In Development this week (Vol. 139, Issue 18)
Posted by Seema Grewal, on 21 August 2012
Here are the highlights from the current issue of Development:
Retinoic acid’s developmental role illuminated
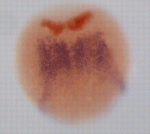
The role of all-trans retinoic acid (RA) in hindbrain development and other developmental processes is usually studied by blocking endogenous RA synthesis and then continuously supplying exogenous RA by soaking embryos in all-trans RA or by implanting RA-soaked beads. Now, on p. 3355, David Bensimon and colleagues use photo-isomerisation of 13-cis RA to all-trans RA and vice versa to manipulate RA activity spatiotemporally in zebrafish embryos. In embryos in which all-trans RA synthesis is impaired, they report, brief incubation in all-trans RA or in 13-cis RA followed by UV illumination before the bud stage rescues hindbrain development. By contrast, rescue is impaired in embryos treated with all-trans RA and then exposed to UV light. Notably, activation of all-trans RA via photo-isomerisation of 13-cis RA at the end of gastrulation in head, but not tail, precursor cells rescues hindbrain development. These results suggest that all-trans RA is sequestered in embryos during normal development. Furthermore, they illustrate how RA activity can be spatiotemporally controlled in developing zebrafish embryos.
Branching out studies of dendritic patterning

The dendritic arbours of neurons in the central nervous system have highly diverse morphologies that determine neuronal connectivity and, consequently, brain function. But how is dendritic architecture sculpted during development? On p. 3442, Kazuto Fujishima, Mineko Kengaku and co-workers investigate this question by combining computer simulations and time-lapse imaging of cultured mouse cerebellar Purkinje cells. The researchers show that the characteristic Purkinje cell space-filling dendritic arbour of non-overlapping branchlets is shaped by several reproducible dynamic processes, including constant tip elongation, stochastic terminal branching and retraction triggered by contacts between growing dendrites. This last process, they report, is regulated by protein kinase D signalling. Their computer simulation of dendrite branch dynamics, which incorporates their experimental measurements, reproduces dendritic development in live Purkinje cells and confirms the important contribution that dendritic retraction makes to the formation of the Purkinje cell dendritic arbour. Further development of this two-pronged approach, suggest the researchers, will help to clarify the fundamental mechanisms of dendrite patterning in the developing brain.
Pancreatic development keeps its Sox on
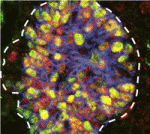
All mature pancreatic cell types arise from a pool of organ-specific multipotent progenitor cells. Cell-intrinsic and cell-extrinsic cues promote the proliferation and cell fate commitment of these progenitor cells but what integrates these cues during pancreatic morphogenesis? Maike Sander and co-workers now report that the transcription factor Sox9 forms the centrepiece of a gene regulatory network that controls pancreatic development (see p. 3363). Pancreatic progenitor-specific ablation of Sox9 during early mouse pancreatic development, they report, leads to cell-autonomous loss of fibroblast growth factor receptor 2b (Fgfr2b), which is required to transduce mesenchymal Fgf10 signals. In turn, Fgf10 is required to maintain progenitor expression of Sox9 and Fgfr2b. Perturbation of this Sox9/Fgfr2b/Fgf10 feed-forward expression loop results in pancreas-to-liver fate conversion. Given that Fgf signalling is necessary for pancreatic progenitor cell proliferation, the researchers propose that a Sox9/Fgf feed-forward loop coordinately controls organ fate commitment and progenitor cell expansion in the developing pancreas, a finding that may advance efforts to generate insulin-producing cells for therapeutic use.
