In Development this week (Vol. 142, Issue 5)
Posted by Seema Grewal, on 24 February 2015
Here are the highlights from the current issue of Development:
pancRNAs in early development
 Promoter-associated noncoding RNAs (pancRNAs) are a class of long noncoding RNAs, transcribed from bidirectional promoters and thought to be involved in promoting expression of the gene transcribed from the opposite strand. Takuya Imamura and colleagues (p.910) now set out to investigate the prevalence and potential functions of such pancRNAs during early mouse development. Focussing on the 2-cell stage, when the major wave of zygotic gene activation (ZGA) occurs, they use directional RNA-seq technology to identify several hundred pancRNAs upregulated in concert with their cognate coding RNA. To assess the potential functional relevance of this co-regulation, the authors analyse three candidates, including Il17d/pancIl17d. In each case, siRNA-mediated knockdown of the pancRNA impairs expression of the mRNA and also prevents the normal DNA demethylation associated with gene upregulation at the ZGA. They further find that depletion of pancIl17d leads to defects in blastocyst survival and in embryonic stem cell colony formation – phenotypes that can be rescued by the provision of recombinant IL17D. Although the mechanisms by which pancRNAs act remain poorly understood, these data provide evidence for an important physiological role for pancRNAs in promoting expression of their partner mRNAs during early development.
Promoter-associated noncoding RNAs (pancRNAs) are a class of long noncoding RNAs, transcribed from bidirectional promoters and thought to be involved in promoting expression of the gene transcribed from the opposite strand. Takuya Imamura and colleagues (p.910) now set out to investigate the prevalence and potential functions of such pancRNAs during early mouse development. Focussing on the 2-cell stage, when the major wave of zygotic gene activation (ZGA) occurs, they use directional RNA-seq technology to identify several hundred pancRNAs upregulated in concert with their cognate coding RNA. To assess the potential functional relevance of this co-regulation, the authors analyse three candidates, including Il17d/pancIl17d. In each case, siRNA-mediated knockdown of the pancRNA impairs expression of the mRNA and also prevents the normal DNA demethylation associated with gene upregulation at the ZGA. They further find that depletion of pancIl17d leads to defects in blastocyst survival and in embryonic stem cell colony formation – phenotypes that can be rescued by the provision of recombinant IL17D. Although the mechanisms by which pancRNAs act remain poorly understood, these data provide evidence for an important physiological role for pancRNAs in promoting expression of their partner mRNAs during early development.Uncovering neurogenic potential of striatal astrocytes
 Adult neurogenesis in mammals is generally confined to specific regions where astrocytic cells produce particular neuronal types throughout life. Outside these areas, the capacity for neurogenesis is limited. However, on p.840, Paolo Peretto, Federico Luzzati and co-workers provide evidence that striatal astrocytes can be activated to a neurogenic program in an adult mouse model of Huntington’s disease. Following treatment with the toxin quinolinic acid to induce a lesion, the authors observe the appearance of proliferating progenitors and neuroblasts. Fate-mapping experiments identify local striatal astrocytes as the source of this neurogenic program. Importantly, neurogenesis is only observed upon lesion, suggesting that these astrocytes are normally quiescent but possess latent neurogenic potential upon damage. Further analysis is required to understand the programs regulating this neurogenesis, to determine the final fate of the newly born neuroblasts and to assess whether a similar phenomenon might exist in humans. However, these data open the possibility of harnessing the neurogenic potential of striatal astrocytes for therapeutic purposes.
