All beauty must die
Posted by PCDLab, on 9 July 2018
Anna Daneva, Zhen Gao and Moritz Nowack tell the story behind their recent paper in Nature Plants
The transience of flowers is proverbial. Degeneration of flowers is elicited after successful pollination by the onset of seed and fruit development. However, also unpollinated flowers do not last forever – on the contrary, the life span of unpollinated flowers is a tightly regulated trait that differs greatly among plant species. Some plant species like orchids of the genus Phalenopsis have very long-lived flowers that can stay receptive for weeks, even months. Most plant species, however, sustain unpollinated flowers for much shorter time spans. In the most extreme cases mere hours pass between flower opening and withering.
In the Programmed Cell Death lab at the VIB Center for Plant Systems Biology we are using the model plant Arabidopsis thaliana to study the longevity and senescence of unpollinated flowers. Although Arabidopsis is the most advanced model species for plant developmental biology, it is actually not particularly convenient to investigate the biology of flower senescence. Arabidopsis flowers are tiny, and moreover they are self-pollinated, so that the male floral parts (anthers) have to be either removed manually by emasculation, or male-sterile mutants have to be used.

Every Arabidopsis researcher knows how to perform emasculation and pollination in order to cross different mutants, marker lines, or accessions. And many of us know that a flower emasculated on Friday might already have started to senesce by Monday, hence making successful pollination and seed set inefficient or impossible.
In order to observe and stage flower withering in Arabidopsis, we set up a macroscopic imaging system using a single lens reflex camera equipped with a macro objective. Taking images every 10 minutes under constant light enabled us to monitor flower senescence in detail. We could observe that concomitant with the withering of petals and sepals, also the floral stigma was degenerating.
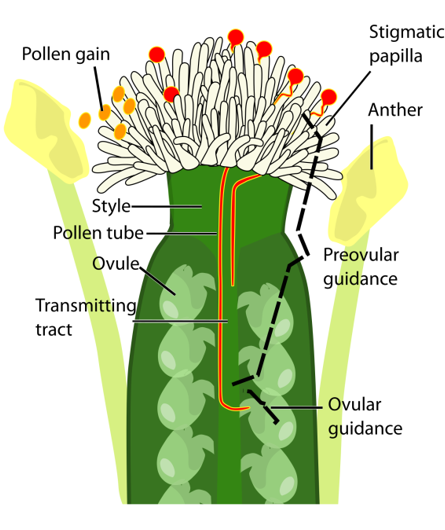
The stigma of flowers is the primary receptive surface for pollen grains. In Arabidopsis the stigma consists of over 200 elongated fingerlike cells, the so-called stigmatic papillae, specialized epidermal cells that serve to intercept the pollen. In Arabidopsis, hundreds of pollen grains compete to fertilize about 50 ovules located inside the pistil of each flower. Once in contact with the stigma cells, pollen grains will hydrate, and germinate to form a single tip growing cell, the pollen tube. The pollen tube penetrates the papilla cell wall, growing in between the cell wall and the plasma membrane to the base of the papilla cell. From there on, guided by chemical cues, the pollen tube grows through the style and the transmitting tract to the ovules. There, it will release its two sperm cells to fertilize the egg cell and the central cell of the female gametophyte, thus initiating embryo and seed development.
The structure of the intact Arabidopsis stigma appears a bit like a little hedgehog due to the hundreds of erect elongated papilla cells. Once flower senescence sets in, individual papilla cells start to break down, leading first to a ragged, and finally to a completely collapsed appearance of the stigma.
Pollination assays at different time points during this process showed that stigma degeneration coincided with a sharp decline in seed set, suggesting that viable papilla cells are necessary for successful reproduction. To test this hypothesis, we specifically ablated papilla cells by expressing diphtheria toxin chain A (DT-A) under a stigma-specific promoter. As we had assumed, DT-A induced stigma degeneration likewise caused a strong reduction of seed set.
In order to quantify stigma longevity, we faced a considerable problem – the macro setup was only able to follow a single flower at a time, and a price tag of about 1400 Euro per unit made the parallel acquisition of time courses impracticable. Coincidence came to our aid: Setting up a skype conference call we noticed that a cheap webcam without autofocus can be brought very close to an object (in that particular case a certain PhD student’s uvula) and amazing magnification can be achieved. After some experimentation with different webcams, we rigged up a phenotyping system with 20 webcams. We used a custom-made script to operate the open source camera surveillance software so that each camera acquires one picture frame every 10 minutes. Although the image quality was of course not comparable to the conventional macro lens setup, in the end of the day we had established a phenotyping platform with 20 imaging units for mere 250 Euros.