COP1 sheds light on root development
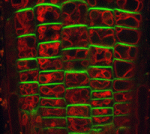
Although the roots of most plant species are not directly exposed to light, root growth is controlled by light and integrated with the growth of above-ground organs. Here (p. 3402), Teva Vernoux, Jian Xu and colleagues uncover a novel long-distance mechanism by which light signals through the master photomorphogenesis repressor COP1 to coordinate root and shoot development in Arabidopsis. The authors show that, in the shoot, COP1 regulates shoot-to-root transport of auxin by controlling transcription of the auxin efflux carrier gene PIN-FORMED1 (PIN1). In addition, they report, COP1 regulates auxin transport and cell proliferation in the root apical meristem by modulating the intracellular distribution of PIN1 and PIN2 in the root, thereby ensuring rapid and precise tuning of root growth to the light environment. Together, these results identify auxin as a long-distance signal in light-regulated plant development and show how spatially separated control mechanisms can converge on a single signalling system to coordinate development at the whole plant level.
Pten-less neuroblasts stop en route
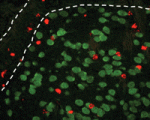
Neuronal precursors in the subventricular zone (SVZ) of the adult rodent brain differentiate into neuroblasts and migrate through the rostral migratory stream (RMS) to the olfactory bulb, where they differentiate into interneurons. Diverse extracellular cues control neuroblast migration but what are the intracellular pathways that respond to these cues? On p. 3422, Suzanne Baker and colleagues identify a role for Pten, a negative regulator of phosphoinositide 3-kinase (PI3K) signalling, in mouse neuroblast development. The PI3K-Akt-mTorc1 pathway is inactivated in migrating neuroblasts in the SVZ and RMS, they report, but activated when the cells reach the olfactory bulb. Postnatal deletion of Pten, they show, causes aberrant activation of PI3K-Akt-mTorc1 signalling, premature termination of neuroblast migration, neuroblast differentiation and enlargement of the SVZ and RMS. Notably, live imaging of slice cultures shows that, although some Pten-null neuroblasts lack directional migration and have a non-polarised morphology, others migrate normally towards the olfactory bulb. Thus, the researchers suggest, the neuroblast migration defect associated with Pten loss is secondary to precocious neuroblast differentiation.
Haploid ESCs: a future in the germline
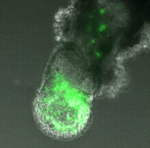
Parthenogenetic haploid embryonic stems cells (ESCs), which have recently been established through chemical activation of unfertilised mouse cells, give rise to a wide range of differentiated cell types in embryos and in vitro. But can they contribute to the germline, which is the defining hallmark of mouse diploid ESCs? Here (p. 3301), Martin Leeb, Anton Wutz and co-workers show that parthenogenetic haploid mouse ESCs have a robust germline potential and that transgenic mouse strains can be produced from genetically modified haploid ESCs. Differentiation of haploid ESCs in chimeric embryos, they report, correlates with the acquisition of a diploid karyotype by the ESCs through endoreduplication. By contrast, haploid ESCs induced to differentiate to an extra-embryonic fate by expression of the transcription factor Gata6 retain their haploid content. Parthenogenetic haploid mouse ESCs, the researchers conclude, are authentic pluripotent ESCs that can, potentially, be used to elucidate new details about developmental pathways through extension of genetic screens and manipulations directly into mouse models.
Plus…
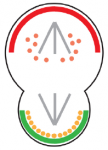
 Left-right patterning: conserved and divergent mechanisms
Left-right patterning: conserved and divergent mechanisms
This Development At a Glance poster by Nakamura and Hamada summarizes the common and divergent mechanisms by which LR asymmetry is established in vertebrates.
 Review: Principles and roles of mRNA localization in animal development
Review: Principles and roles of mRNA localization in animal development
Intracellular targeting of mRNAs, which has long been recognised as a means to produce proteins locally, has recently emerged as a mechanism used to polarise various cell types. Here, Florence Besse and colleagues review the regulation and functions of RNA localisation during animal development. See Review on p. 3263
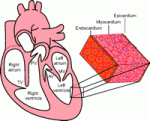 Review: Partitioning the heart: mechanisms of cardiac septation and valve development
Review: Partitioning the heart: mechanisms of cardiac septation and valve development
On p. 3277, Zhou, Chang, and colleagues review the morphogenetic events and genetic networks that regulate spatiotemporal interactions between the cells that partition the heart.


 (No Ratings Yet)
(No Ratings Yet)