Ph.D. Position in Computer Science and Developmental Biology
Posted by Léo Guignard, on 16 January 2021
Closing Date: 15 March 2021
We are seeking a highly motivated candidate strongly interested in interdisciplinary science, to join our groups (van de Pavert and Guignard labs) and work on a project at the crossroad between Computer Science and Developmental Biology as a Ph.D student.
Deadline: February 26 2021
Quantification and modelling of embryonic lymph node organogenesis at the single cell scale.
The role
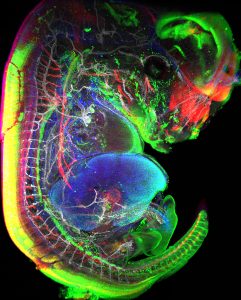
The project aims at studying lymph node (LN) formation during mouse embryonic development. LN formation requires hematopoietic lymphoid tissue inducer cells (LTi) to interact with mesenchymal cells at precise locations within the embryo, where they subsequently form aggregates. We have postulated that the peripheral nervous system outgrowth initiates the earliest events in LN formation. Indeed, preliminary data show that LTi aggregates morphology and cell density is affected in mouse embryos lacking neuronal subsets.
To understand the relationship between neuronal outgrowth and lymph node formation, the successful candidate will work between the van de Pavert lab and the Guignard lab to develop new computational methods to reconstruct and quantify LTi aggregates and peripheral nervous system morphology at the single cell scale. The reconstructions will then be used to develop a machine learning framework to systematically quantify phenotypes in perturbed mouse embryos. These quantifications will in turn allow to model the effects of neuronal outgrowth on LN formation.
The successful candidate will work towards:
- building a library of whole-mount mouse embryo images
- developing computational methods for the analysis of the generated library, including:
- reconstruction and mapping of the neuronal network
- quantification of LTi aggregate morphologies and positions
- developing computational methods to automatically stage mouse embryos
- modelling LN formation in relation to neuronal network morphology
Keywords
Quantitative embryogenesis, whole-mount analysis, peripheral nervous system, immune system, image analysis, machine learning, big data analysis
Whom would we like to hire?
The student should be enthusiastic, creative and ambitious, have good communication skills and be eager to learn. A master degree with major or minor in computer science is required. Affection for developmental biology is preferred. Some experience in developmental biology is also preferred but not required. Note: exception can be made for students who have not studied computer science if the student can prove coding skills.
The offer
- 3-year Ph.D. position in the van de Pavert lab and the Guignard lab
- Target start date: from October 2021
Application procedure
All applications must be done through the CENTURI portal at this address: https://centuri-livingsystems.org/application-form-phd2021-26/
Questions can be addressed to Serge van de Pavert or to Léo Guignard by email with the mention [Job-2021] in the title.
Serge van de Pavert: vandepavert@ciml.univ-mrs.fr
Léo Guignard: leo.guignard+lab@gmail.com
Selection process and calendar
- Call open for applications: January 15 – February 26
- Interviews of shortlisted candidates by the evaluation committee: April 20 to April 22
- The pre-selection process will be based on qualifications and expertise reflected on the candidates CV and motivation letter. It will be merit-based. All candidates will be informed whether they have been pre-selected or not.
- Final candidate selection: April 27
- Expected decision from the candidate: May 4
Relevant publications
- Simic A., et al., Distinct Waves from the Hemogenic Endothelium Give Rise to Layered Lymphoid Tissue Inducer Cell Ontogeny, Cell Reports, 32(6):108004, (2020)
- van de Pavert, et al., Chemokine CXCL13 is essential for lymph node initiation and is induced by retinoic acid and neuronal stimulation, Nature Immunology, 10(11):1193-9, (2009)
- van de Pavert, et al., Maternal retinoids control type 3 innate lymphoid cells and set the offspring immunity, Nature, 508(7494):123-7, (2014)
- McDole K.†, Guignard L.†, et al., In Toto Imaging and Reconstruction of Post-Implantation Mouse Development at the Single-Cell Level, Cell, 175(3):859-876, (2018)
About the teams
Guignard lab:
We are a group of computer scientists with a strong interested in biology in general and more specifically in embryonic development. We develop novel computational methods and models that allow the analysis of very large 3D movies of animal embryonic development (up to 2TB per movie). We work in close relationship with biologists to tailor our methods so that they help to address fundamental biological questions.
The developmental biology question that mainly but not only animates us is to better understand the mechanisms driving embryogenesis to robustly form a complex organism despite genetic polymorphism and variable environmental conditions.
For further information about the lab you can visit our website guignardlab.com, our twitter @guignardlab or contact directly Léo Guignard via email: leo.guignard+lab@gmail.com
van de Pavert lab:
We are interested to study the first cues required to form lymph nodes at specific locations within the embryo. We are fascinated by the interactions between different cell types, such as hematopoietic cells, mesenchymal cells and neurons which eventually will generate a highly organized LN. To study these interactions, we use a multi-disciplinary approach and combine techniques such as 3D immunofluorescence imaging, flow cytometry and single cell sequencing on embryos from different mouse models.
For further information, you can visit the lab’s website www.ciml.univ-mrs.fr/science/lab-serge-van-de-pavert, obtain an impression of the lab on Instagram @splab_ciml or contact Serge van de Pavert directly via email: vandepavert@ciml.univ-mrs.fr
The Institute
The Turing Centre for Living Systems (CENTURI) is an interdisciplinary project located in Marseille (France).
CENTURI aims at developing an integrated interdisciplinary community, to decipher the complexity of biological systems through the understanding of how biological function emerges from the organization and dynamics of living systems.
The project federates 15 teaching and research institutes in biology, physics, mathematics, computer science, engineering and focuses on Research, Education and Engineering, 3 missions that hold interdisciplinary as their core principle.
The research and training programmes implemented under the auspices of CENTURI will foster new collaborations, will transform practices, will attract new talents and thereby contribute to making the Luminy campus a leading site for the interdisciplinary study of biological systems.


 (No Ratings Yet)
(No Ratings Yet)