Featured resource: The National Avian Research Facility, UK
Posted by Lindsay Henderson, on 9 November 2021
Doing great science depends on teamwork, whether this is within the lab or in collaboration with other labs. However, sometimes the resources that support our work can be overlooked. In our new series, we aim to shine a light on these unsung heroes of the science world. The first article in the series is by Lindsay Henderson (NARF Academic Liaison and Post-Doctoral Scientist at Roslin Institute) who describes the work of the National Avian Research Facility, UK.
What is the NARF and where is it based?
The National Avian Research Facility (NARF) was founded in 2013 and is based at The Roslin Institute, on the University of Edinburgh’s Easter Bush campus, UK. The NARF produces and maintains poultry lines for use in scientific research and consists of two units, a conventional facility and a facility that has specified pathogen free (SPF) status.
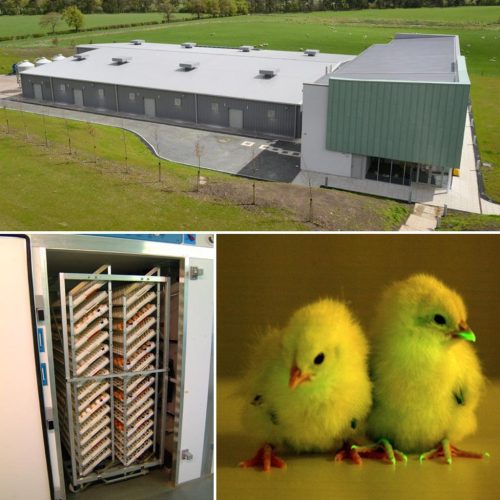

What services does the NARF offer for developmental biologists?
The NARF supplies fertile eggs from our range of chicken lines that have a broad utility in developmental biology. These include, commercial layer and broiler chicken lines, and Japanese quail.
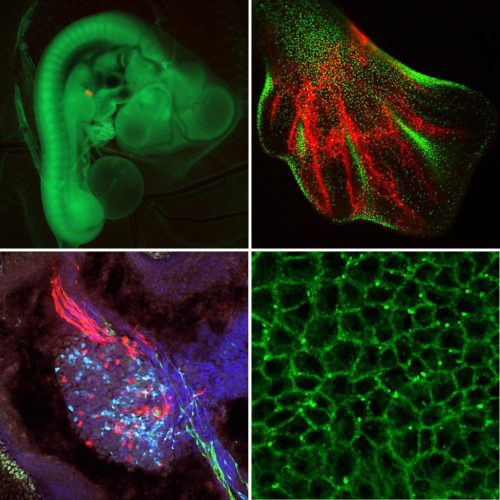
Researchers within the Roslin Institute Chicken Embryology (RICE) group have generated a number of genetically-altered (GA) fluorescent reporter chicken lines that are valuable tools for developmental biology2. These lines carry fluorescent reporter transgenes that are expressed ubiquitously or are inducible in cells of the developing embryo and are used for fate mapping, cell lineage tracing and tissue grafting. At this time the NARF has three ubiquitous reporter chicken lines; the Roslin Green (eGFP)3, Flamingo (TdTomato)4 and Membrane GFP5, and an inducible reporter chicken line, the Chameleon (Cytbow)2.
The NARF also maintains fluorescent reporter chicken lines that are gene specific. This includes a number of immune cell reporter lines, such as the CSF1R-reporter lines6, that can be used to visualise immune cells, including macrophages and dendritic cells.
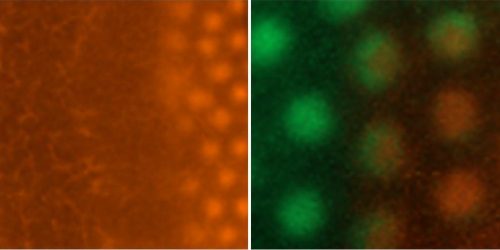

