A Day in the Life of… a Zebrafish lab
Posted by Andrew Mathewson, on 8 October 2013
My name is Andrew Mathewson and I am a fourth year graduate student in the University of Washington’s Molecular and Cellular Biology program. Whereas most my friends from college are pursuing productive careers in medicine, social justice, or environmental preservation (or just trying to get by), I spend most my days deep in a basement encouraging small fish to mate so that I can steal their babies for science. I am currently pursuing a PhD in the lab of Dr. Cecilia Moens in the Basic Sciences division at the Fred Hutchinson Cancer Research Center (FHCRC) in Seattle, Washington. The Moens lab exclusively studies the zebrafish as a model for vertebrate development, focusing primarily on hindbrain patterning and cell migration. My project investigates the interplay between basic cell polarity signaling and the ability of neurons to move throughout the developing brain.
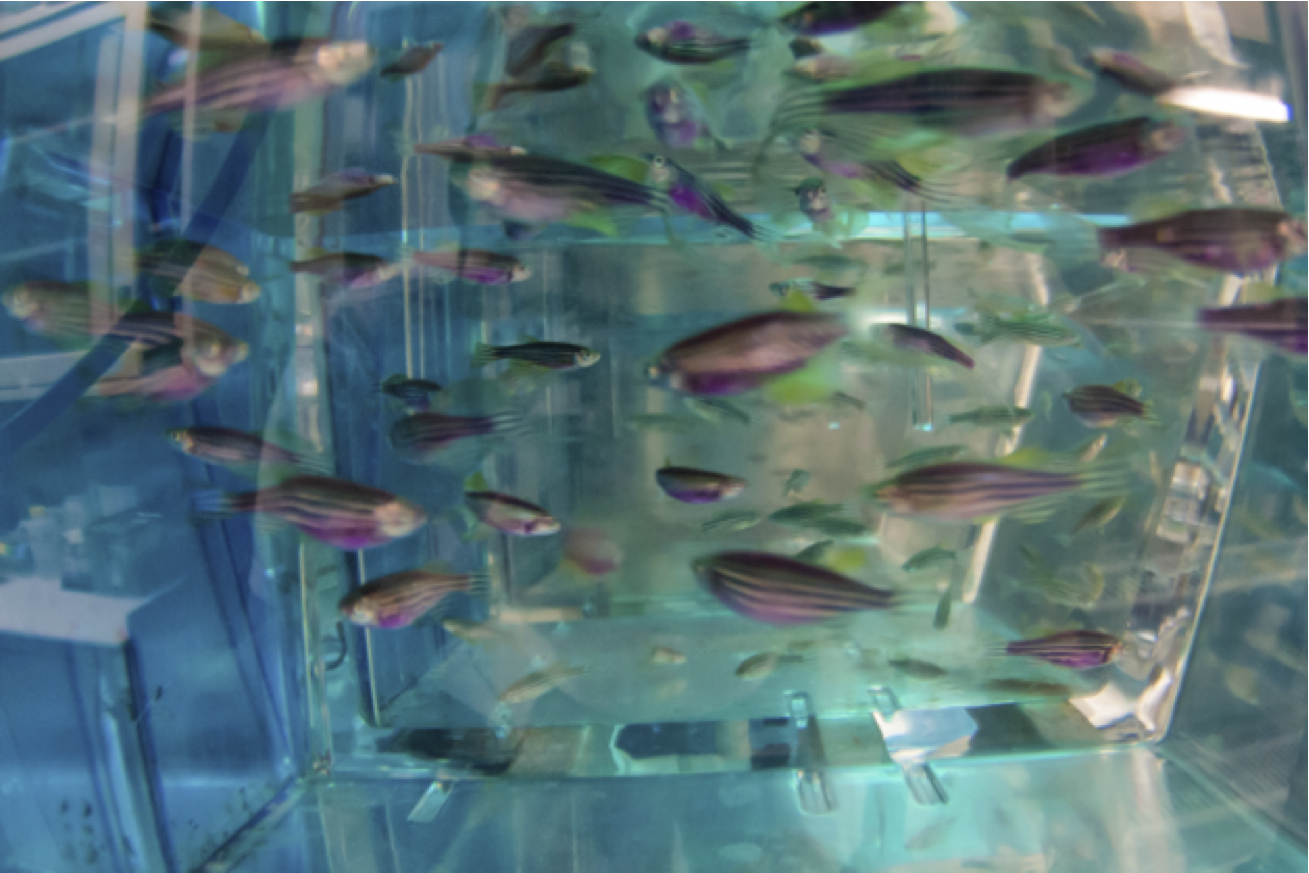
A tank of adult zebrafish expressing a red fluorescent protein in all muscle tissues bustles with excitement during their morning feeding.
