Layered patterning systems in hair development
Posted by jamesglover1, on 28 September 2017
The story behind our recent paper ‘Hierarchical patterning modes orchestrate hair follicle morphogenesis‘ , finding that distinct patterning mechanisms can co-exist during embryonic organ formation.
From the spots of a leopard and stripes on a zebra to the pigmentation of sea shells and arrangement of sand dunes in a desert, repeating patterns are present at vastly different scales throughout the natural world. During embryonic development, repeating patterns are also prevalent and lead to diverse structures such as the digits of the limb, intestinal villi, cartilaginous rings of the trachea and rugae of the palate. But how do these patterns form in the first place?
A common feature with repeating patterns is that they arise through a process of self-organisation from local interactions within the system. The ability to self-organize is nowhere more apparent than in an embryo, where a single fertilised cell can develop into a complex organism with many different structures. To the mathematician and famous war-time code breaker Alan Turing, this emergence of a body plan (morphogenesis) was an example of symmetry breaking; where homogenous or ‘symmetric’ tissue, in terms of it being developmentally equivalent, transitions from a uniform to an organised heterogeneous state. This phenomenon inspired Turing to publish his seminal 1952 work ‘The chemical basis of morphogenesis’[1]. In his paper, Turing theorised that a pair of interacting chemical molecules (which he termed ‘morphogens’) that are both diffusible but at different rates, when acting across an entire field with local fluctuations can break symmetry and produce a regularly spaced pattern. This concept is now commonly referred to as the Turing reaction-diffusion model.
Some 20 years later two German scientists, Hans Meinhardt and Alfred Gierer, provided a general mechanism for such pattern formation [2]. Meinhardt and Gierer showed that a system only requires a network where a self-enhancing molecule with a short diffusible range known as the ‘activator’ stimulates the production of its own ‘inhibitor’ molecule that operates over a larger spatial range. Simply put, for a pattern to form, there needs only be local self-enhancing activation coupled with long-range inhibition.
Although this Meinhardt-Gierer model originally described biological molecules, the fundamental interactions of the network can explain the formation of non-biological patterns. Indeed, a nice example to introduce their model is during the formation of sand dunes. In this system, grains of sand are blown across a desert by the prevailing wind. Tiny random fluctuations in the system (such as a small rock making a bump) will cause sand grains from the wind to be deposited and soon a small mound will begin to form. This deposition of sand is the activator. Now as the wind continues to blow, this new mound will trap more sand and consequently will increase in size. This can be thought of as local self-activation. However, this increasing deposition of sand at the growing mound means that the number of sand grains remaining immediately downwind is vastly depleted. As a result, no sand is deposited in the area immediately behind the mound, generating a space until the next dune can form. This is the long-range inhibition in the system.
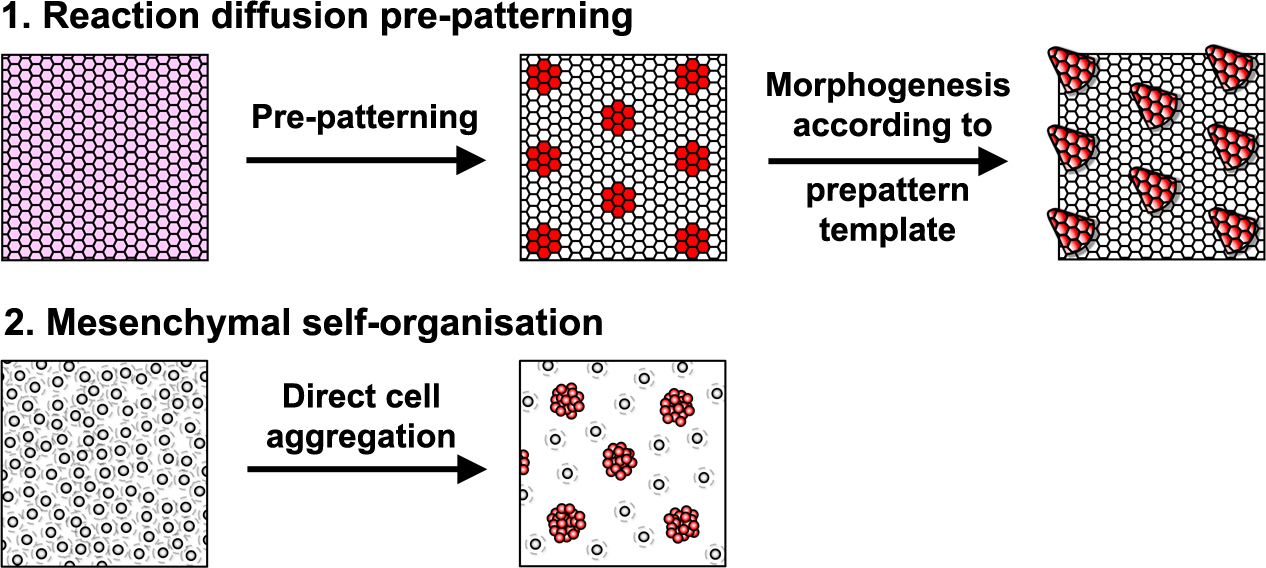
