In Development this week (Vol. 140, Issue 2)
Posted by Seema Grewal, on 18 December 2012
Here are the highlights from the current issue of Development:
Against the segmentation clock
 A gene expression oscillator called the segmentation clock controls the periodic formation of somites in vertebrate embryos. In zebrafish, negative autoregulation of the transcriptional repressor genes her1 and her7 is thought to control the clock’s oscillations. Delays in this negative-feedback loop, including transcriptional delay (the time taken to make each her1 or her7 mRNA) should thus control the clock’s oscillation period. On p. 444, Ertuǧrul Özbudak, Julian Lewis and colleagues report that, unexpectedly, mutants in which only her1 or her7 is functional have almost identical segmentation clock oscillation periods – because the her1 and her7 genes are very different lengths, the researchers had anticipated that the two mutants would have different transcriptional delays and thus different oscillation periods. The researchers resolve this paradox by showing that the RNA polymerase II elongation rate is extremely fast in zebrafish embryos. They suggest, therefore, that the time taken for her1 and her7 transcript elongation is relatively insignificant, and that other sources of delay (e.g. splicing delay) may instead determine the oscillation period.
A gene expression oscillator called the segmentation clock controls the periodic formation of somites in vertebrate embryos. In zebrafish, negative autoregulation of the transcriptional repressor genes her1 and her7 is thought to control the clock’s oscillations. Delays in this negative-feedback loop, including transcriptional delay (the time taken to make each her1 or her7 mRNA) should thus control the clock’s oscillation period. On p. 444, Ertuǧrul Özbudak, Julian Lewis and colleagues report that, unexpectedly, mutants in which only her1 or her7 is functional have almost identical segmentation clock oscillation periods – because the her1 and her7 genes are very different lengths, the researchers had anticipated that the two mutants would have different transcriptional delays and thus different oscillation periods. The researchers resolve this paradox by showing that the RNA polymerase II elongation rate is extremely fast in zebrafish embryos. They suggest, therefore, that the time taken for her1 and her7 transcript elongation is relatively insignificant, and that other sources of delay (e.g. splicing delay) may instead determine the oscillation period.
Non-catalytic role for Mmp14 in stromal invasion
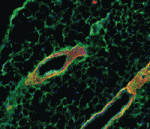 During mammary gland branching morphogenesis, mammary epithelial cells (MECs) invade the surrounding stroma. Here (p. 343), Mina Bissell and co-workers investigate whether the matrix metalloproteinase Mmp14, which is expressed in mammary glands during branching and which has a key role in cancer cell invasion, is involved in branching morphogenesis. The researchers report that the catalytic activity of Mmp14 is required for MEC branching in dense but not sparse three-dimensional collagen gels, but that, surprisingly, a non-proteolytic function of Mmp14 is required for branching in both conditions. They show that silencing Mmp14 reduces the levels of integrin β1 (Itgb1), which is required for branching in vivo, and that Mmp14 associates directly with Itgb1 through a transmembrane/cytoplasmic domain in Mmp14. Notably, this non-catalytic domain is required for branching in collagen gels. These results indicate that non-proteolytic activities of Mmp14 modulate the Itgb1-dependent signals that mediate MEC invasion during branching morphogenesis and provide a possible explanation for why cancer therapy drugs that target Mmp14’s catalytic function have failed in clinical trials.
During mammary gland branching morphogenesis, mammary epithelial cells (MECs) invade the surrounding stroma. Here (p. 343), Mina Bissell and co-workers investigate whether the matrix metalloproteinase Mmp14, which is expressed in mammary glands during branching and which has a key role in cancer cell invasion, is involved in branching morphogenesis. The researchers report that the catalytic activity of Mmp14 is required for MEC branching in dense but not sparse three-dimensional collagen gels, but that, surprisingly, a non-proteolytic function of Mmp14 is required for branching in both conditions. They show that silencing Mmp14 reduces the levels of integrin β1 (Itgb1), which is required for branching in vivo, and that Mmp14 associates directly with Itgb1 through a transmembrane/cytoplasmic domain in Mmp14. Notably, this non-catalytic domain is required for branching in collagen gels. These results indicate that non-proteolytic activities of Mmp14 modulate the Itgb1-dependent signals that mediate MEC invasion during branching morphogenesis and provide a possible explanation for why cancer therapy drugs that target Mmp14’s catalytic function have failed in clinical trials.