Analysis of the time lapse images revealed a remarkable reproducibility of stigma collapse on average around 56 hours after emasculation of a flower at the developmental stage 12c1.
The apparently precisely timed collapse of papilla cells suggested that an active cell death process might be occurring in Arabidopsis stigmata. Based on a large-scale meta-analysis of mRNA transcriptome profiles, our lab has established promoter-reporter lines of genes transcriptionally upregulated prior to a number of developmentally regulated programmed cell death (PCD) events2. Microscopic analyses revealed that most promoter-reporters come up during stigma senescence, thus linking stigma degeneration with established forms of developmentally controlled PCD processes3. Microscopic imaging of papilla cells is not trivial; although the individual cells are thankful objects for cell biological analyses, the stigma as a whole is a rather large and very sensitive three-dimensional structure. In order to investigate cell biological hallmarks of PCD in living stigmata, we could not use conventional slide-and-coverslip setups lest we create injuries and mounting artefacts in papilla cells. After testing diverse setups, we found that mounting the entire flower in an agar block and viewing the stigma from top with a long working-distance water dipping lens on an upright Zeiss 710 confocal microscope allowed a minimally invasive way to perform live-cell imaging of papilla cell death. In order to visualize cell death, we used a live/death stain in which fluorescein-diacetate (FDA) fluorescence indicates living cells, while nuclear staining of the membrane impermeable stain propidium iodide (PI) reveals plasma-membrane permeation as a committing step of cell death.
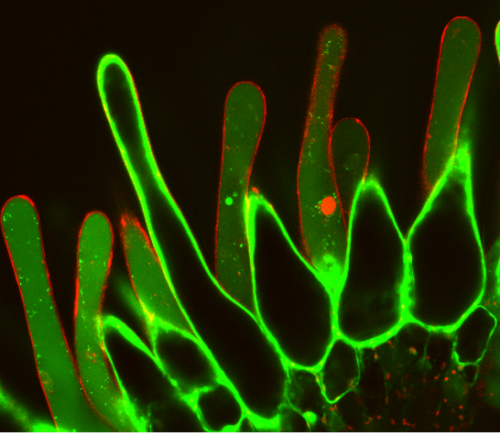
Alternatively, we used a tool developed in our lab, a tonoplast integrity marker (ToIM) line4. This plant line expresses a cytoplasmic green fluorescent protein (cGFP) and a polycistronically produced vacuolar red fluorescent protein (vRFP). Living cells display clearly separated GFP and RFP domains indicating an intact vacuolar membrane (tonoplast). Collapse of the large central vacuole in mature plant cells, another hallmark of plant programmed cell death5, is visible as merging of red and green fluorescence signals.
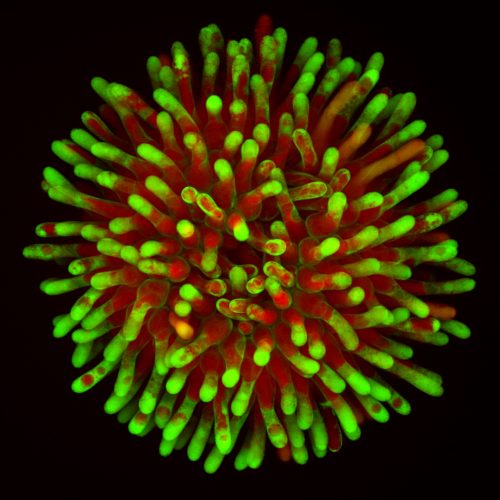
Our analyses revealed that as in the macroscopic setup, stigma degeneration occurred over a time span of 12 hours. Fascinatingly, the death of individual papilla cells followed a pattern from the periphery towards the center of the stigma, and often small clusters of individual cells underwent vacuolar collapse almost synchronously. On the level of individual cells, the PCD program occurred surprisingly fast; vacuolar collapse, plasma membrane permeation, nuclear fragmentation, and finally cellular collapse occurred within about one hour. The reproducibility of these successive events confirmed our hypothesis of a tightly regulated cell death program.
In order to discover regulators of this cell death process, we set out to perform an RNA-seq analysis monitoring the transcriptional changes occurring in the stigma over time. We manually dissected the stigmata from flowers at 1, 2, and 3 days after emasculation, corresponding to young, mature, and senescent flower stages. In total we had to stage, emasculate, and dissect way over individual 2000 flowers, a feat that we only managed with lots of training and a great team effort.

