene expression and are involved in many developmental process. Much of what we understand about them within invertebrates comes from studies in insects. These studies have found interesting patterns of microRNA evolution within particular lineages such as Drosophila. However, there is a lack of a broader understanding of microRNA evolution and function in development across arthropod species.
To date, the characterisation of microRNA repertoires in chelicerates has been limited to just mites and ticks. Our ongoing research aims to characterise the microRNAs present in Parasteatoda to provide new comparative insights into the evolution of these genes among chelicerate orders and other metazoans. This will allow us to then identify their functions and further investigate their involvement in the evolution of morphological diversity.
The rest of the Lab
Parasteatoda is not the only model organism being studied in the McGregor lab. Our colleagues are investigating the genetic and developmental bases of differences in morphology within and between Drosophila species. This includes studying the evolution and development of compound eyes, which vary in the number and size of ommatidia, and differ in the distribution of rhodopsin proteins. Members of our group also investigate the rapid evolution of male Drosophila genitalia, which always kindles some interesting conversations.
Our lab is very international and it is excellent to have a mixture of people and projects running so that we can learn about experimental approaches and biological processes in different organisms. While spiders and flies may not get along, the humans in the lab often end the evening altogether in one of the amazing pubs in Oxford, over a refreshing pint! If you want to know more about our projects, feel free to visit our website (http://mcgregor-evo-devo-lab.net) and if you have any questions, please email us!
 This post is part of a series on a day in the life of developmental biology labs working on different model organisms. You can read the introduction to the series here and read other posts in this series here.
This post is part of a series on a day in the life of developmental biology labs working on different model organisms. You can read the introduction to the series here and read other posts in this series here.
2 thoughts on “A day in the life of a Spider Lab”
-
Pingback: Morsels For The Mind – 19/02/2016 › Six Incredible Things Before Breakfast
-
Pingback: Spiderday (#25) – February | Arthropod Ecology


 (8 votes)
(8 votes)