The long road to understanding homeobox genes in the nervous system
Posted by Oliver Hobert, on 1 October 2020
Following the initial discovery of the homeobox in the 1980s in invertebrates and then vertebrates, it became quickly clear that homeobox genes come in two flavors – that of the Antennapedia-like HOX cluster genes and that of the many more non-clustered genes with diverse sequence and expression features (Gehring, 1998). One theme that became evident through expression and mutant analysis in a variety of organisms was the selective expression and function of homeobox genes within the nervous system (Gehring, 1998).
When I started to look for postdoctoral positions in the early 1990s, I was particularly intrigued by mutant phenotypes of several fly and worm homeobox genes (Blochlinger et al., 1988; Doe et al., 1988; Finney and Ruvkun, 1990; Way and Chalfie, 1988), but also by the work of the late Tom Jessell, who proposed a LIM homeobox code in the vertebrate spinal cord (Tsuchida et al., 1994). The simplicity and well-characterized nature of the C. elegans nervous system, as well as its genetic amenability was very appealing to me and, in 1996, I decided to join Gary Ruvkun’s lab. Gary’s lab had not only characterized one of the first C. elegans homeobox genes, unc-86 (Finney and Ruvkun, 1990; Finney et al., 1988); Thomas Bügrlin in Gary’s lab had also used library screening with degenerate probes, a method that now, in the post-genome era, seems quite archaic, to discover the abundance of homeobox genes in this simple organism (Burglin et al., 1989).
In Gary’s lab, I set out to study the expression and function of the LIM homeobox subfamily, which were discovered initially by Marty Chalfie (Way and Chalfie, 1988) and implicated further in neuronal identity specification by Tom Jessell’s lab (Tsuchida et al., 1994). Using emerging GFP reporter technology (Chalfie et al., 1994) and mutant analysis, I determined what turned out to be mostly incomplete expression patterns (owing to the shortcomings of “classic” reporter genes which often just contained fractions of their surrounding gene regulatory regions) and mutant phenotypes that could only be very superficially analyzed (owing to a shortage of markers that allowed for a more in-depth analysis of mutant phenotypes)(Hobert et al., 1998; Hobert et al., 1997; Hobert et al., 1999).
After starting my own lab at Columbia University in 1999, a string of students and postdocs (Zeynep Altun, Adam Wenick, Ephraim Tsalik, Feifan Zhang, Pat Gordon, Vincent Bertrand, Maria Doitsidou, Nuria Flames, Rich Poole, Paschalis Kratsios, Marie Gendrel, Esther Serrano-Saiz, Laura Pereira, among others) continued to work on a small number of specific homeobox genes, digging much deeper into what these genes did in the nervous system. One theme that continued to emerge throughout this analysis was that not only the classic unc-86 and mec-3 genes, studied in impressive depth by Marty Chalfie over the years (Chalfie, 1995), but other homeobox genes as well had a remarkably broad effect on the differentiation of specific neuron types. Rather than regulating only some subset of specific identity features in a neuron, several homeobox genes fulfilled a “master regulatory” role in controlling most, if not all, known identity features of a neuron, through direct initiation and maintenance of terminal differentiation gene batteries. This led me to propose the concept of “terminal selectors” of neuronal identity, a term extended from the Drosophila field where “selector genes” were coined as genes that act earlier in development to specify the identity of developing fields and tissues (Hobert, 2016).