Illumina RNA-seq revealed a large number of differentiallty regulated genes (1180 genes out of more than 25 000 predicted genes in Arabidopsis) over the course of stigma senescence. Interestingly far more genes were upregulated (897) than downregulated (283), again indicating an actively controlled program controlling stigma senescence and papilla cell death.
RNA sequencing also confirmed the strong upregulation of developmental PCD-associated genes. In order to identify key regulators of this process, we investigated the expression profiles of transcription factors (TFs). Interestingly, the plant specific NAC TF family was strongly overrepresented in senescence-associated TFs. Among the most strongly upregulated NAC TFs was the well-established leaf senescence regulator ORESARA1/ANAC092 (ORE1, Korean for “long-living”), and the flower-specific ANAC074, a previously uncharacterized NAC TF that we dubbed KIRA1 (KIR1) after the killer “Kira” in the Japanese manga “Death Note”.
Using our webcam-phenotyping platform, we investigated an established knock-out mutant of ORE1 and found a slight, but not significant delay of stigma collapse. Interestingly, a newly established KIR1 knock-out mutant showed modest, but significant extension of stigma life span. We crossed the mutants and the resulting kir1 ore1 double mutant showed a clear synergistic effect which led to a doubled life span in comparison to the wild type. We also generated lines overexpressing dominant repressive mutant versions of KIR1 and ORE1, and some of these lines had an even stronger effect with some stigmata only collapsing at 11 days after emasculation. Intriguingly, both recessive kir1 ore1 loss-of-function mutations, as well as expression of dominant repressive alleles showed an uncoupling of stigma life span from floral organ longevity. Although the stigma life span was considerably increased, sepals and petals senesced similar to the wild type.
In a complementary approach, we investigated KIR1 gain-of-function mutants. Estradiol-inducible systemic misexpression of KIR1 in seedlings caused a rapid growth arrest caused by a widespread ectopic cell death. This demonstrates that the transcriptional program controlled by KIR1 is sufficient to elicit cell death outside of the stigma context. When inducing precipitate KIR1 expression specifically in the stigma, the papilla cell death and loss of receptivity occurred significantly earlier than in estradiol treated wild types or mock treated mutant lines. These results demonstrated that KIR1 functions to actively terminate the receptive life span of the flower by promoting a cell death program in the stigma.

A central question arising from this investigation was of course whether aged but viable kir1 ore1 loss-of-function mutants could be successfully pollinated. When performing a pollination time series, we discovered that in dominant repressive KIR1 and ORE1 mutants, there was as significant, but rather modest extension of flower receptivity. In the recessive kir1 ore1 double mutant, there was no extension of floral receptivity at all. We used pollen from a transgenic Arabidopsis line that expresses β-glucuronidase (GUS) under a pollen-tube specific promoter to visualize pollen tube growth on pollinated stigmata after addition of the GUS substrate X-Gluc6. Microscopic analyses showed that pollen hydrated and germinated, but that pollen tube growth on aged mutant stigmata was strongly reduced in comparison to younger wild type or mutant stigmata. No pollen tubes could be detected to enter the style or the transmitting tract of the aged mutant flowers, suggesting that KIR1 / ORE1 loss of function is sufficient to suppress age-induced stigma PCD, but not sufficient to maintain stigma function in a corresponding fashion.
In summary, our research on stigma senescence suggests that a KIR1-ORE1 dependent cell death program actively terminates stigma and flower receptivity. However, as suppression of this cell death program is not sufficient to extend stigma function, we assume there must be additional, KIR1 – ORE1 independent pathways that either passively or actively terminate stigma function in the absence of cell death. With research going on in our lab we attempt to research these pathways in order to effectively modulate flower receptivity.
While Arabidopsis serves as a model system to discover these pathways, our research might also be applicable to outcrossing crop species. An extension of floral receptivity, especially under spells of environmental stresses, which are deleterious to plant reproduction, might be a key strategy to stabilize the yield of seed and fruit bearing crops7.
References
1 Christensen, C. A., King, E. J., Jordan, J. R. & Drews, G. N. Megagametogenesis in Arabidopsis wild type and the Gf mutant. Sex. Plant Reprod. 10, 49-64 (1997).
2 Olvera-Carrillo, Y. et al. A conserved core of programmed cell death indicator genes discriminates developmentally and environmentally induced programmed cell death in plants. Plant physiology 169, 2684-2699 (2015).
3 Daneva, A., Gao, Z., Van Durme, M. & Nowack, M. K. Functions and Regulation of Programmed Cell Death in Plant Development. Annual Review of Cell and Developmental Biology (2015).
4 Fendrych, M. et al. Programmed cell death controlled by ANAC033/SOMBRERO determines root cap organ size in Arabidopsis. Current Biology 24, 931-940 (2014).
5 Van Doorn, W. G. Classes of programmed cell death in plants, compared to those in animals. Journal of Experimental Botany 62, 4749-4761 (2011).
6 Johnson, M. A. et al. Arabidopsis hapless mutations define essential gametophytic functions. Genetics 168, 971-982, doi:10.1534/genetics.104.029447 (2004).
7 Ferradás, Y., López, M., Rey, M. & González, M. V. Programmed cell death in kiwifruit stigmatic arms and its relationship to the effective pollination period and the progamic phase. Ann. Bot. 114, 35-45, doi:10.1093/aob/mcu073 (2014).


 (4 votes)
(4 votes)