Reaction-diffusion systems are signal driven processes based on diffusing elements (e.g extracellular proteins, such as growth factors) that lead to a spatially patterned change in cell state usually resulting in altered gene expression. This arrangement of cell states then acts as a template or ‘prepattern’ from which anatomical structures are produced by inducing local cell aggregation, growth or survival (Fig. 1). However, importantly, these models are not the only type of mechanism that can produce pattern. A second class of model relies upon the ability of cells to self-organise directly, requiring no prepattern. In these systems periodic focal points of high cell density are created through chemotaxis, mechanical deformation of the environment, or cell-cell adhesion (Fig. 1). Although no prepattern is required, this type of pattern formation shares the same constraints as the first class of reaction-diffusion systems (local activation coupled with long range inhibition), but in these systems local cell clustering provides the activation whereas the inhibition results from depletion of cells around the emerging aggregates. Thus, these two classes of patterning mechanism are distinguished by the entity that moves to break symmetry – whether diffusible signals (reaction-diffusion) or cells (mesenchymal self-organisation).
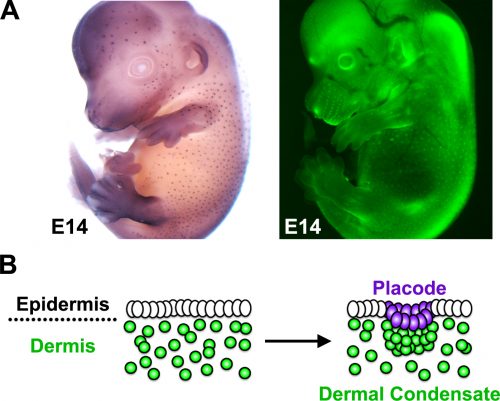
The developing skin is a good model in which to study periodic pattern formation as it is structurally quite simple and readily produces easily appreciated repeating patterns. At embryonic day 13, mouse skin is composed simply of an epithelial sheet (epidermis) lying atop a loose mesenchyme (dermis). As far as we can tell no pre-programmed information relevant to defining hair follicle positions is in there at the start and so the tissue can be deemed symmetric at this stage. Yet, only a day later (either in vivo or in culture) the skin has partitioned itself into a dotted pattern of altered gene expression and cell aggregations that form the hair follicle primordia (Fig. 2). This model system allows us to investigate pattern formation in a tightly framed way as we do not need to worry about the (admittedly interesting) earlier problem of how the skin develops to that stage, nor the (also interesting) question of how the primordia go on to make fully functional hair follicles and the intervening space becomes mature skin. So how is the symmetry of the homogeneous tissue broken to give a repeating spatial pattern of cell clusters and divergent cell fates? More specifically, instead of concerning ourselves with determining how the pattern produces an exact density and size of spots, we wanted to understand how the tissue symmetry is broken to give a spatially ordered arrangement.
A more complete reaction-diffusion system
As several signalling pathways have been shown to be essential for early hair primordium formation, we were interested in the integration between them and whether this integration might constitute a pattern forming reaction-diffusion system. We restricted our consideration to molecules that are members of three major pathways (BMP, FGF and WNT) known to be involved in hair follicle development [4-6], and to those mRNAs that have short half-lives and so can undergo rapid regulation at the timescale of the primary hair pattern formation. We assessed the transcriptional regulatory interactions between these molecules to define the network of interactions between them. To test whether our experimentally derived network was capable of creating a periodic pattern we sought the expertise of Dr Vaclav Klika (Czech Technical University, Prague). Vaclav’s mathematical analysis showed that this multiple species reaction-diffusion system is capable of breaking symmetry to produce a periodic pattern. From this it is plausible that interactions between WNT, FGF and BMP pathways are sufficient to generate the hair follicle pattern.
