In Development this week (Vol. 143, Issue 14)
Posted by Seema Grewal, on 19 July 2016
Here are the highlights from the current issue of Development:
Gestational stress: at the heart of birth defects

Congenital heart disease (CHD) is the most common form of human birth defect, yet the genetic and environmental factors that contribute to CHD remain poorly understood. Here, Sally Dunwoodie and colleagues investigate how gestational hypoxia affects heart development in mouse embryos (p. 2561). They reveal that the exposure of developing mouse embryos to short-term hypoxia in uteroresults in heart defects, notably perturbations to the outflow tract (OFT). These changes are mediated by altered cell proliferation and FGF signalling in the secondary heart field (SHF), which contains progenitor cells that contribute to the OFT. The authors further report that hypoxia leads to rapid induction of the unfolded protein response (UPR) in SHF cells. This, in turn, causes a global decrease in protein translation and may contribute to the reduced levels of FGFR1, and hence FGF signalling, observed in SHF cells following exposure to hypoxia. Together, these results suggest that hypoxia-mediated UPR induction during pregnancy can give rise to CHD. Given the key role of FGF signalling during embryogenesis, these findings also have important implications for understanding birth defects that affect other organs.
Shox2 goes out on a limb

Hox-TALE factors are involved in patterning the vertebrate limb but precisely how they regulate specific and regional gene expression patterns is unclear. Here, on p. 2548, YiPing Chen and co-workers uncover a limb patterning transcriptional programme that is coordinated by the transcription factor Shox2. The researchers demonstrate that, although Shox2 is expressed in mesenchymal progenitors of multiple cell types in the proximal limb, its deletion specifically in the osteogenic lineage causes limb defects and loss of the stylopod – the most proximal region of the limb. ChIP-Seq analyses indicate that Shox2 binds predominantly to limb-specific enhancers that are involved in skeletogenesis; these regions are also co-occupied by Hox-TALE factors. Finally, the authors show that Shox2 is expressed in a gradient that is complementary to that of TALE factors and that it represses the expression of TALE factors in the stylopod. Overall, these observations, together with other findings, highlight the existence of a Shox2-coordinated transcriptional programme that functions to pattern the vertebrate limb and provide insights into the ‘enhancer grammar’ that is used to mediate specific transcriptional outputs.
Pinning down spindle orientation

Spindle orientation is regulated by a conserved mechanism in which Pins/LGN anchors Mud/NuMA to the cell cortex. This pathway has been assumed to operate in nearly all animal cell types but here (p. 2573) Daniel St Johnston and colleagues reveal that Pins is not required for spindle orientation in the Drosophila wing disc. Using live imaging, they first discover that spindle angles in the wing disc vary widely; spindle angles are initially random but gradually align with the plane of the tissue as cells enter anaphase, highlighting that spindle angles are not accurate predictors of division orientation until this point. Importantly, the researchers reveal that spindle orientation does not require Pins, or aPKC, Dlg or Lgl. They further report that Mud is able to localize to the cell cortex in the absence of Pins, suggesting that a parallel but as yet unknown mechanism must act to localize Mud in wing disc cells. In summary, these surprising results indicate that a Pins-independent mechanism can orient the mitotic spindle in the Drosophila wing disc and lead the authors to propose that this system provides robustness to this rapidly developing epithelial tissue.
A sticky situation in plants

Cell-cell adhesion in plants is known to be regulated by pectins, and levels of homogalacturonan (HG; the main component of pectins) within the cell wall have generally been linked to cell adhesion. But how is cell-cell adhesion maintained and regulated in the face of the dynamic cell wall remodelling that takes place during cell growth and division? Here, Grégory Mouille and colleagues investigate this issue (p. 2536). Using a cell adhesion defect suppressor screen, they identify a putative O-fucosyltransferase – an enzyme that mediates the transfer of sugar residues onto substrates – that regulates cell adhesion in Arabidopsis thaliana. They further reveal that mutations in the gene encoding this enzyme or another putative O-fucosyltransferase perturb cell adhesion. Importantly, a comparison of mutant and suppressor lines suggests that cell adhesion does not rely on HG content per se. Based on their findings, the authors propose a model in which a pectin-related signalling pathway, rather than simply HG levels, contributes to the control and maintenance of cell adhesion during plant growth and development.
Plus:
An interview with Enrico Coen
 Enrico Coen CBE FRS is a Project Leader at the John Innes Centre in Norwich, UK, who uses a variety of approaches to study patterning and morphogenesis in plants. We met with Enrico at the Spring Meeting of the British Society for Developmental Biology, where he was awarded the Waddington Medal, to ask him more about his career and his passion for art and book-writing. See the Spotlight article on p. 2479.
Enrico Coen CBE FRS is a Project Leader at the John Innes Centre in Norwich, UK, who uses a variety of approaches to study patterning and morphogenesis in plants. We met with Enrico at the Spring Meeting of the British Society for Developmental Biology, where he was awarded the Waddington Medal, to ask him more about his career and his passion for art and book-writing. See the Spotlight article on p. 2479.
Exosomes in developmental signalling
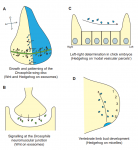 Cells can signal by releasing exosomes – extracellular vesicles containing bioactive molecules such as RNA, DNA and enzymes. Recent work has suggested that exosomes can also carry signalling proteins, including ligands of the Notch receptor and secreted proteins of the Hedgehog and WNT families. Here, describe various types of exosomes and their biogenesis and critically assess the role of exosomes in developmental signalling. See the Review on p. 2482.
Cells can signal by releasing exosomes – extracellular vesicles containing bioactive molecules such as RNA, DNA and enzymes. Recent work has suggested that exosomes can also carry signalling proteins, including ligands of the Notch receptor and secreted proteins of the Hedgehog and WNT families. Here, describe various types of exosomes and their biogenesis and critically assess the role of exosomes in developmental signalling. See the Review on p. 2482.
Direct neuronal reprogramming: learning from and for development
![]() The key signalling pathways and transcriptional programmes that instruct neuronal diversity during development have largely been identified. Here, and co-workers discuss how this knowledge has been used to successfully reprogramme various cell types into distinct types of functional neurons, and the extent to which direct neuronal reprogramming recapitulates embryonic development. See the Review on p. 2494.
The key signalling pathways and transcriptional programmes that instruct neuronal diversity during development have largely been identified. Here, and co-workers discuss how this knowledge has been used to successfully reprogramme various cell types into distinct types of functional neurons, and the extent to which direct neuronal reprogramming recapitulates embryonic development. See the Review on p. 2494.


 (No Ratings Yet)
(No Ratings Yet)