October in preprints
Posted by the Node, on 2 November 2016
Our latest monthly trawl for developmental biology (and other cool) preprints. See June’s post for background, and let us know if we missed anything
This month features quite a bit of cell biology, both in early embryos and in a dish, with work on microtubules from the labs of Andreas Prokop, Tim Mitchison and Manuel Thery. We also found investigations into zebrafish cell migration and gastrulation, lots of new C. elegans work, a new view on superenhancers, some genomed Swedes, and some cats.
As in the last months, the preprints predominantly came from bioRxiv, but we also found some in PeerJ and arXiv. If we missed anything out, let us know. Happy preprinting!
Developmental biology
Collective epithelial migration drives timely morphogenesis of the retinal neuroepithelium. Jaydeep Sidhaye, Niklas Ifflaender, Caren Norden
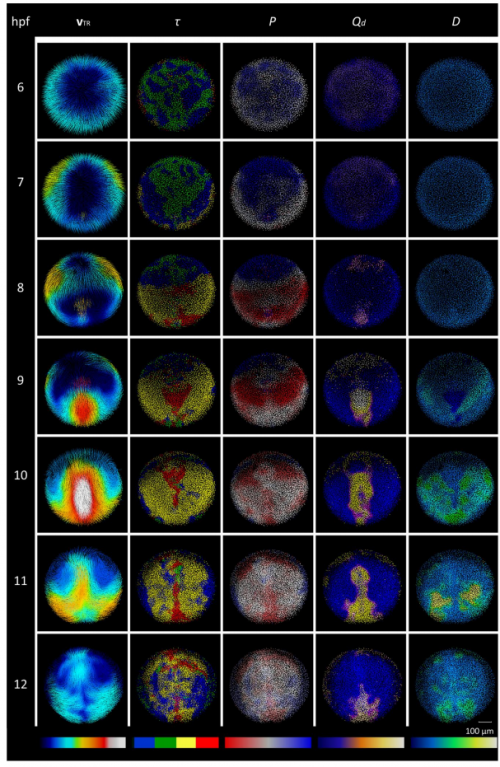
Kinematic analysis of cell lineage reveals coherent and robust mechanical deformation patterns in zebrafish gastrulation. David Pastor-Escuredo, Benoit Lombardot, Thierry Savy, Adeline Boyreau,Jose M. Goicolea, Andres Santos, Paul Bourgine, Juan C. del Alamo, Nadine Peyrieras, Maria J. Ledesma-Carbayo.
A microRNA family exerts maternal control on sex determination in C. elegans. Katherine McJunkin, Victor Ambros
Casein Kinase II is Required for Proper Cell Division and Acts as a Negative Regulator of Centrosome Duplication in C. elegans Embryos. Jeffrey C Medley, Megan M Kabara, Michael D Stubenvoll, Lauren E DeMeyer, Mi Hye Song
Sexually dimorphic differentiation of a C. elegans hub neuron is cell-autonomously controlled by a conserved transcription factor. Esther Serrano Saiz, Meital Oren-Suissa, Emily Ann Bayer, Oliver Hobert
The nematode homologue of Mediator complex subunit 28, F28F8.5, is a critical regulator of C. elegans development. Markéta Kostrouchová, David Kostrouch, Ahmed A Chughtai, Filip Kaššák, Jan P. Novotný, Veronika Kostrouchová, Aleš Benda, Michael W. Krause, Vladimír Saudek, Marta Kostrouchová, Zdenek Kostrouch
Transcriptomic Description of an Endogenous Female State in C. elegans. David Angeles-Albores, Daniel H. W. Leighton, Tiffany Tsou, Tiffany H Khaw, Igor Antoshechkin, Paul Sternberg
Mcmdc2, a minichromosome maintenance (MCM) paralog, is required for repair of meiotic DNA breaks and homolog pairing in mouse meiosis. Adrian J McNairn, Vera D Rinaldi, John C Schimenti
Dynamic changes in Sox2 spatio-temporal expression direct the second cell fate decision through Fgf4/Fgfr2 signaling in preimplantation mouse embryos. Tapan Kumar Mistri, Wibowo Arindrarto, Wei Ping Ng, Choayang Wang, Hiong Lim Leng, Lili Sun, Ian Chambers, Thorsten Wohland, Paul Robson
Mice deficient of Myc super-enhancer region reveal a differential control mechanism between normal and pathological growth. Kashyap Dave, Inderpreet Sur, Jian Yan, Jilin Zhang, Eevi Kaasinen, Fan Zhong, Leander Blaas, Xiaoze Li, Robert Mansson, Jussi Taipale
Neuropilin 1 sequestration by neuropathogenic mutant glycyl-tRNA synthetase is permissive to vascular development and homeostasis. James Nicholas Sleigh, Adriana Gomez-Martin, Giampietro Schiavo
