The 12 GIFs of Christmas
Posted by the Node, on 21 December 2018
Over on Twitter we’ve been having fun with our third instalment of the 12 GIFs of Christmas. For those not on Twitter, here are the GIFs – they represent some of the most cutting edge and inventive developmental biology of 2018, and also showcase the beauty of timelapse microscopy.
Transcription overlaid onto the rapid cell divisions of the early Drosophila embryo.
Jeremy Dufourt, Antonio Trullo, Jennifer Hunter, Carola Fernandez, Jorge Lazaro, Matthieu Dejean, Lucas Morales, Saida Nait-Amer, Katharine N. Schulz, Melissa M. Harrison, Cyril Favard, Ovidiu Radulescu & Mounia Lagha
Nature Communications
Arabidopsis under lightsheet
Multiscale imaging of plant development by light-sheet fluorescence microscopy
Miroslav Ovečka, Daniel von Wangenheim, Pavel Tomančák, Olga Šamajová, George Komis & Jozef Šamaj
Nature Plants
Mouse development like you’ve never seen it before (including PGC migration)
In Toto Imaging and Reconstruction of Post-Implantation Mouse Development at the Single-Cell Level
Katie McDole, Léo Guignard, Fernando Amat, Andrew Berger, Grégoire Malandain, Loïc A. Royer, Srinivas C. Turaga, Kristin Branson, Philipp J. Keller
Cell
Amphipod embryogenesis