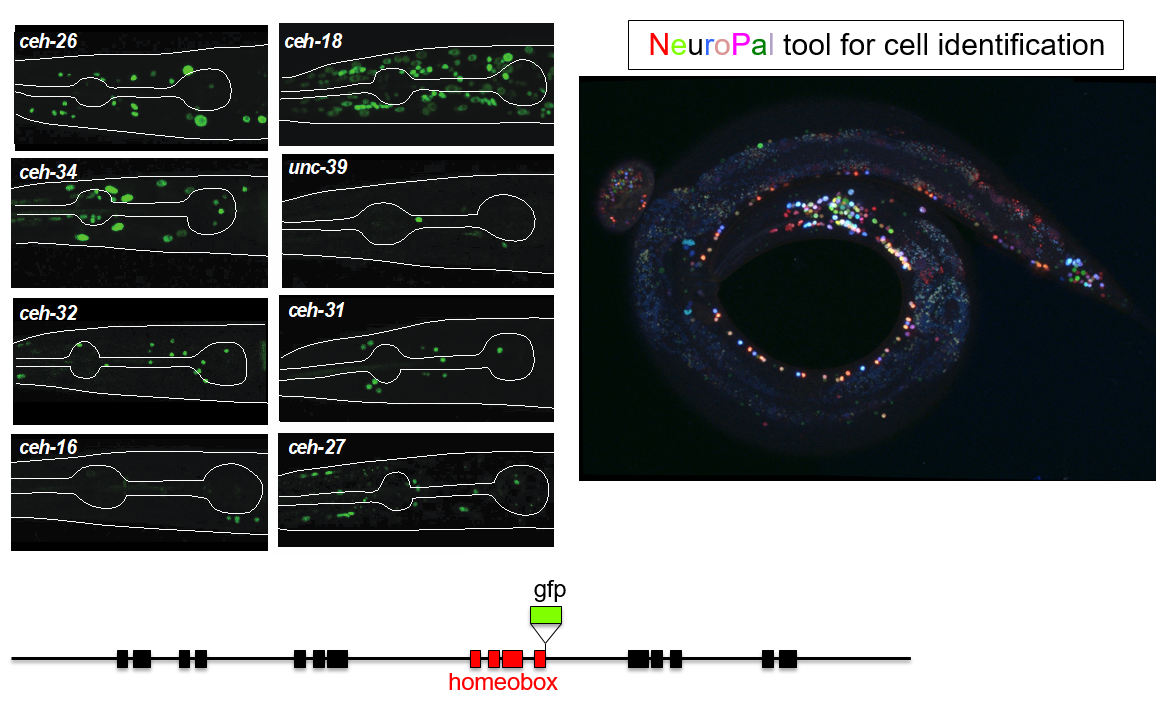
This trajectory finally led to the work of Molly Reilly, a graduate student in my lab, who recently set out to achieve the ambitious goal of describing the expression patterns of the entire homeobox gene family across the entire C. elegans nervous system (Reilly et al., 2020). This tremendous leap forward was, as so often is the case, enabled by novel technology. First, gene expression patterns, or even better, protein expression patterns, can now be much more reliably identified by not just extracting some arbitrary small regulatory region adjacent to your gene of interests to drive a reporter gene. Rather, bacterial recombineering technology enables the reporter tagging of genes in the context of very large genomic intervals containing many genes up- and downstream of the gene of interest (Tursun et al., 2009). Moreover, CRISPR/Cas9 technology even allowed for reporter tagging of an entire locus in the endogenous context (Dickinson et al., 2013). But even with good reagents at hand, identifying sites of expression of a reporter gene across the entire nervous system has traditionally not been a small feat because neurons in C. elegans are tightly packed and their position can be locally variable. Here is where Eviatar Yemini, a postdoc in my lab, came in to solve the long-standing problem of neuronal cell identification. Using multiple distinct fluorophores (excluding GFP), Eviatar built a multicolor landmark strain, NeuroPAL, which unlike Brainbow-style technology, assigned neurons a strictly deterministic color code (Yemini et al., 2019). Crossing NeuroPAL with a GFP reporter strain enables unambiguous identification for the sites of gene expression, anywhere in the nervous system (Figure 1).
Molly exploited these technological advances to (a) tag all but one of the 102 homeobox genes of C. elegans with a fluorescent reporter and (b) identify their sites of expression throughout the entire nervous system. What she found was something I could barely have dreamed of when starting my postdoc in Gary’s lab: Most of the conserved homeobox genes are not only sparsely expressed throughout the nervous system of the worm, but each of the 118 different neuron classes displayed a unique combination of homeobox genes (Figure 2).