A dual-strategy expression screen for candidate connectivity labels in the developing thalamus. Olivia Bibollet-Bahena, Tatsuya Okafuji, Karsten Hokamp, Kevin J. Mitchell
Oligodendrocytes in the mouse optic nerve originate in the preoptic area. Katsuhiko Ono, Kengo Yoshii, Hiroyuki Tominaga, Hitoshi Gotoh, Tadashi Nomura, Hirohide Takebayashi, Kazuhiro Ikenaka
High temporal resolution of gene expression dynamics in developing mouse embryonic stem cells. Brian S Gloss, Bethany Signal, Seth W Cheetham, Franziska Gruhl, Dominik Kaczorowski, Andrew C Perkins, Marcel E Dinger
Periodic actin structures in neuronal axons are required to maintain microtubules. Yue Qu, Ines Hahn, Stephen Webb, Simon P. Pearce, Andreas Prokop
A Novel function for Cactus/IκB inhibitor to promote Toll signals in the Drosophila embryo. Maira A Cardoso, Marcio R Fontenele, Bomyi Lim, Paulo M Bisch, Stanislav Shvartsman, Helena M Araujo
A shielded irradiation assay to investigate mechanisms of in vivo stem cell migration in planarians. Prasad Abnave, Ellen Aboukhatwa, Nobuyoshi Kosaka, James Thompson, Mark Hill, Aziz Aboobaker
Computational study of the impacts of mechanical and physical cell properties on mitotic cell rounding in developing epithelia. Ali Nematbakhsh, Wenzhao Sun, Pavel A Brodskiy, Aboutaleb Amiri, Cody Narciso, Zhiliang Xu, Jeremiah J Zartman, Mark S Alber
Exploratory bioinformatics analysis reveals importance of “junk” DNA in early embryo development. Steven Xijin Ge
Frequency of mosaicism points towards mutation-prone early cleavage cell divisions. Chad Harland, Carole Charlier, Latifa Karim, Nadine Cambisano, Manon Deckers, Erik Mullaart, Wouter Coppieters, Michel Georges
Single Cell Expression Data Reveal Human Genes that Escape X-Chromosome Inactivation. Kerem Wainer-Katsir, Michal Linial
Morphological plant modeling: Unleashing geometric and topologic potential within the plant sciences. Alexander Bucksch, Acheampong Atta-Boateng, Akomian Fortune Azihou, Mathilde Balduzzi, Dorjsuren Battogtokh, Aly Baumgartner, Brad Binder, Siobhan Braybrook, Cynthia Chang, Viktoriya Coneva, Thomas DeWitt, Alexander Fletcher, Malia Gehan, Diego Hernan Diaz Martinez, Lilan Hong, Anjali Iyer-Pascuzzi, Laura Klein, Samuel Leiboff, Mao Li, Jonathan Lynch, Alexis Maizel, Julin Maloof, RJ Cody Markelz, Ciera Martinez, Laura Miller, Washington Mio, Wojtek Palubicki, Hendrik Poorter, Christophe Pradal, Charles Price, Eetu Puttonen, John Reese, Ruben Rellan-Alvarez, Edgar Spalding, Erin Sparks, Chris Topp, Joseph Williams, Daniel Chitwood
Multi-Compartmental Biomaterial Scaffolds for Patterning Neural Tissue Organoids in Models of Neurodevelopment and Tissue Regeneration. Richard J. McMurtrey
Cell Biology
Physical basis of large microtubule aster growth. Keisuke Ishihara, Kirill S Korolev, Timothy J Mitchison
CENP-F couples cargo to growing and shortening microtubule ends. Gil Kanfer, Martin Peterka, Vladimir K. Arzhanik, Alexei L. Drobyshev, Fazly I. Ataullakhanov, Vladimir A. Volkov, Benoît Kornmann