Carsten Wolff, Jean-Yves Tinevez, Tobias Pietzsch, Evangelia Stamataki, Benjamin Harich, Léo Guignard, Stephan Preibisch, Spencer Shorte, Philipp J Keller, Pavel Tomancak, Anastasios Pavlopoulos
eLife
The fly adult midgut over many hours
Judy Lisette Martin, Erin Nicole Sanders, Paola Moreno-Roman, Leslie Ann Jaramillo Koyama, Shruthi Balachandra, XinXin Du, Lucy Erin O’Brien
eLife
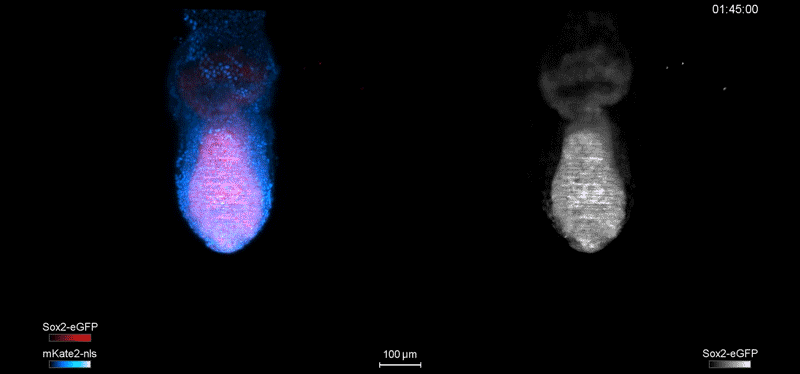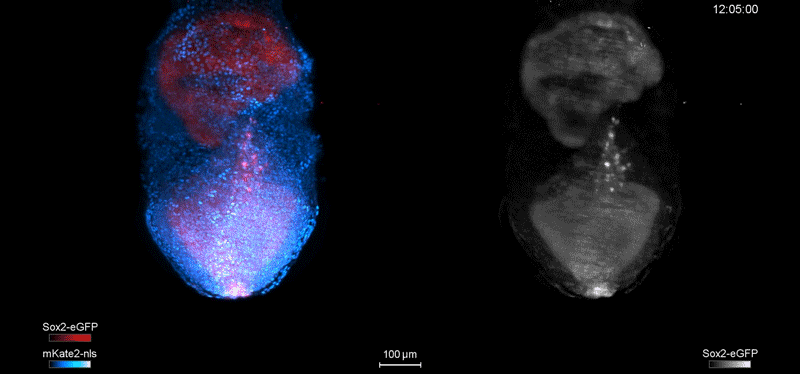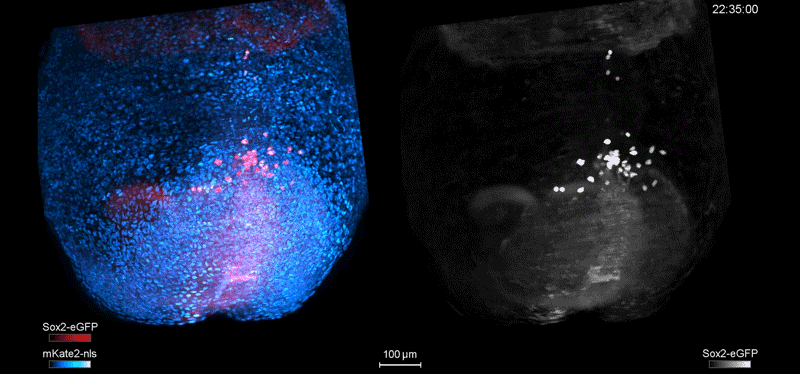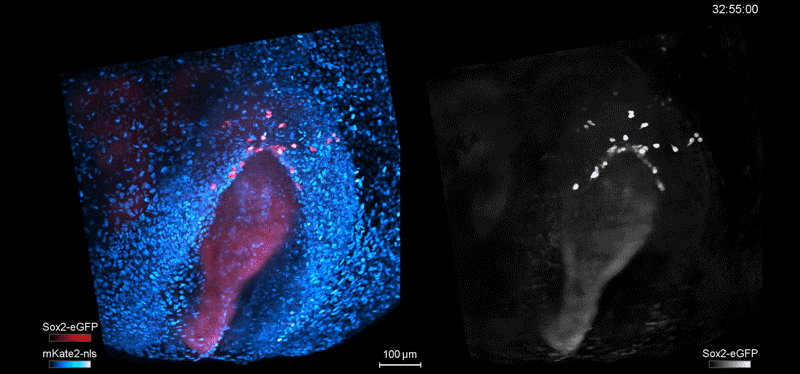
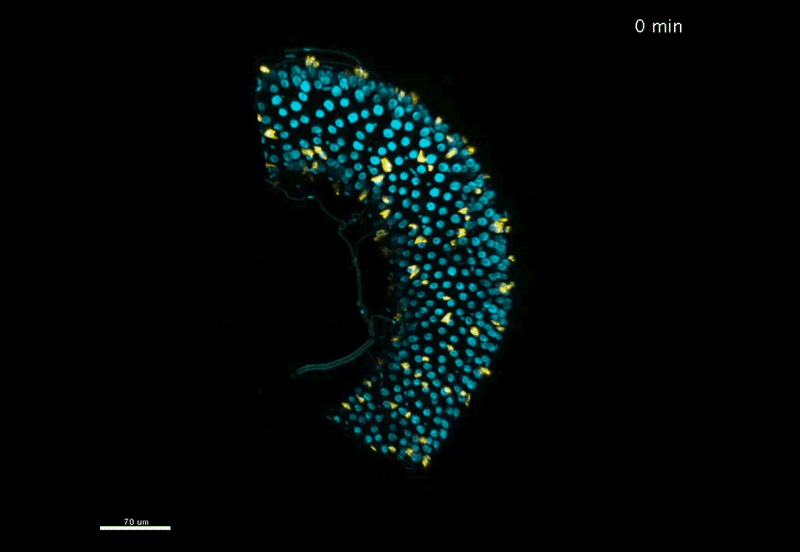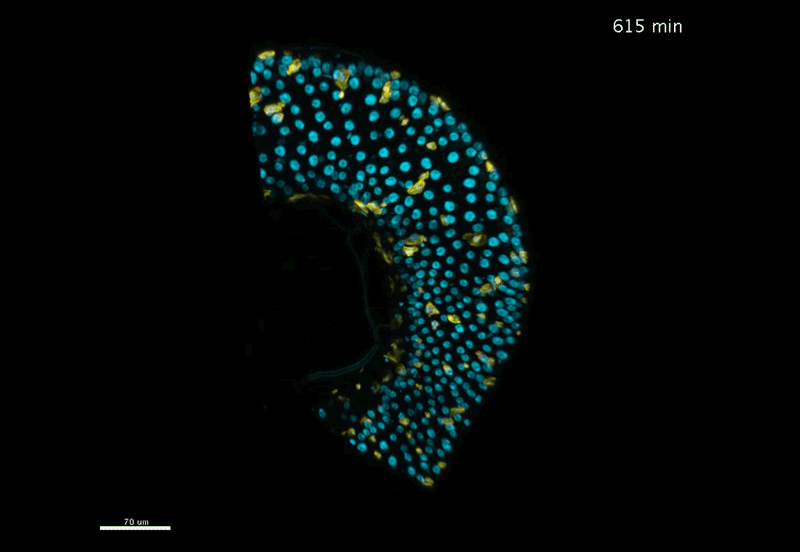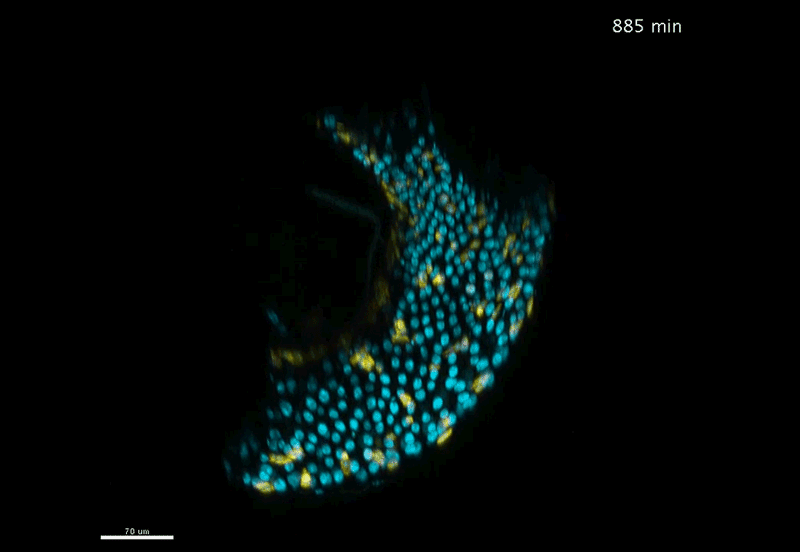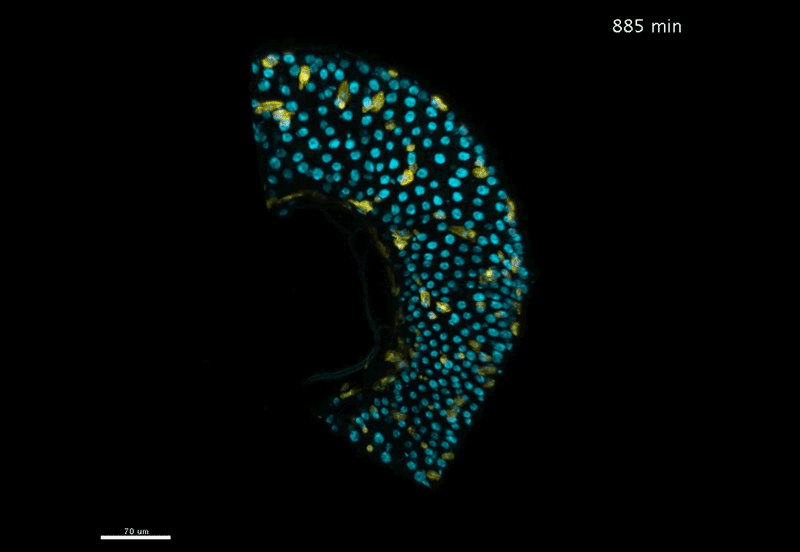