Zebrafish are small tropical freshwater fish species native to the Indian subcontinent that were popularized as aquarium fish many years ago. They were pioneered as a model system for scientific research by George Streisinger and colleagues at the University of Oregon, where the Zebrafish International Resource Center (ZIRC) is now located. Many developmental biologists have adopted these robust little critters for their research in the last several decades due to the fact that their biology is very amenable to experimentation and observation. Zebrafish develop externally from a single cell to a rather complex larval fish in just a couple of days, meaning that in just a few hours, zebrafish speed through developmental processes that take weeks or months in other vertebrate models. Most incredibly – they remain entirely transparent while doing so. A single mating pair of zebrafish can produce upwards of a thousand embryos in a single morning, making zebrafish a powerful system for generating large experimental datasets. Zebrafish has triumphed as a system for forward genetic screens and researchers have developed innumerable transgenic lines for the scientific community in recent years. It is the unique combination of developmental speed, optical transparency, and the rapidly growing array of genetic tools that have made zebrafish a popular system for modern developmental research.
If you have never worked in a fish lab you might wonder what goes on behind the locked doors of our facility deep in the bowels of the FHCRC. In the evening before most of my experiments I set aside a lucky few male and female fish with the mutations and/or transgenes I am using and place them into crossing tanks that allow the fish to see and smell but not touch one another. At 9AM the next morning the facility lights turn on and the fish are frisky after a long night’s rest. One of my favorite parts of my job is when I get to remove the transparent dividers from crossing tanks, allowing the fish to finally interact after a long night of frustrated solitude. Healthy males will almost immediately begin aggressively courting any willing (or not) female in sight. This courting mostly looks like a game of tag, where the female is “it”. Usually in a matter of minutes the females will begin to release eggs that the males dutifully fertilize and I patiently collect for my morning experiments.
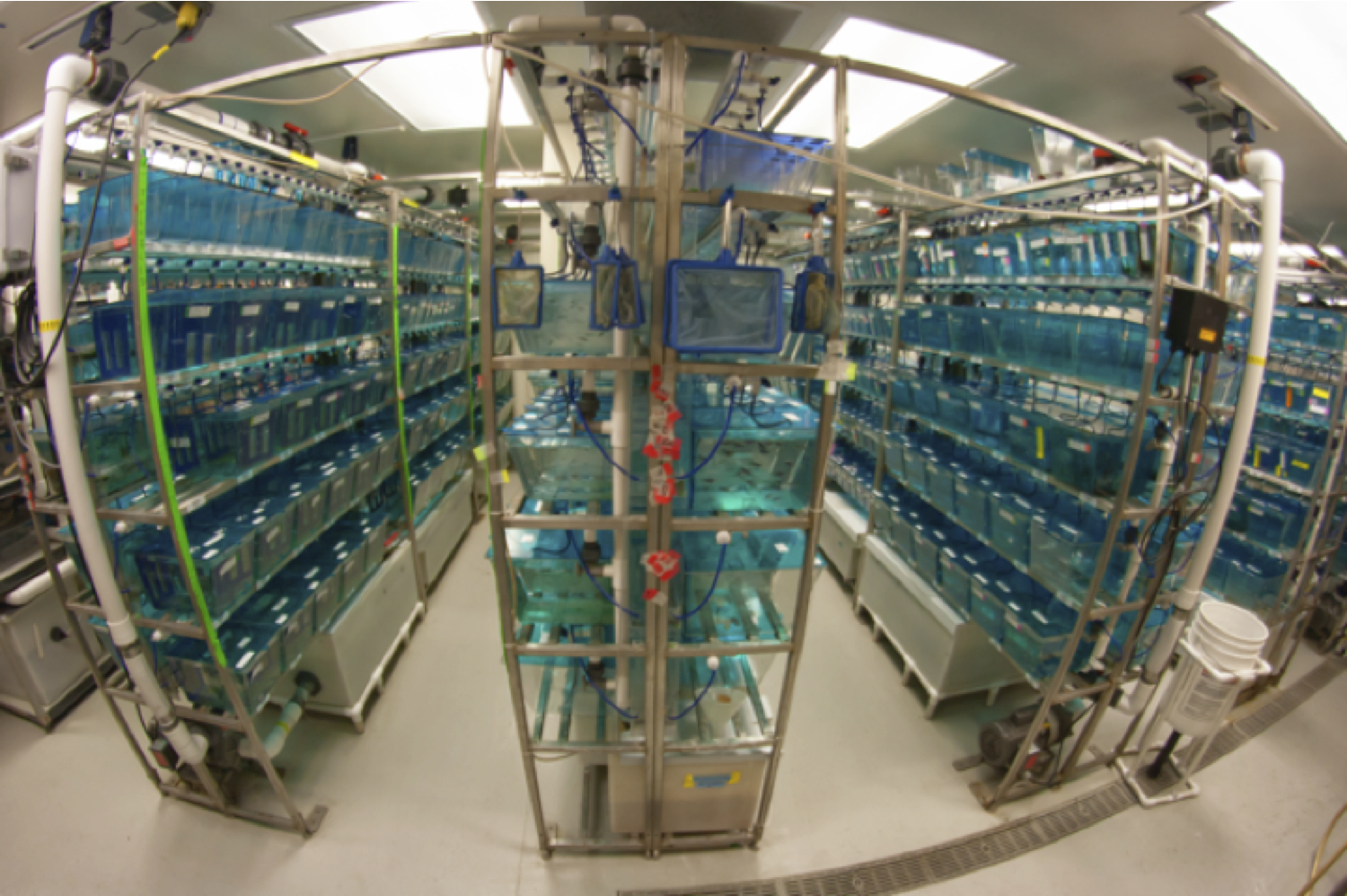
The Moens Lab main fish facility