Bioelectric signals size up regeneration
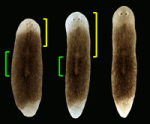 Regenerative medicine aims to replace lost or damaged tissues and organs with functional parts of the correct size and shape. To achieve this goal, we need to understand what determines the scale and form of regenerating tissues. Michael Levin and colleagues have been tackling this issue by investigating the regulation of organ size during planarian regeneration (see p. 313). During this process, existing tissues are remodelled concurrently with new tissue growth to maintain the correct relative tissue proportions. The researchers show that, in Schmidtea mediterranea, membrane voltage-dependent bioelectric signalling determines head size and organ scaling during regeneration. Specifically, RNA interference of the H+,K+-ATPase ion pump causes membrane hyperpolarisation and produces regenerated animals with shrunken heads and oversized pharynges, but does not inhibit the production of new tissue (blastema). Other experiments indicate that the H+,K+-ATPase ion pump maintains proportionality during regeneration by mediating apoptotic sculpting of the original tissues. Thus, bioelectric signalling regulates the cellular mechanisms that control organ size and shape during regeneration.
Regenerative medicine aims to replace lost or damaged tissues and organs with functional parts of the correct size and shape. To achieve this goal, we need to understand what determines the scale and form of regenerating tissues. Michael Levin and colleagues have been tackling this issue by investigating the regulation of organ size during planarian regeneration (see p. 313). During this process, existing tissues are remodelled concurrently with new tissue growth to maintain the correct relative tissue proportions. The researchers show that, in Schmidtea mediterranea, membrane voltage-dependent bioelectric signalling determines head size and organ scaling during regeneration. Specifically, RNA interference of the H+,K+-ATPase ion pump causes membrane hyperpolarisation and produces regenerated animals with shrunken heads and oversized pharynges, but does not inhibit the production of new tissue (blastema). Other experiments indicate that the H+,K+-ATPase ion pump maintains proportionality during regeneration by mediating apoptotic sculpting of the original tissues. Thus, bioelectric signalling regulates the cellular mechanisms that control organ size and shape during regeneration.
Fat-tening up planar cell polarity
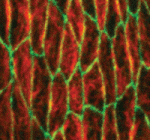 The conserved atypical cadherin Fat regulates planar cell polarity, but the mechanisms by which Fat controls cell shape and tissue organisation are unclear. Emily Marcinkevicius and Jennifer Zallen (p. 433) now show that Fat is required for the planar polarised organisation of denticle precursors, adherens junction proteins and microtubules in the Drosophila embryo epidermis. Adherens junction remodelling and cell shape are disrupted in fat mutants, they report, and in flies carrying mutations in Expanded (a regulator of the Hippo pathway) and in Hippo and Warts (two kinases in the Hippo pathway). Mutations in the Hippo/Warts pathway do not recapitulate the effects of Fat loss on denticle planar organisation, however, and the cell shape and planar polarity defects in fat mutants do not require transcriptional regulation by Yorkie, a target of the Hippo pathway. These results suggest that a common upstream signal provided by Fat regulates junctional and cytoskeletal planar polarity in the Drosophila embryo and that Fat influences tissue organisation by regulating polarised junctional remodelling.
The conserved atypical cadherin Fat regulates planar cell polarity, but the mechanisms by which Fat controls cell shape and tissue organisation are unclear. Emily Marcinkevicius and Jennifer Zallen (p. 433) now show that Fat is required for the planar polarised organisation of denticle precursors, adherens junction proteins and microtubules in the Drosophila embryo epidermis. Adherens junction remodelling and cell shape are disrupted in fat mutants, they report, and in flies carrying mutations in Expanded (a regulator of the Hippo pathway) and in Hippo and Warts (two kinases in the Hippo pathway). Mutations in the Hippo/Warts pathway do not recapitulate the effects of Fat loss on denticle planar organisation, however, and the cell shape and planar polarity defects in fat mutants do not require transcriptional regulation by Yorkie, a target of the Hippo pathway. These results suggest that a common upstream signal provided by Fat regulates junctional and cytoskeletal planar polarity in the Drosophila embryo and that Fat influences tissue organisation by regulating polarised junctional remodelling.