Microtubule stabilization drives centrosome migration to initiate primary ciliogenesis. Amandine Pitaval, Fabrice Senger, Gaelle Letort, Xavier Gidrol, James Sillibourne, Manuel Thery
The duration of mitosis and daughter cell size are modulated by nutrients in budding yeast. Ricardo Mendes Leitao, Annie Pham, Douglas Kellogg
Crossover in the dynamics of cell wall growth controls Caulobacter division times. Shiladitya Banerjee, Klevin Lo, Alan Selewa, Thomas Kuntz, Matthew K Daddysman, Aaron R Dinner, Norbert F Scherer
Probing the effect of uniaxial compression on cell migration. Nishit Srivastava, Robert R Kay, Alexandre J Kabla
Disease models
Zika infection of neural progenitor cells perturbs transcription in neurodevelopmental pathways. Lynn Yi, Harold Pimentel, Lior Pachter
Single Cell Phenotyping Reveals Heterogeneity among Haematopoietic Stem Cells Following Infection. Adam L MacLean, Maia A Smith, Juliane Liepe, Aaron Sim, Reema Khorshed, Nico Scherf, Axel Krinner, Ingo Roeder, Cristina Lo Celso, Michael PH Stumpf
A Saposin deficiency model in Drosophila: lysosomal storage, progressive neurodegeneration, sensory physiological decline and defective calcium homeostasis. Samantha J Hindle, Sarita Hebbar, Dominik Schwudke, Christopher J Elliott, Sean T Sweeney
Distal axotomy enhances retrograde presynaptic excitability onto injured pyramidal neurons via trans-synaptic signaling. Tharkika Nagendran, Rylan Larsen, Rebecca Bigler, Shawn Frost, Benjamin Philpot, Randolph J. Nudo, Anne Marion Taylor
Resources & Techniques
Mocap: Large-scale inference of transcription factor binding sites from chromatin accessibility. Xi Chen, Bowen Yu, Nicholas Carriero, Claudio Silva, Richard Bonneau
Pooled CRISPR screening with single-cell transcriptome read-out. Paul Datlinger, Christian Schmidl, Andre F Rendeiro, Peter Traxler, Johanna Klughammer, Linda Schuster, Christoph Bock
Transcriptional landscape of cell lines and their tissues of origin. Camila Miranda Lopes-Ramos, Joseph N Paulson, Cho-Yi Chen, Marieke L Kuijjer, Maud Fagny, John Platig, Abhijeet R Sonawane, Dawn L DeMeo, John Quackenbush, Kimberly Glass
Whole genome resequencing of a laboratory-adapted Drosophila melanogaster population sample. William P. Gilks, Tanya M Pennell, Ilona Flis, Matthew T Webster, Edward H. Morrow
SweGen: A whole-genome map of genetic variability in a cross-section of the Swedish population. Adam Ameur, Johan Dahlberg, Pall Olason, Francesco Vezzi, Robert Karlsson, Par Lundin, Huiwen Che, Jessada Thutkawkorapin, Andreas Kusalananda Kahari, Mats Dahlberg, Johan Viklund, Jonas Hagberg, Niclas Jareborg, Inger Jonasson, Asa Johansson, Sverker Lundin, Daniel Nilsson, Bjorn Nystedt, Patrik Magnusson, Ulf Gyllensten
Semi-automated genome annotation using epigenomic data and Segway. Eric G. Roberts, Mickaël Mendez, Coby Viner, Mehran Karimzadeh, Rachel C. W. Chan,Rachel Ancar, Davide Chicco, Jay R. Hesselberth,Anshul Kundaje, Michael M. Hoffman
SCORPIUS improves trajectory inference and identifies novel modules in dendritic cell development. Robrecht Cannoodt, Wouter Saelens, Dorine Sichien, Simon Tavernier, Sophie Janssens, Martin Guilliams, Bart N Lambrecht, Katleen De Preter, Yvan Saeys.