While any research demands practice and patience, zebrafish work in particular demands exceedingly steady hands, sharp eyes, and an even sharper pair of forceps. A typical day can involve anything from molecular cloning and sequence analysis, confocal microscopy and imaging, visual or genetic screening of mutant embryos, all the way up to actual wrangling of adult fish. Most of my current work involves studying developmental processes that occur within the first two days of life. This means that I spend the first half of most work weeks crossing fish lines and manipulating embryos and the second half imaging and analyzing my results. One technique I’ve tried to master is cell transplantation between embryos of different genotypes in order to ask questions about cell-specific gene function. Zebrafish is a great system for this type of work since embryos develop externally, are produced in large numbers, and can tolerate a lot of manipulation. However, most of my current experiments revolve around cloning interesting bits of DNA together and injecting them into newly laid single-cell embryos with infinitesimally small glass needles. I can efficiently create mosaic embryos (where some cells express my DNA and others do not) by mixing my DNA with transposase to increase genomic integration. Most people can comfortably inject hundreds of embryos per hour making it possible to rapidly conduct multiple experiments in parallel. Once I’ve injected my embryos, I spend the rest of my days in dark rooms peering through dissection scopes trying to discern if my manipulations change how cells behave during development. Even though these cells tend to be labeled with at least one type of fluorescent protein, this process can be difficult because zebrafish embryos are quite small. Therefore, once I’ve identified embryos that express my injected DNA or contain successfully transplanted cells, I’ll typically anesthetize the embryos, mount them in agar, and image them on one of our confocal microscopes for closer analysis. The coolest part of this is that I do most of my imaging on living embryos, so I can see how cells behave in their natural environment as the fish continue to develop. This also allows me to use the fish in further experimentation or I can raise them to generate new transgenic lines. Depending on the type or combination of transgenes I want for my future experiments, it can sometimes take several generations and many months to actually generate a new stable transgenic line.