Recapitulating striatal neurogenesis
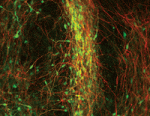 Medium-sized spiny neurons (MSNs) are the only neostriatum projection neurons and are among the first neurons to degenerate in Huntington’s disease. Here (see p. 301), Elena Cattaneo and co-workers report that human pluripotent stem (hPS) cells can be induced to differentiate into MSNs using an ontogeny-recapitulating protocol. The researchers induce ventral telencephalic specification of hPS cells in feeder-free adherent cultures using BMP/TGFβ inhibition followed by sonic hedgehog/WNT pathway modulation. They then induce terminal differentiation of the telencephalic progenitors in the presence of brain-derived neurotrophic factor, thereby generating MSNs that express DARPP-32 and other striatal markers. These MSNs carry dopamine and adenosine receptors, elicit a typical firing pattern, and show dopamine-dependent neuromodulation and synaptic integration ability in vivo. Finally, when transplanted into the striatum of a rat model for Huntington’s disease, hPS cell-derived neurons survive and correct motor deficits. This ontogeny-recapitulating method provides a platform for human neurodevelopmental biology studies, suggest the researchers, and could be used to develop a Huntington’s disease model for drug screening.
Medium-sized spiny neurons (MSNs) are the only neostriatum projection neurons and are among the first neurons to degenerate in Huntington’s disease. Here (see p. 301), Elena Cattaneo and co-workers report that human pluripotent stem (hPS) cells can be induced to differentiate into MSNs using an ontogeny-recapitulating protocol. The researchers induce ventral telencephalic specification of hPS cells in feeder-free adherent cultures using BMP/TGFβ inhibition followed by sonic hedgehog/WNT pathway modulation. They then induce terminal differentiation of the telencephalic progenitors in the presence of brain-derived neurotrophic factor, thereby generating MSNs that express DARPP-32 and other striatal markers. These MSNs carry dopamine and adenosine receptors, elicit a typical firing pattern, and show dopamine-dependent neuromodulation and synaptic integration ability in vivo. Finally, when transplanted into the striatum of a rat model for Huntington’s disease, hPS cell-derived neurons survive and correct motor deficits. This ontogeny-recapitulating method provides a platform for human neurodevelopmental biology studies, suggest the researchers, and could be used to develop a Huntington’s disease model for drug screening.
Cell-cycle pattern predicts stem cell fate
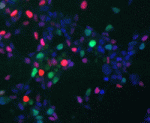 Cell-cycle progression and lineage commitment of stem cells seem to be tightly linked but it has not been possible to study this relationship in live stem cells. Now, Matthias Lutolf and colleagues (p. 459) describe a single-cell tracking approach that enables the automatic detection of cell-cycle phases in live stem cells that express fluorescent ubiquitylation-based cell-cycle indicator (FUCCI) probes. The researchers use their approach to identify distinctive changes in the length of cell-cycle phases and in the fluorescence intensity of the G1 (red) and S/G2-M (green) FUCCI probes during the differentiation of adult neural stem/progenitor cells (NSCs) and embryonic stem cells. Moreover, they use these changes in fluorescence intensity to purify NSCs from a heterogeneous population and to increase the proportion of reprogrammed cells obtained during NSC reprogramming to an induced pluripotent stem cell-like state. These findings shed new light on the relationship between cell-cycle progression and cell fate choice and introduce a tool that could advance our understanding of stem cell biology.
Cell-cycle progression and lineage commitment of stem cells seem to be tightly linked but it has not been possible to study this relationship in live stem cells. Now, Matthias Lutolf and colleagues (p. 459) describe a single-cell tracking approach that enables the automatic detection of cell-cycle phases in live stem cells that express fluorescent ubiquitylation-based cell-cycle indicator (FUCCI) probes. The researchers use their approach to identify distinctive changes in the length of cell-cycle phases and in the fluorescence intensity of the G1 (red) and S/G2-M (green) FUCCI probes during the differentiation of adult neural stem/progenitor cells (NSCs) and embryonic stem cells. Moreover, they use these changes in fluorescence intensity to purify NSCs from a heterogeneous population and to increase the proportion of reprogrammed cells obtained during NSC reprogramming to an induced pluripotent stem cell-like state. These findings shed new light on the relationship between cell-cycle progression and cell fate choice and introduce a tool that could advance our understanding of stem cell biology.