More than signals alone
Because complete hair follicle development also requires the formation of dermal condensates that underlie each placode, we wanted to explore how these mesenchymal cells aggregate and how this is regulated by the pathways and molecules we identified in our gene regulatory network. To answer this question we enlisted the help of Dr Richard Mort (based at the University of Lancaster) to track dermal cell movement during condensate formation. Using live cell imaging we found that the mesenchymal cells that form the dermal condensate are those located in its immediate vicinity, suggesting that local cues arising during patterning guide the dermal cells. As it was likely that any such signals would come from the epidermis we examined the timing between the epidermal signal driven pattern and corresponding dermal cell rearrangement. By analysing the gene expression of early placode markers and comparing this with the cellular organisation we found the existence of a molecular prepattern in the epidermis that precedes condensate formation. Further investigation revealed that this prepattern provides a template of local FGF sources that attract dermal cells ultimately leading to condensate formation.
Making everything a hair follicle
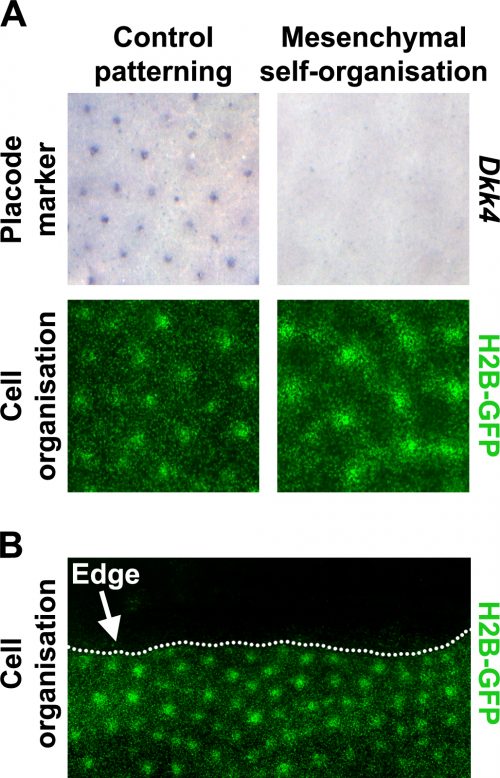
Prior literature indicated that hair follicle primordia are distinguished by their high FGF and low BMP activity [5, 7]. We wondered what would happen if we imposed these conditions across the entire skin. Using a pharmacological inhibitor of BMP signalling and a recombinant FGF protein, we treated unpatterned skins to achieve this effect. The resulting pattern of the skins cultured in these conditions looked rather similar to control skins, based on dermal cell aggregation, but we found that expression of epidermal placode marker genes was absent in these skins (Fig. 3A). We realised that this condition had revealed a previously unrecognised developmental potential in the skin; that the dermis has the capability to pattern by itself without instruction from of an epidermal prepattern.
To analyse the formation of these mesenchyme-only patterns we enlisted the help of Dr Franziska Matthäus (FIAS in Frankfurt) who specializes in methods to analyse cell motility. Through particle image velocimetry (PIV) analysis we began to identify distinct differences, such as far greater cell movement and incorporation into condensates, between the mesenchyme-only and the unperturbed patterning processes.
Living on the edge
To work out whether the mesenchymal cell only patterns arise through a fundamentally different patterning system we used the fact that although the skin appears developmentally homogeneous, it is unavoidable that it has edges where dissected away from the embryo to put into culture. The behaviour of patterns at these tissue edges can be very informative when trying to distinguish the underlying mechanism of a pattern’s generation (Fig. 3B).
Patterning systems relying on active inhibitory signals will form a row of foci close to the edge as the inhibitor diffuses off the edge, thereby giving a competitive advantage to those cells at the perimeter. Conversely, in cell driven patterning systems the pattern will stay away from the edge as cell number becomes limiting for the nucleation of new cell aggregates, and at the edge there are simply fewer cells available to recruit. By inserting cuts into skins cultured in both control and in our conditions recreating the hair follicle primordium, we see that the mesenchymal only patterning is characteristic of a cell driven mechanism rather than a signal driven reaction-diffusion system as is observed for the control experiments. This suggested that these two systems rely on fundamentally different mechanisms.
We then searched for a molecular mechanism for mesenchyme-only patterning. We found that disruption of TGFβ signalling abolished mesenchyme-only patterning, but that these perturbations had only modest effects on normal patterning, which would fit the idea that these patterning processes occur through divergent mechanisms with a different reliance on TGFβ signalling. In addition, we found that local sources of TGFβ2 attract mouse dermal cells, revealing that TGFβ2 serves as a widely expressed attractant that draws mesenchymal cells together.