Not surprisingly, it can take a little bit of work keep my experimental stocks of hundreds (or thousands) of fish alive and happy throughout my experimental timeline which in some cases can extend for years. Fortunately the Moens lab is able to hire a full-time fish technician that cares for our facility and generally keeps things running smoothly. She maintains cultures of brine shrimp (aka sea monkeys, a zebrafish’s favorite snack) and rotifers (microscopic water creatures that baby fish love to munch on), feeds the entire facility both live and dry food several times a day, cleans used tanks and nets, helps keep track of our stocks as they mature and age, and euthanizes aging or diseased fish, to name just a few of the amazing things she does for us. Occasionally our technician takes a day off work (like any normal human being), leaving us researchers responsible for our own fish. It’s usually not a big deal—everyone in the lab is at least nominally trained in fish facility maintenance for our own work, and ever since the funding drop, everyone has gotten used to sacrificing the odd weekend to keep our stocks alive. Yet when things go wrong it forces one to recognize the sheer amount of effort required to generate the massive amounts of data produced by the zebrafish research community. Just the other day it was my turn to feed our baby tanks but it turned out that our rotifer cultures had crashed. Our babies went hungry that day—slowing their growth and decreasing their chances of survival, meaning that we can’t count on them for future experiments. Fortunately these small disasters are pretty rare because we have such an awesome fish technician and everyone in lab pays attention to the needs of the fish.
Disease and parasites are a major concern when running a large facility of animals. We try to minimize introducing unwanted pests into our facility by keeping an entirely separate “quarantine” facility for housing fish sent from other labs as well as regular sterilization of media and equipment. Even with these precautions it is impossible to prevent the occasional unwanted pest from getting in. It’s not uncommon to discover an odd arthropod swimming in one’s dishes while sorting embryos. Though tiny creatures like these are usually a normal part of a complex water environment in nature, they can devastate young (or even adult) zebrafish populations in lab. Instances like these remind us that the hard work we do to keep our facility clean and our fish happy isn’t just because the FHCRC requires us to—it’s so we all can continue to do our research.