Development of a modular automated system for maintenance and differentiation of adherent human pluripotent stem cells. Duncan E Crombie, Maciej Daniszewski, Helena H Liang, Tejal Kulkarni, Fan Li, Grace E Lidgerwood, Alison Conquest, Damian Hernandez, Sandy S Hung, Katherine P Gill, Elisabeth De Smit, Lisa S Kearns, Linda Clarke, Valentin M Sluch, Xitiz Chamling, Donald J Zack, Raymond CB Wong, Alex Hewitt, Alice Pebay
Evolution etc
Recombination Suppression is Unlikely to Contribute to Speciation in Sympatric Heliconius Butterflies. John W. Davey, Sarah L. Barker, Pasi M. Rastas, Ana Pinharanda, Simon H. Martin, Richard Durbin, Richard M. Merrill, Chris D. Jiggins
Evolutionary determinants of morphological polymorphism in colonial animals. Carl Simpson, Jeremy B.C. Jackson, Amalia Herrera-Cubilla
Cytoplasmic-nuclear incompatibility between wild-isolates of Caenorhabditis nouraguensis. Piero Lamelza, Michael Ailion
Transition from environmental to partial genetic sex determination in Daphnia through the evolution of a female-determining incipient W-chromosome. (Revision) Celine M. O. Reisser, Dominique Fasel, Evelin Hurlimann, Marinela Dukic, Cathy Haag-Liautard, Virginie Thuillier, Yan Galimov, Christoph R Haag
The genome of the crustacean Parhyale hawaiensis: a model for animal development, regeneration, immunity and lignocellulose digestion. (Revision) Damian Kao, Alvina G Lai, Evangelia Stamataki, Silvana Rosic, Nikolaos Konstantinides, Erin Jarvis, Alessia Di Donfrancesco, Natalia Pouchkina-Stantcheva, Marie Semon, Marco Grillo, Heather Bruce, Suyash Kumar, Igor Siwanowicz, Andy Le, Andrew Lemire, Michael Eisen, Cassandra Extavour, William Browne, Carsten Wolff, Michalis Averof, Nipam H Patel, Peter Sarkies, Anastasios Pavlopoulos, Aziz Aboobaker
Spatiotemporal patterns of desiccation tolerance in natural populations of Drosophila melanogaster. Subhash Rajpurohit, Eran Gefen, Alan Bergland, Dmitri Petrov, Allen Gibbs, Paul Schmidt
Evolving Notch polyQ tracts reveal possible solenoid interference elements. Albert J Erives
Pre-metazoan origin of animal miRNAs. (Revision) Jon Brate, Ralf Stefan Neumann, Bastian Fromm, Arthur Alexander Blorstad Haraldsen, Paul Grini, Kamran Shalchian-Tabrizi
Challenges and solutions for transcriptome assembly in non-model organisms with an application to hybrid specimens. Arnaud Ungaro, Nicolas Pech, Jean-Francois Martin, Scott RJ McCairns, Remi Chappaz, Andre Gilles
Why not…
Of cats and men: the paleogenetic history of the dispersal of cats in the ancient world. Claudio Ottoni, Wim van Neer, Bea De Cupere, Julien Daligault, Silvia Guimaraes, Joris Peters, Nikolai Spassov, Mary E. Pendergast, Nicole Boivin, Arturo Morales-Muniz, Adrian Balasescu, Cornelia Becker, Norbert Benecke, Adina Boronenant, Hijlke Buitenhuis, Jwana Chahoud, Alison Crowther, Laura Llorente, Nina Manaseryan, Herve Monchot, Vedat Onar, Marta Osypinska, Olivier Putelat, Jacqueline Studer, Ursula Wierer, Ronny Decorte, Thierry Grange, Eva-Maria Geigl


 (1 votes)
(1 votes)