The NARF also holds a spontaneous mutation line, the talpid3 line that is a classical recessive embryonic lethal chicken mutant with abnormal limb patterning and malformations7–9. The talpid3 line has provided direct insights into clinical genetics and specifically the genes responsible for limb abnormalities and ciliopathies in humans9,10.
The NARF maintains one of the only SPF inbred lines, Line 0, that is free of Avian Sarcoma-Leukosis Virus (ASLV) ev loci. Embryos from this line may be particularly suitable for research using RCAS retrovirus vectors11.
Who are the developmental biologists based at the Roslin Institute that use the NARF?
RICE is a collective of research groups based at the Roslin Institute that use the chicken embryo to study vertebrate embryonic growth and development. RICE includes; the Balic Group that has produced the first transgenic chicken lines which allow specific sub-population of chicken immune cells to be visualised. The Davey Group examines the causative alterations of gene expression which lead to variations in phenotype using comparative anatomy, genomics and embryonic manipulation of avian species, with a particular interest in the molecular anatomy of the developing limb bud. The Headon Group focuses on the development, maintenance and repair of the skin and its appendages. The Rainger Group’s research is focused on the fusion of tissues in the developing retina using the chicken embryo as an experimental model system. The McGrew Group has made major advances in and continues to explore gene editing of avian germ cells and biobanking. The Clinton Group investigates the molecular control of sex determination and gonadal development, and the mechanisms underlying sexual dimorphisms in birds. The Sang Group has a broad focus on the development and application of technologies for genome engineering of the chicken.
What are the strengths of the chick model system in today’s developmental biology?
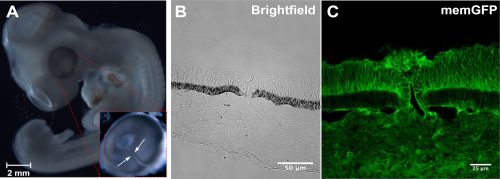
Recent advances in genome modification in the chicken led by Mike McGrew using the NARF facilities, have made possible the rapid and cost-effective production of GA chicken lines. In addition to new methods to create precise and targeted gene modifications, like CRISPR/Cas9 technologies, this greatly advances the potential of gene editing in chickens for developmental biologists. These advances in gene editing and transgenesis in conjunction with the experimental accessibility of chick embryos, and the ease of live culture and imaging, make the chick a powerful model for future developmental biology research.
Gene editing in chickens can be used to create targeted mutations, to knock-out genes of interest and enable inducible ablation of cell types that contain specific genes12. Recently, the NARF’s gene editing technologies have made possible research investigating avian sex determination using targeted mutations in the DMRT1 gene13. Gene editing in the chicken will soon be used to explore the genes responsible for defects of the optic fissure closure in the eye that cause ocular coloboma14.
How can researchers get involved with the NARF?
The NARF can supply fertile eggs for research and can supply a quote for eggs for inclusion in grant applications. Please find the full list of the lines we provide here. To order eggs from any of the poultry lines or to obtain costings for grant applications, contact narf@roslin.ed.ac.uk.
We continue to develop new GA chicken lines to aid in research under both conventional and SPF conditions. We welcome new collaborations with developmental biologists based at other research institutes. If you have any questions or are interested in a collaboration please contact Lindsay Henderson the NARF Academic Liaison.
The NARF is supported by the University of Edinburgh and the UKRI-Biotechnology and Biological Sciences Research Council.
Publications
- Macdonald, J. et al. Efficient genetic modification and germ-line transmission of primordial germ cells using piggyBac and Tol2 transposons. Proc. Natl. Acad. Sci. U. S. A. 109, 8803 (2012).
- Davey, M. G., Balic, A., Rainger, J., Sang, H. M. & McGrew, M. J. Illuminating the chicken model through genetic modification. Int. J. Dev. Biol. 62, 257–264 (2018).
- McGrew, M. J. et al. Localised axial progenitor cell populations in the avian tail bud are not committed to a posterior Hox identity. Development 135, 2289–2299 (2008).
- Ho, W. K. W. et al. Feather arrays are patterned by interacting signalling and cell density waves. PLoS Biol. 17, (2019).
- Rozbicki, E. et al. Myosin-II-mediated cell shape changes and cell intercalation contribute to primitive streak formation. Nat. Cell Biol. 17, 397–408 (2015).
- Balic, A. et al. Visualisation of chicken macrophages using transgenic reporter genes: Insights into the development of the avian macrophage lineage. Development 141, 3255–3265 (2014).
- Ede, D. A. & Kelly, W. A. Developmental abnormalities in the head region of the talpid3 mutant of the fowl. J. Embryol. Exp. Morphol. 12, 161–182 (1964).
- Ede, D. A. & Kelly, W. A. Developmental abnormalities in the trunk and limbs of the talpid3 mutant of the fowl. J. Embryol. Exp. Morphol. 12, 339–356 (1964).
- Fraser, A. M. & Davey, M. G. TALPID3 in Joubert syndrome and related ciliopathy disorders. Current Opinion in Genetics and Development 56, 41–48 (2019).
- Davey, M. G., Towers, M., Vargesson, N. & Tickle, C. The chick limb: Embryology, genetics and teratology. Int. J. Dev. Biol. 62, 253–263 (2018).
- McNally, M. M., Wahlin, K. J. & Canto-Soler, M. V. Endogenous expression of ASLV viral proteins in specific pathogen free chicken embryos: Relevance for the developmental biology research field. BMC Dev. Biol. 10, 106 (2010).
- Ballantyne, M. et al. Direct allele introgression into pure chicken breeds using Sire Dam Surrogate (SDS) mating. Nat. Commun. 12, 659 (2021).
- Ioannidis, J. et al. Primary sex determination in birds depends on DMRT1 dosage, but gonadal sex does not determine adult secondary sex characteristics. Proc. Natl. Acad. Sci. 118, e2020909118 (2021).
- Rainger, J. Novel approaches to define tissue fusion mechanisms in embryonic development. Funder: UK Research and Innovation Available at: https://gtr.ukri.org/projects?ref=MR%2FS033165%2F1.


 (No Ratings Yet)
(No Ratings Yet)