Application of Fucci technology to analysis of endoreplicating cells
Posted by matsushi, on 18 November 2013
Endoreplication (endoreduplication or endomitosis) is the process by which a cell undergoes successive rounds of DNA replication without an intervening mitosis and its accompanying cytokinesis. Developmentally programmed endoreplication causes differentiating cells, such as TGCs and megakaryocytes (MKCs), to become highly polyploid. Our recent study published in Development (Sakaue-Sawano et al., 2013) comparatively characterized endoreduplication (TGCs) and endomitosis (MKCs) using Fucci (Fluorescent Ubiquitination-based Cell Cycle Indicator) technology. Our long-term imaging experiments also enabled monitoring of endoreplication for weeks, which revealed that the transition from endoG2 to endoG1 in MKCs varied with the polyploidy level.
The Fucci technology harnesses the cell cycle dependent proteolysis of two ubiquitin oscillators, human Cdt1 and Geminin, which are the direct substrates of the SCFSkp2 and APCCdh1 E3 ubiquitin ligase complexes, respectively (Sakaue-Sawano et al., 2008).
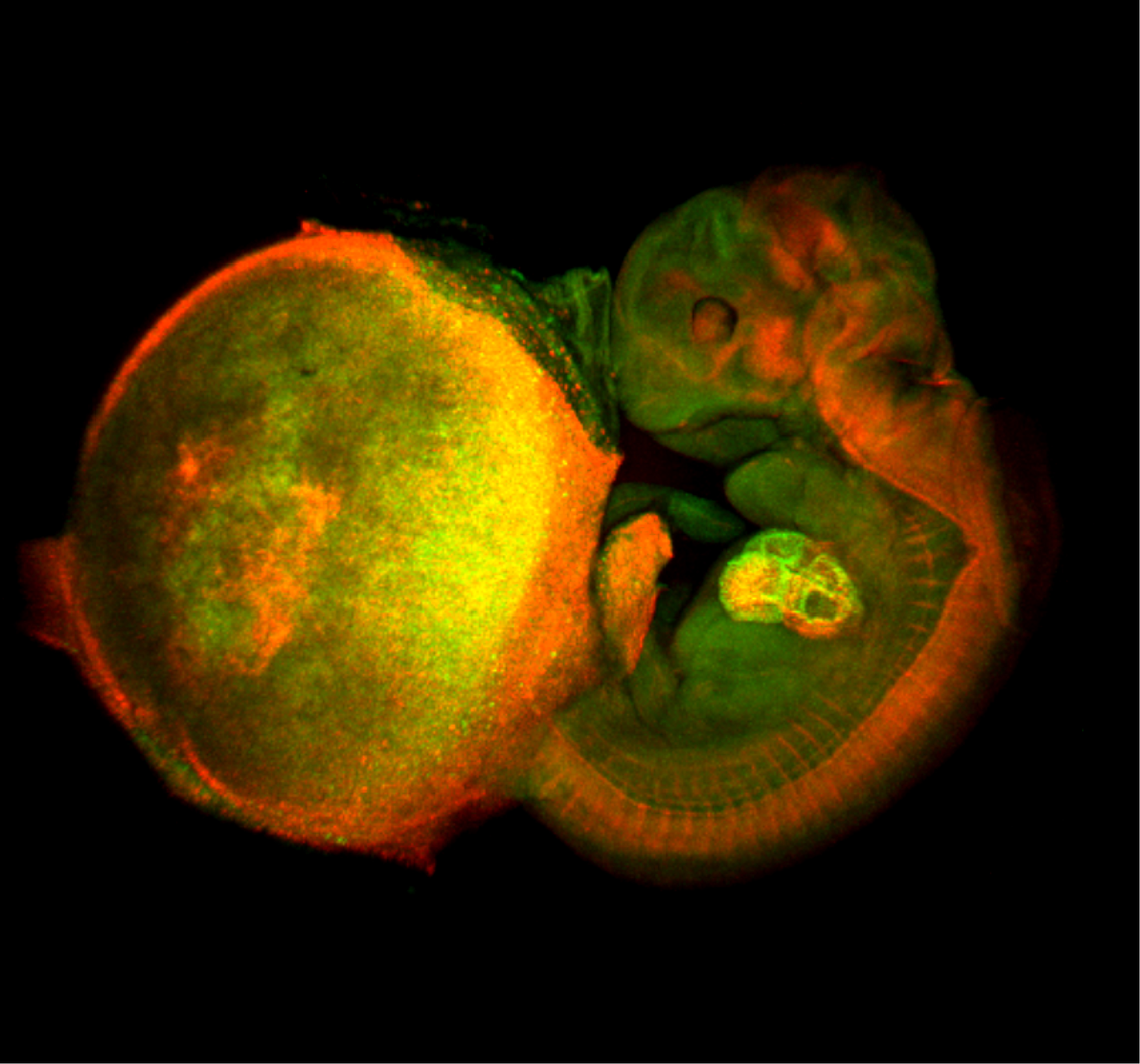
The image shown above is a comprehensive perspective view of cell-cycle progression in a mouse fetal-placental unit (embryonic day 11.5) that was fixed with 4% formaldehyde and then optically cleared by the Scale technique (Hama et al., 2011). The fluorescent cell cycle indicator Fucci was expressed ubiquitously in the tissues. Green and red signals indicate the presence of FucciS/G2/M (mAG-hGem(1/110)) and FucciG1(G0) (mKO2-hCdt1(30/120)) probes, and signify cell proliferation and differentiation, respectively. The developing heart inside the fetus is most clearly illuminated. Remarkably, the enormous nuclei of endoreplicating trophoblast giant cells (TGCs) scattered in the placenta are highlighted in green and red, indicating endoS/G2 and endoG1 phases, respectively.