Adult neurogenesis in mammals is generally confined to specific regions where astrocytic cells produce particular neuronal types throughout life. Outside these areas, the capacity for neurogenesis is limited. However, on p.840, Paolo Peretto, Federico Luzzati and co-workers provide evidence that striatal astrocytes can be activated to a neurogenic program in an adult mouse model of Huntington’s disease. Following treatment with the toxin quinolinic acid to induce a lesion, the authors observe the appearance of proliferating progenitors and neuroblasts. Fate-mapping experiments identify local striatal astrocytes as the source of this neurogenic program. Importantly, neurogenesis is only observed upon lesion, suggesting that these astrocytes are normally quiescent but possess latent neurogenic potential upon damage. Further analysis is required to understand the programs regulating this neurogenesis, to determine the final fate of the newly born neuroblasts and to assess whether a similar phenomenon might exist in humans. However, these data open the possibility of harnessing the neurogenic potential of striatal astrocytes for therapeutic purposes.Timing degradation to tune development
 A key issue in developmental biology is how particular processes are coordinated in time: for example, what determines when development of a particular organ starts and how quickly it proceeds? Jennifer Nemhauser and colleagues explore this problem in Arabidopsis, using auxin-mediated regulation of lateral root development as a model (p. 905). Specifically, they investigate the consequence of manipulating the degradation rate of the auxin-responsive transcriptional co-repressor IAA14, the mutation of which is known to affect lateral root formation. Using a synthetic biology approach, the authors generate several versions of IAA14 that show varying degradation kinetics upon auxin stimulation, and then generate transgenic plants expressing each version from the wild-type promoter. They find that both the density and timing of lateral root emergence inversely correlates with IAA14 stability: more stable variants show fewer lateral roots and these roots take longer to initiate. IAA14 is part of a large family of Aux/IAA repressors and these data suggest that regulating the degradation rate of these proteins could act as a tunable timer for developmental progression in plants.
A key issue in developmental biology is how particular processes are coordinated in time: for example, what determines when development of a particular organ starts and how quickly it proceeds? Jennifer Nemhauser and colleagues explore this problem in Arabidopsis, using auxin-mediated regulation of lateral root development as a model (p. 905). Specifically, they investigate the consequence of manipulating the degradation rate of the auxin-responsive transcriptional co-repressor IAA14, the mutation of which is known to affect lateral root formation. Using a synthetic biology approach, the authors generate several versions of IAA14 that show varying degradation kinetics upon auxin stimulation, and then generate transgenic plants expressing each version from the wild-type promoter. They find that both the density and timing of lateral root emergence inversely correlates with IAA14 stability: more stable variants show fewer lateral roots and these roots take longer to initiate. IAA14 is part of a large family of Aux/IAA repressors and these data suggest that regulating the degradation rate of these proteins could act as a tunable timer for developmental progression in plants.Getting to the heart of heart cell identity
 Cardiac progenitor cells differentiate into multiple cell types that make up the functional heart: cardiomyocytes (CMs), smooth muscle cells (SMCs), endothelial cells (EDCs) and fibroblasts. These lineage decisions can be modelled by differentiation of embryonic stem cells (ESCs), but it is not fully clear how closely these in vitrosystems reflect in vivo developmental progression, or how much variability there is within the progenitor population – either in culture or in the embryo. On p.846, Sean Wu and co-workers use single-cell quantitative PCR and lineage-tracing assays on embryonic and adult mouse cardiac cells, as well as mouse ESCs differentiated down the cardiac lineage, to define a gene expression signature for each of the various cell types. Amongst the wealth of data generated, a number of key findings emerge. Firstly, the authors find that ESC-derived CMs closely resemble embryonic and neonatal endogenous CMs, but adult CMs diverge. Secondly, embryonic and ESC-derived cardiac progenitors show different potential: both generate CMs, but embryonic cells can differentiate to EDCs while ESC-derived progenitors produce SMCs. These data demonstrate the power of the single-cell profiling approach and provide valuable insights into lineage choices during cardiac development.
