Featured resource: MorphoNet 2.0
Posted by the Node, on 14 April 2026
Our ‘Featured resource’ series aims to shine a light on the resources that support our research – the unsung heroes of the science world. In this post, we learn about the data and functionalities available at MorphoNet 2.0, and hear about new initiatives they are developing.
MorphoNet 2.0: breaking the glass ceiling of 3D+time bioimage curation

Modern microscopy now allows us to image living systems in three dimensions over time. From early embryonic divisions to complex tissue morphogenesis, we can follow every cell with remarkable spatial and temporal resolution.
Yet a critical bottleneck remains. Even with powerful AI-based segmentation tools, such as Cellpose or StarDist, errors are inevitable. In large 3D+time (3D+t) datasets, a seemingly small error rate quickly translates into thousands of mis-segmented cells, broken lineages, and ultimately misleading biological conclusions.
Correcting these errors — a process known as curation — is often so time-consuming that it becomes the real “glass ceiling” of quantitative developmental biology.
MorphoNet 2.0 was designed to break this glass ceiling.
From segmentation to curation
MorphoNet 2.0 is a major evolution of the original platform. Rather than proposing yet another segmentation algorithm, it focuses on a problem that is just as critical, but far less addressed: how to efficiently assess, correct, and validate 3D and 4D segmentations at scale.
The key idea is simple: automated segmentation is only useful if biologists can trust and refine its output. MorphoNet 2.0 makes this process fast, efficient, interactive, and accessible to non-programmers.
A hybrid architecture built for 3D+t data
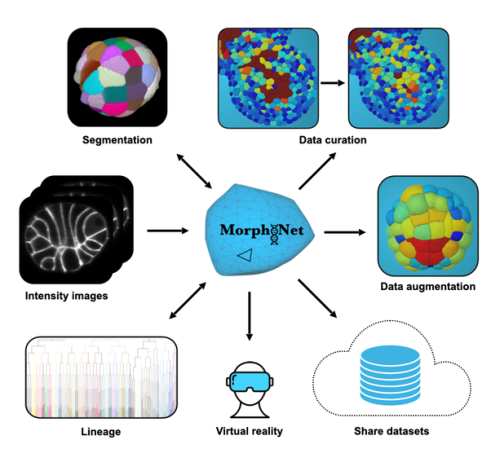
Unlike the original web-based MorphoNet, version 2.0 is a standalone application designed to run on standard research workstations and laptops. Its architecture deliberately bridges two complementary worlds:
- a high-performance 3D interface, powered by the Unity game engine, for smooth exploration of thousands of objects in real time
- a Python backend integrated with powerful scientific image-processing libraries, enabling advanced image analysis, editing, and AI-based segmentation directly on raw data.
Letting the data highlight its own problems
A major challenge in large 3D datasets is simply knowing where to look. Manually inspecting every cell is unrealistic.
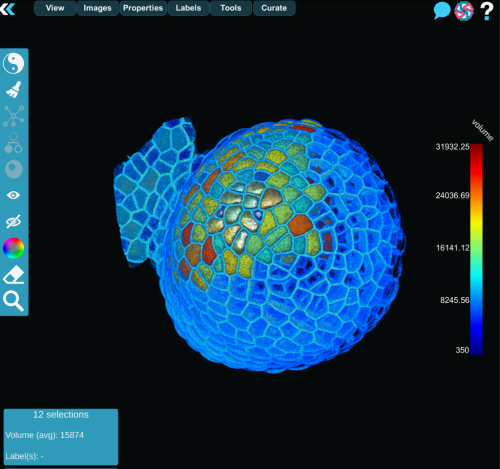
MorphoNet 2.0 addresses this by computing unsupervised quality metrics for each segmented object, such as volume, shape regularity, boundary intensity, or temporal stability. These properties are projected, as color maps, directly onto the 3D meshes or cell lineages, turning potential errors into visible outliers.
While these metrics are not absolute measures of correctness, they act as intelligent guides, directing experts toward regions that deserve closer inspection.
Curation as a local, interactive process
Once a potential error is identified, MorphoNet 2.0 enables rapid local, targeted corrections rather than global reprocessing.
This is achieved through a modular plug-in system that allows users to perform a wide range of operations, including (but not limited to):
- re-segment specific regions using AI or classical methods
- split fused cells or merge over-segmented fragments
- remove specifically small objects
- propagate corrections across time in 3D+t datasets
- edit and repair cell lineages.
Because plug-ins operate only on user-defined regions of interest, most corrections take seconds rather than hours. This transforms curation from a batch process into a true human-in-the-loop workflow, with immediate visual feedback.
MorphoNet 2.0 provides an open plug-in architecture that enables contributors to develop and share custom plug-ins tailored to their own segmentation, tracking, or lineage-repair challenges.
Tested on real biological datasets
This paper demonstrates MorphoNet 2.0’s successful use on five previously published datasets spanning insects, plants, nematodes, echinoderms, and ascidians. These case studies show how the platform can:
- reveal hidden segmentation errors in datasets long considered “finished”
- significantly improve segmentation quality through iterative correction
- polish cell lineages to a level suitable for studying subtle biological variability.
In one example, rare lineage errors scattered across thousands of cells were identified and corrected in a few hours — even in datasets comprising several thousand segmented objects — a task that would have been practically infeasible with traditional tools.
Why this matters for AI and data reuse
High-quality 3D ground truth data are essential for training and benchmarking modern AI models. Yet producing such datasets is extremely costly when curation tools do not scale.
By making 3D+t curation feasible, traceable, and accessible to biologists, MorphoNet 2.0 directly addresses this gap. It helps turn raw automated outputs (“silver truth”) into reliable, reusable datasets (“gold truth”) that can support reproducible analysis, fair algorithm comparison, and community challenges.
Who is MorphoNet 2.0 for?
MorphoNet 2.0 is designed for:
- developmental and cell biologists working with complex 3D or 4D imaging data
- imaging facilities producing reference datasets
- researchers developing or benchmarking segmentation and tracking algorithms.
No programming skills are required to use the platform, but its open, Python-based plug-in architecture allows advanced users to extend it and share new tools with the community.
Looking ahead
MorphoNet 2.0 positions complex 3D curation as a fast and manageable activity rather than a never-ending technical burden. By combining intuitive 3D interaction, unsupervised quality assessment, and local correction, it offers a practical solution to one of the most persistent bottlenecks in modern bioimage analysis.
Find out more
📄 Read the paper
💾 Download MorphoNet 2.0
📚 Documentation & use cases


 (No Ratings Yet)
(No Ratings Yet)