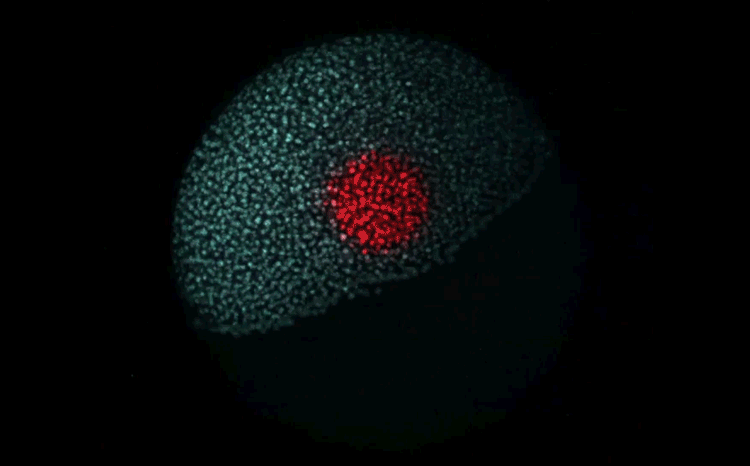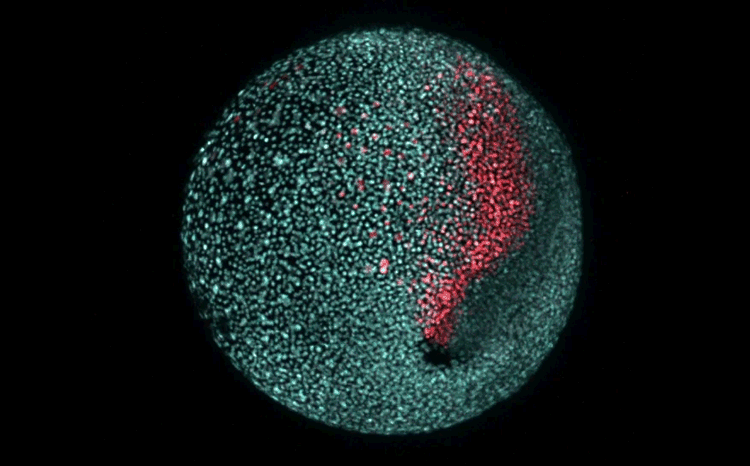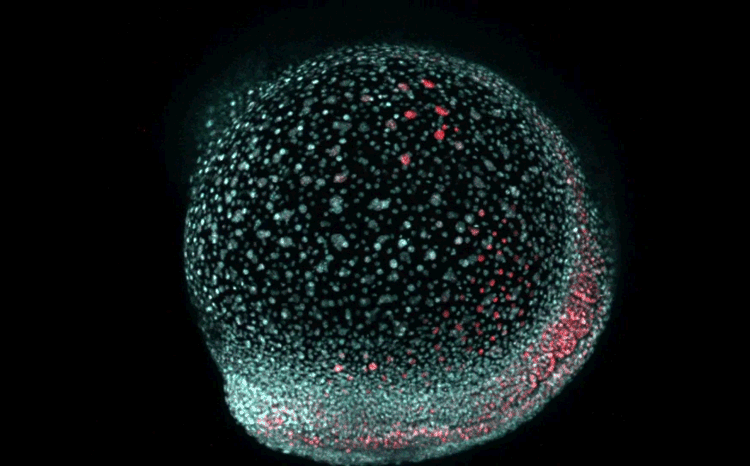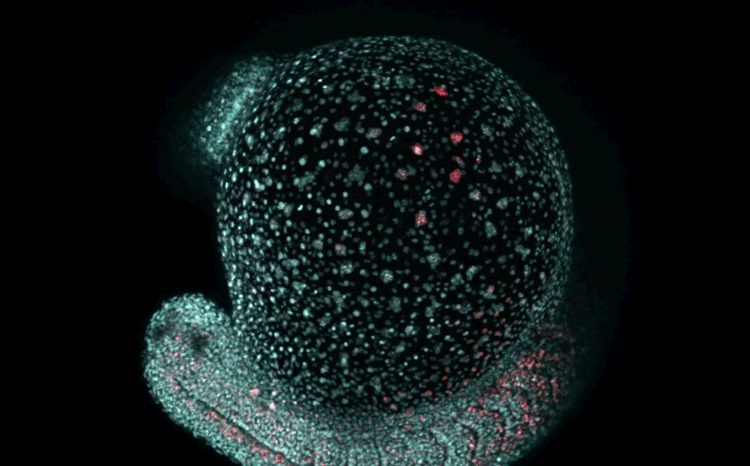
Fate mapping the zebrafish embryo with the photoconvertible protein Kikume
Neuromesodermal progenitors are a conserved source of spinal cord with divergent growth dynamics
Andrea Attardi, Timothy Fulton, Maria Florescu, Gopi Shah, Leila Muresan, Martin O. Lenz, Courtney Lancaster, Jan Huisken, Alexander van Oudenaarden, Benjamin Steventon
Development
Colour-coded ascidian embryogenesis
Contact-dependent cell communications drive morphological invariance during ascidian embryogenesis
Leo Guignard, Ulla-Maj Fiuza, Bruno Leggio, Emmanuel Faure, Julien Laussu, Lars Hufnagel,Gregoire Malandain, Christophe Godin, Patrick Lemaire