Finally, having determined that restricted TGFβ signalling was critical for mesenchyme-only patterning we wanted to determine its role in the interplay between normal and mesenchymal patterning mechanisms during hair development. We found that TGFβ2 treatment substantially enhances dermal cell attraction to FGF sources and that when TGFβ signalling is inhibited dermal cells migrate very poorly towards to local FGF sources. Thus, in normal development TGFβ signalling creates an environment conducive to cell recruitment to the local epidermal FGF sources. These sources, defined by the signal driven network as the prepattern, create restricted microenvironments that provide the conditions for mesenchyme-only patterning to occur such that a dermal condensate is formed. This demonstrates that fundamentally distinct patterning systems can operate together during embryonic organ formation, but in this case a hierarchy exists wherein one system guides the other.
Conclusions
Research into biological pattern formation has been enjoying a resurgence in recent years. Examples of Turing systems underpinning the formation of the limb digits [8], villi of the intestine [9] or the transition of colour spots on lizard skin [10] have been described. In addition to signal focused patterning mechanisms, during the weeks following the release of our work, two new papers [11, 12] have described cell processes driving chicken feather patterning and mouse hair follicle assembly, highlighting the ability of mesenchymal cells to self-organise. Studying the interplay between cell and signal driven processes during embryogenesis promises to be an exciting field of investigation in the future, which could provide fresh insight in the fundamental areas of developmental biology.
This work would not have been possible with the input of many collaborators. The mathematical guidance, simulations and models provided to us by Kevin Painter, Vaclav Klika and Franziska Matthäus has allowed us to confirm our preliminary laboratory findings and provide a more comprehensive answer to the initial question we asked. The imaging experiments and tools for cell tracking designed by Richard Mort enabled us to accurately follow dermal cells during hair follicle formation making it possible for us reject a hypothesis of pre-determined cell sorting, highlighting the utility of live cell imaging when studying developmental processes. Collaborations such as these, where scientists specialising in a range of different fields and from different locations work together on a single problem is the basis for outstanding research and I strongly encourage establishing similar relationships to help advance your own research.
This work was funded by the BBSRC and carried primarily in Dr Denis Headon’s group at The Roslin Institute near Edinburgh. You can discover the full story, findings and experiments in our paper:
Hierarchical patterning modes orchestrate hair follicle morphogenesis. J. D. Glover et al., PLOS Biology, 2017.15 (7):p.e2002117 http://journals.plos.org/plosbiology/article?id=10.1371/journal.pbio.2002117
James D. Glover
References
- Turing, A.M., The Chemical Basis of Morphogenesis. Philosophical Transactions of the Royal Society of London Series B-Biological Sciences., 1952. 237(641): p. 37-72.
- Gierer, A. and H. Meinhardt, A theory of biological pattern formation. Biological Cybernetics, 1972. 12(1): p. 30-39.
- Glover, J.D., et al., Hierarchical patterning modes orchestrate hair follicle morphogenesis. PLoS Biol, 2017. 15(7): p. e2002117.
- Mou, C., et al., Generation of the primary hair follicle pattern. Proc Natl Acad Sci U S A, 2006. 103(24): p. 9075-80.
- Huh, S.H., et al., Fgf20 governs formation of primary and secondary dermal condensations in developing hair follicles. Genes Dev, 2013. 27(4): p. 450-8.
- Andl, T., et al., WNT signals are required for the initiation of hair follicle development. Dev Cell, 2002. 2(5): p. 643-53.
- Botchkarev, V.A., et al., Noggin is a mesenchymally derived stimulator of hair-follicle induction. Nat Cell Biol, 1999. 1(3): p. 158-64.
- Raspopovic, J., et al., Modeling digits. Digit patterning is controlled by a Bmp-Sox9-Wnt Turing network modulated by morphogen gradients. Science, 2014. 345(6196): p. 566-70.
- Walton, K.D., et al., Villification in the mouse: Bmp signals control intestinal villus patterning. Development, 2016. 143(3): p. 427-36.
- Manukyan, L., et al., A living mesoscopic cellular automaton made of skin scales. Nature, 2017. 544(7649): p. 173-179.
- Shyer, A.E., et al., Emergent cellular self-organization and mechanosensation initiate follicle pattern in the avian skin. Science, 2017. 357(6353): p. 811-815.
- Lei, M., et al., Self-organization process in newborn skin organoid formation inspires strategy to restore hair regeneration of adult cells. Proceedings of the National Academy of Sciences, 2017. 114(34): p. E7101-E7110.


 (8 votes)
(8 votes)