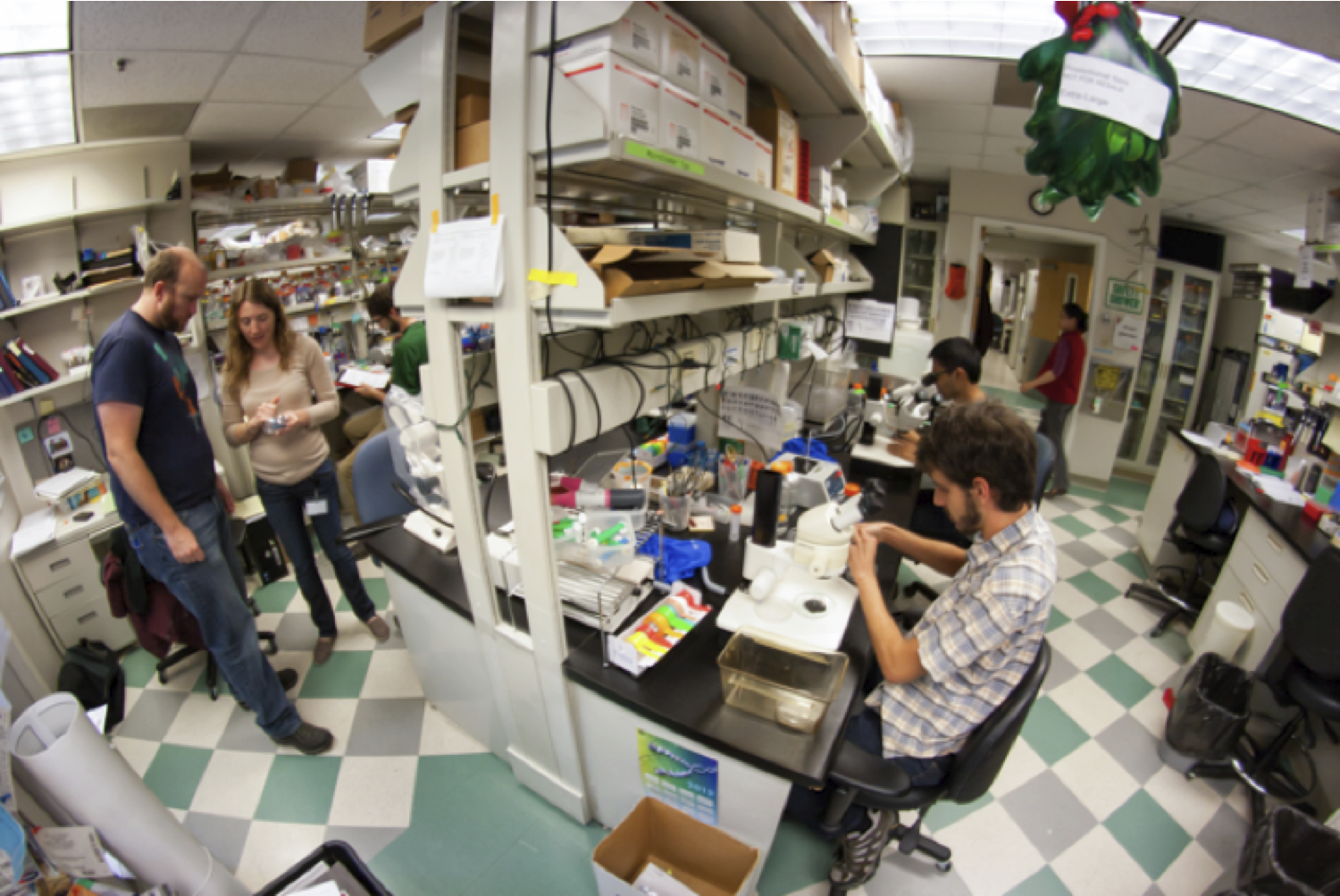
Lab mates cluster around our central work bench most afternoons. Adam Miller and Crystal Davey inspect injection needles (on the left) for cell transplantations while Daniel Berman and Arish Shah (on the right) sort zebrafish embryos.
I like working with zebrafish. While I find the pace of yeast and tissue culture almost frenetic, and the speed of Arabidopsis growth to be at times overly meditative, zebrafish grow and live at a rate that I can relate to. Most experiments fall into a natural daily rhythm in sync with our fish facility’s diurnal cycle. From my experience, zebrafish labs tend to foster a friendly and collaborative lab atmosphere due to the tiny bit of mutual dependence required for keeping stocks alive and experiments running (or maybe zebrafish work just attracts friendly people). Given that we usually want our fish to stay healthy, we have a pronounced lack of the toxic chemicals one usually associates with this type of research, which means the lab can be a comfortable space for everyone. One nice thing about zebrafish work in general is that the fish do most of the work. Once I’ve done my manipulations early in the week I place my embryos in a water filled dish in an incubator and don’t touch them until they are old enough to study. Each fish has its own yolk sac meaning that they don’t need food (or much of anything) for the first week of development. Though zebrafish is a great system for cell transplantation and other active manipulations, the fish really shine as a model organism once they are old enough to look at. Whether you are tagging an interesting protein with GFP, tracking transplanted cells with viable dyes, screening for mutant phenotypes, or just watching the precise choreography of development, zebrafish is the system to turn to. The fact that I can mount living transgenic embryos and image their (often glowing and multicolored) cells as they grow and interact with one another just blows my mind. This aspect is what caught my attention early in graduate school and is a big part what keeps me working towards my Ph.D. It’s hard to describe the beauty of a living, glowing brain as seen through a powerful confocal microscope, and even after several years of works it never fails to leave me in awe.