bioRxiv
Convergence and extension in Xenopus
Spatial and temporal analysis of PCP protein dynamics during neural tube closure
Mitchell T Butler, John B Wallingford
eLife
Modelling development in silico
Theoretical tool bridging cell polarities with development of robust morphologies
Silas Boye Nissen, Steven Rønhild, Ala Trusina , Kim Sneppen
eLife
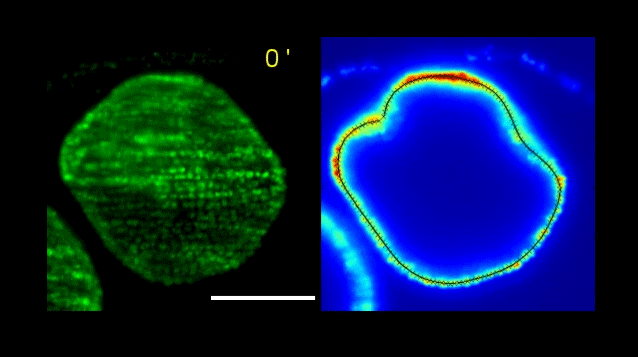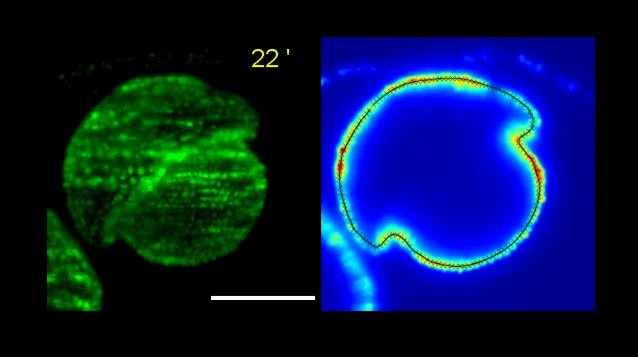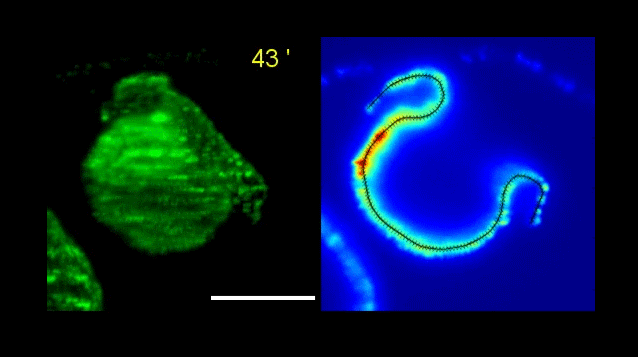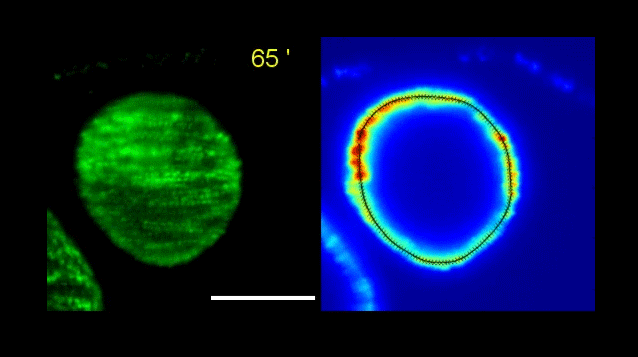
How Volvox turns itself inside out