Cardiac progenitor cells differentiate into multiple cell types that make up the functional heart: cardiomyocytes (CMs), smooth muscle cells (SMCs), endothelial cells (EDCs) and fibroblasts. These lineage decisions can be modelled by differentiation of embryonic stem cells (ESCs), but it is not fully clear how closely these in vitrosystems reflect in vivo developmental progression, or how much variability there is within the progenitor population – either in culture or in the embryo. On p.846, Sean Wu and co-workers use single-cell quantitative PCR and lineage-tracing assays on embryonic and adult mouse cardiac cells, as well as mouse ESCs differentiated down the cardiac lineage, to define a gene expression signature for each of the various cell types. Amongst the wealth of data generated, a number of key findings emerge. Firstly, the authors find that ESC-derived CMs closely resemble embryonic and neonatal endogenous CMs, but adult CMs diverge. Secondly, embryonic and ESC-derived cardiac progenitors show different potential: both generate CMs, but embryonic cells can differentiate to EDCs while ESC-derived progenitors produce SMCs. These data demonstrate the power of the single-cell profiling approach and provide valuable insights into lineage choices during cardiac development.Asymmetric division and fate in the retina
 In many systems, neural progenitor cells divide asymmetrically to generate a self-renewing progenitor and a committed neuron. How is this fate segregation controlled, and what defines the balance of proliferation and differentiation? Using live imaging in the developing zebrafish retina, Lucia Poggi and colleagues (p. 832) address these questions, focussing on the role of Anillin – a protein involved in cytokinesis – in the cell divisions that generate retinal ganglion cells (RGCs). By following individual divisions, the authors find that Anillin is itself asymmetrically inherited between daughter cells, and directs the asymmetric inheritance of actin and the polarity protein Par3. Cells with reduced Anillin levels tend to divide symmetrically, generating two RGCs rather than a progenitor and a RGC. Globally, this results in a retina with more RGCs. The authors further show that anillin expression is itself regulated by the RGC fate determinant Ath5, suggesting that there may be feedback loops involving Ath5 and Anillin that control the balance of proliferation and differentiation. How Anillin acts to regulate asymmetric division and fate choice remains unclear, but this technically challenging study demonstrates the importance of this protein in the control of neurogenesis in the retina.
In many systems, neural progenitor cells divide asymmetrically to generate a self-renewing progenitor and a committed neuron. How is this fate segregation controlled, and what defines the balance of proliferation and differentiation? Using live imaging in the developing zebrafish retina, Lucia Poggi and colleagues (p. 832) address these questions, focussing on the role of Anillin – a protein involved in cytokinesis – in the cell divisions that generate retinal ganglion cells (RGCs). By following individual divisions, the authors find that Anillin is itself asymmetrically inherited between daughter cells, and directs the asymmetric inheritance of actin and the polarity protein Par3. Cells with reduced Anillin levels tend to divide symmetrically, generating two RGCs rather than a progenitor and a RGC. Globally, this results in a retina with more RGCs. The authors further show that anillin expression is itself regulated by the RGC fate determinant Ath5, suggesting that there may be feedback loops involving Ath5 and Anillin that control the balance of proliferation and differentiation. How Anillin acts to regulate asymmetric division and fate choice remains unclear, but this technically challenging study demonstrates the importance of this protein in the control of neurogenesis in the retina.PLUS…
Neural development and regeneration: it’s all in your spinal cord
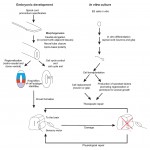 The latest advances in the field of spinal cord development and regeneration were discussed at a recent EMBO workshop entitled ‘Spinal cord development and regeneration’, which was held in Sitges, Spain, in October, 2014. Here, and Ruth Diez del Corral provide a review of the workshop. See the Meeting Review on p. 811
The latest advances in the field of spinal cord development and regeneration were discussed at a recent EMBO workshop entitled ‘Spinal cord development and regeneration’, which was held in Sitges, Spain, in October, 2014. Here, and Ruth Diez del Corral provide a review of the workshop. See the Meeting Review on p. 811
A pathway to bone: signaling molecules and transcription factors involved in chondrocyte development and maturation
 Decades of work have identified the signaling pathways that regulate the differentiation of chondrocytes during bone formation, from their initial induction from mesenchymal progenitor cells to their terminal maturation into hypertrophic chondrocytes. Here, , Elazar Zelzer review how multiple signaling molecules, mechanical signals and morphological cell features are integrated to activate a set of key transcription factors that determine and regulate the genetic program that induces chondrogenesis and chondrocyte differentiation. See the Review on p. 817
Decades of work have identified the signaling pathways that regulate the differentiation of chondrocytes during bone formation, from their initial induction from mesenchymal progenitor cells to their terminal maturation into hypertrophic chondrocytes. Here, , Elazar Zelzer review how multiple signaling molecules, mechanical signals and morphological cell features are integrated to activate a set of key transcription factors that determine and regulate the genetic program that induces chondrogenesis and chondrocyte differentiation. See the Review on p. 817
Beautiful imaging of zebrafish development
This issue’s featured movie shows the development of a zebrafish, from early embryo to larval stage, imaged using a combination of optical tomography and SPIM. Read the Techniques and Resources article by Huisken and colleagues here.


 (No Ratings Yet)
(No Ratings Yet)