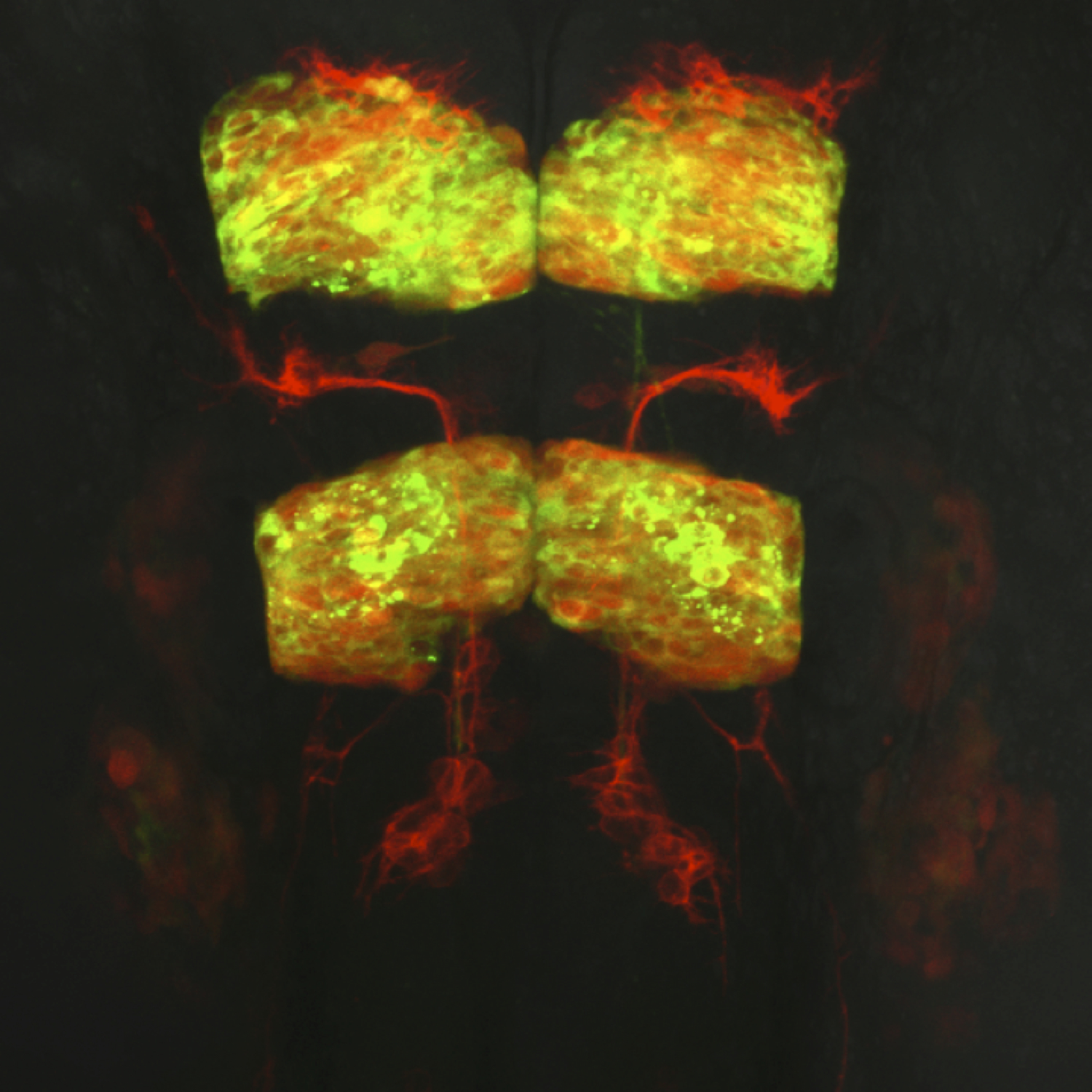
Dorsal view of a live transgenic zebrafish embryo’s hindbrain at 24 hours post fertilization. Facial branchiomotor neurons express membrane red fluorescent protein and migrate posteriorly through hindbrain segment rhombomere 5 while leaving behind trailing axons. Rhombomeres 3 and 5 expresses a red fluorescent protein marker as well as a cytoplasmic protein tagged with green fluorescent protein.
 This post is part of a series on a day in the life of developmental biology labs working on different model organisms. You can read the introduction to the series here and read other posts in this series here.
This post is part of a series on a day in the life of developmental biology labs working on different model organisms. You can read the introduction to the series here and read other posts in this series here.





 (17 votes)
(17 votes)
I have fond memories of my days of injecting and transplanting zebrafish – thanks for bringing them back Andrew!
Thank you for making me apply for a research assistant position working with zebrafish. This post truly inspired me. Keep doing what you are doing, you and your entire lab members (including the wonderful fish technician) are amazing human beings. Xxx