Pierre A. Haas, Stephanie S. M. H. Höhn , Aurelia R. Honerkamp-Smith, Julius B. Kirkegaard, Raymond E. Goldstein
PLOS Biology
From symmetrical to asymmetrical: the making of the heart tube
Hinako Kidokoro, Sayuri Yonei-Tamura, Koji Tamura, Gary C. Schoenwolf, Yukio Saijoh
Development
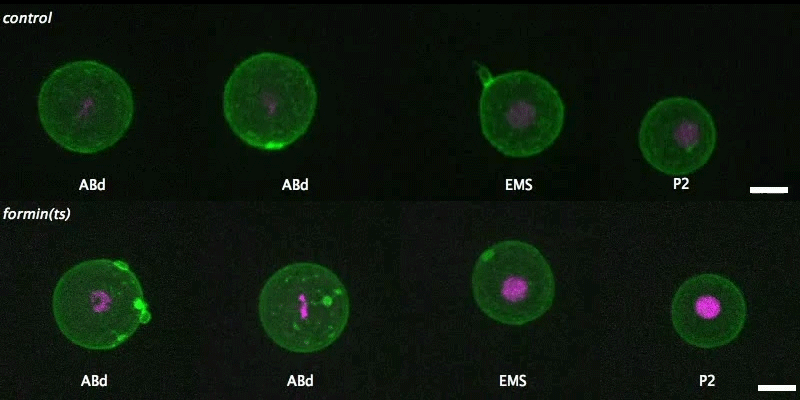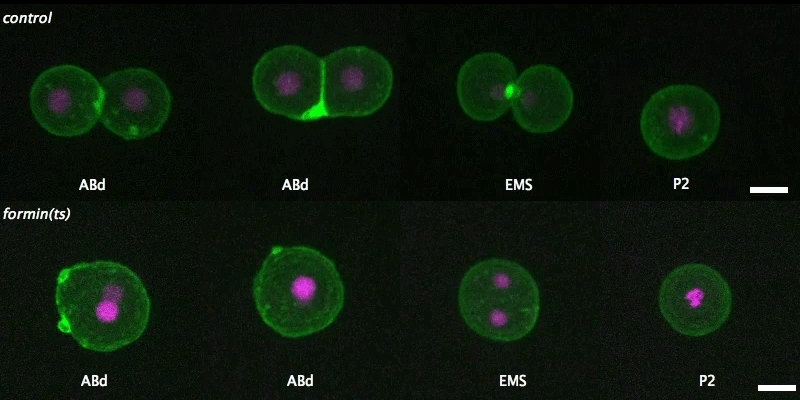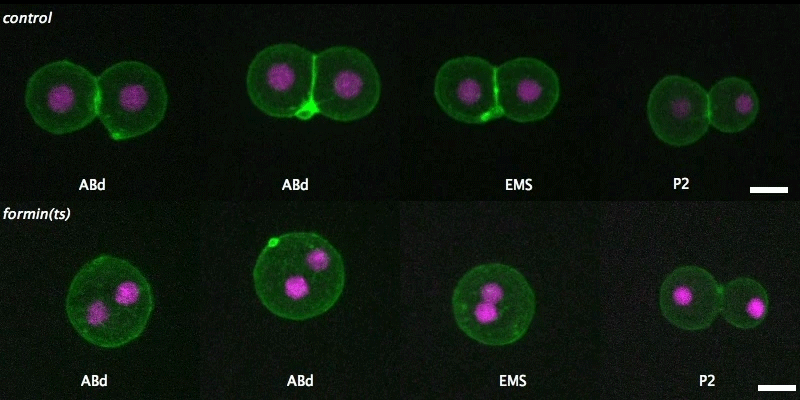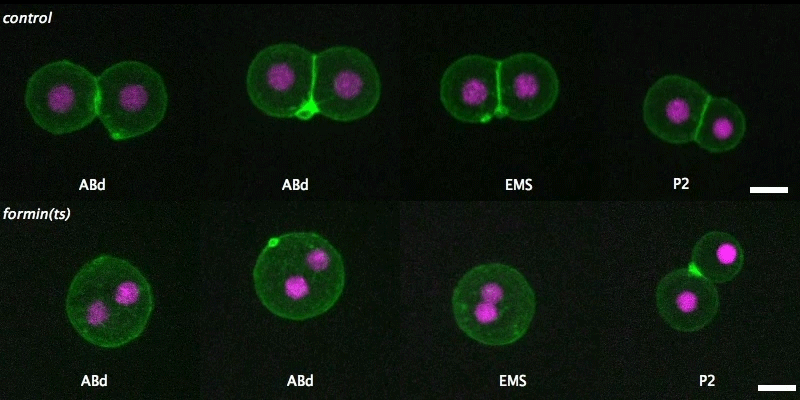
Isolated blastomeres in C. elegans
Cell-intrinsic and -extrinsic mechanisms promote cell-type-specific cytokinetic diversity
Tim Davies, Han X Kim, Natalia Romano Spica, Benjamin J Lesea-Pringle, Julien Dumont, Mimi Shirasu-Hiza, Julie C Canman
eLife









 (4 votes)
(4 votes)