We would like to express our concern about the inaccurate usage of Fucci terminology in recent literature. While an increasing number of papers report cell cycle dynamics by use of the Fucci technology, the two Fucci probes, mKO2-hCdt1(30/120) and mAG-hGem(1/110), are often called mKO2-Cdt1 and mAG-Geminin, respectively, without annotation. However, this terminology misleads readers into believing that the Fucci probe contains the entire Cdt1 or Geminin protein. In fact, quite a few researchers are wondering whether such probes perturb the cell cycle regulation in recipient cells. In our original study of Fucci (Sakaue-Sawano et al. 2008), we made considerable effort to extract the regulatory domains (ubiquitination domains) from Cdt1 and Geminin. After performing long-term time-lapse imaging of numerous constructs, we demonstrated that amino acid residues 30-120 of human Cdt1 (hCdt1(30/120)) and amino acid residues 1-110 of human Geminin (hGem(1/110)) are both necessary and sufficient for this purpose. We would like to request that future studies using Fucci technology should include the full names of the probes, mKO2-hCdt1(30/120) and mAG-hGem(1/110), in the methods.
Hama, H., Kurokawa, H., Kawano, H., Ando, R., Shimogori, T., Noda, H., Fukami, K., Sakaue-Sawano, A. and Miyawaki, A. (2011). ‘Scale: a chemical approach for fluorescence imaging and reconstruction of transparent mouse brain’, Nat. Neurosci. 14: 1481-1488.
Sakaue-Sawano, A., Kurokawa, H., Morimura, T., Hanyu, A., Hama, H., Osawa, H., Kashiwagi, S., Fukami, K., Miyata, T., Miyoshi, H., Imamura, T., Ogawa, M., Masai, H. And Miyawaki, A. (2008). ‘Visualizing spatiotemporal dynamics of multicellular cell-cycle progression’, Cell 132: 487-498.
Sakaue-Sawano, A,. Hoshida, T., Yo, M., Takahashi, R., Ohtawa, K., Arai, T., Takahashi, E., Miyoshi, H. and Miyawaki, A. (2013) ‘Visualizing developmentally programmed endoreplication in mammals using ubiquitin oscillators’, Development. 140:4624-4632.


 (5 votes)
(5 votes)