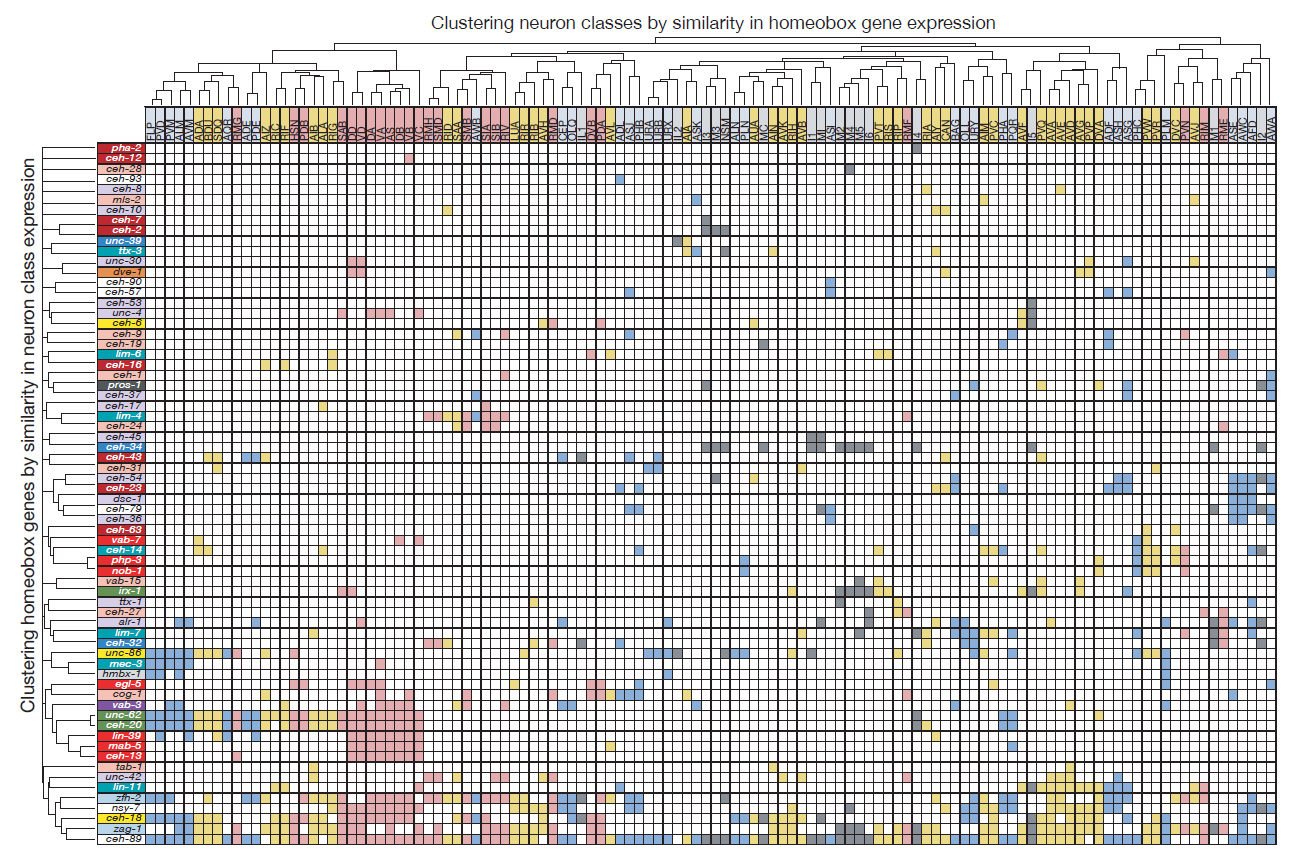
Homeobox genes are thus a comprehensive “descriptor” of neuronal diversity throughout an entire nervous system – a homeobox code for all neurons! Furthermore, the mapping of these homeobox genes led another graduate student, Cyril Cros, to find that neurons previously not known to express or require a homeobox gene, do indeed also require a homeobox gene for their identity specification (Reilly et al., 2020).
This is not the end of the road. The lab remains motivated to test whether indeed every single C. elegans neurons not only expresses, but requires a homeobox gene for their identity specification. Moreover, it remains little explored to what extent we can reprogram the identity of neurons by respecifying their homeobox codes. I am looking forward to see whether work in other systems with more complex brains will also uncover the broad employment of homeobox codes. Recent transcriptome analysis in restricted parts of the flies and mice CNS indeed provides tantalizing hints for similar specificity and selectivity of homeobox gene expression in more complex nervous systems (Allen et al., 2020; Davis et al., 2020; Sugino et al., 2019).
References
Hobert, O. (2016). Terminal Selectors of Neuronal Identity. Curr Top Dev Biol 116, 455-475.


 (2 votes)
(2 votes)