Plus…
DEVELOPMENT AT A GLANCE: Phyllotaxis
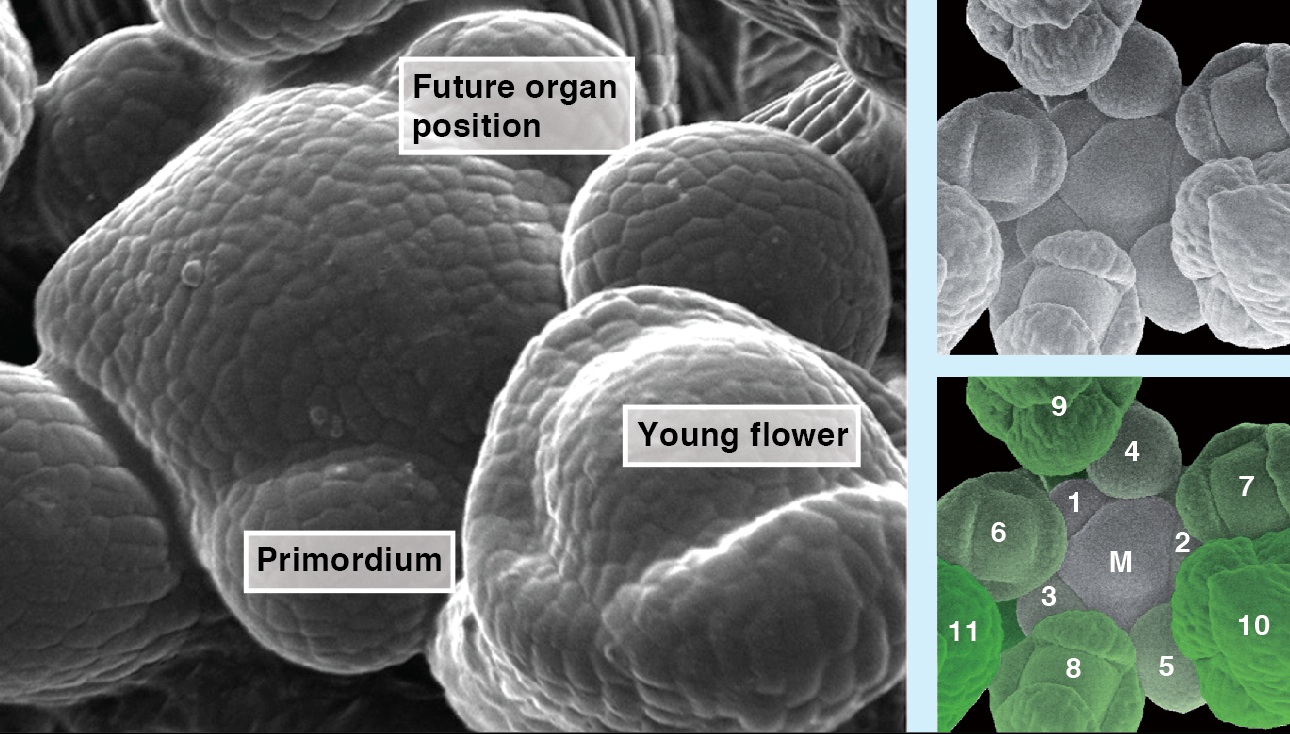 The precise arrangement of plant organs, also called phyllotaxis, has fascinated scientists from multiple disciplines. In this Issue, Jan Traas reviews new insights into the regulation of phyllotactic patterning and provides an overview of the various factors that can drive these robust growth patterns. See the Development at a Glance poster article on p. 249
The precise arrangement of plant organs, also called phyllotaxis, has fascinated scientists from multiple disciplines. In this Issue, Jan Traas reviews new insights into the regulation of phyllotactic patterning and provides an overview of the various factors that can drive these robust growth patterns. See the Development at a Glance poster article on p. 249
Adhesion in the stem cell niche: biological roles and regulation
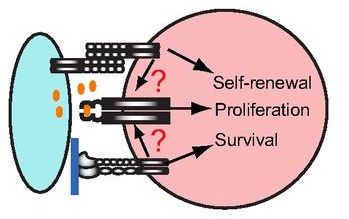 Stem cells are often anchored to their niche via adhesion molecules. However, as reviewed here by Xie and colleagues, recent studies have revealed other important roles for adhesion molecules in the regulation of stem cell function. See the Review article on p. 255
Stem cells are often anchored to their niche via adhesion molecules. However, as reviewed here by Xie and colleagues, recent studies have revealed other important roles for adhesion molecules in the regulation of stem cell function. See the Review article on p. 255


 (No Ratings Yet)
(No Ratings Yet)