September in preprints
Posted by the Node, on 5 October 2016
Our monthly trawl for preprints from developmental biology and related research. Previous posts can be found here.
Preprints roll on, and this month’s selection is a reminder of how developmental biology is based on a variety of model organisms. We’ve got two investigations into zebrafish mesoderm from Deborah Yelon’s lab, quite a bit of cell mechanics (in zebrafish, fly and worm), updates on the underlying cell biology of cell division in c. elegans, a transcriptome for the spiny mouse, and much more besides.
As in the last months, the preprints predominantly came from bioRxiv, but we also found a couple from PeerJ. If we missed anything out, let us know. Happy preprinting!
Developmental biology
PDGF signaling directs cardiomyocyte movement toward the midline during heart tube assembly. Joshua Bloomekatz, Reena Singh, Owen W.J. Prall, Ariel C. Dunn, Megan Vaughan, Chin-San Loo, Richard P. Harvey, Deborah Yelon
Hand2 Inhibits Kidney Specification While Promoting Vein Formation Within the Posterior Mesoderm. Elliot A. Perens, Zayra V. Garavito-Aguilar, Gina P. Guio-Vega, Karen T. Pena, Yocheved L. Schindler, Deborah Yelon
Kinematic analysis of cell lineage reveals coherent and robust mechanical deformation patterns in zebrafish gastrulation. David Pastor-Escuredo, Benoit Lombardot, Thierry Savy, Adeline Boyreau, Jose M. Goicolea, Andres Santos, Paul Bourgine, Juan C. del Alamo, VNadine Peyrieras, Maria J. Ledesma-Carbayo
Zfp423 regulates Sonic hedgehog signaling via primary cilium function. Chen-Jei Hong, Bruce A. Hamilton
Heterozygous mutation to Chd8 causes macrocephaly and widespread alteration of neurodevelopmental transcriptional networks in mouse. Andrea L Gompers, Linda Su-Feher, Jacob Ellegood, Tyler W Stradleigh, Iva Zdilar, Nycole A Copping, Michael C Pride, Melanie D Schaffler, M Asrafuzzaman Riyadh, Gaurav Kaushik, Brandon Mannion, Ingrid Plajzer-Frick, Veena Afzal, Axel Visel, Len A Pennacchio, Diane Dickel, Jason P Lerch, Jacqueline N Crawley, Konstantinos S Zarbalis, Jill L Silverman, Alex S Nord
Landscape of X chromosome inactivation across human tissues. Taru Tukiainen, Alexandra-Chloé Villani, Angela Yen, Manuel A. Rivas, Jamie L. Marshall, Rahul Satija, Matt Aguirre, Laura Gauthier, Mark Fleharty, Andrew Kirby, Beryl B. Cummings, Stephane E. Castel, Konrad J. Karczewski, François Aguet, Andrea Byrnes, GTEx Consortium, Tuuli Lappalainen, Aviv Regev, Kristin G. Ardlie, Nir Hacohen, Daniel G. MacArthur
Genome-wide binding of posterior HOXA/D transcription factors reveals subgrouping and association with CTCF. Ivana Jerković, Daniel Murad Ibrahim, Guillaume Andrey, Stefan Haas, Peter Hansen, Catrin Janetzki, Irene González Navarrete, Peter Nick Robinson, Jochen Hecht, Stefan Mundlos
Differential gene expression in blastocysts following pronuclear transfer. Edward H Morrow, Fiona C Ingleby
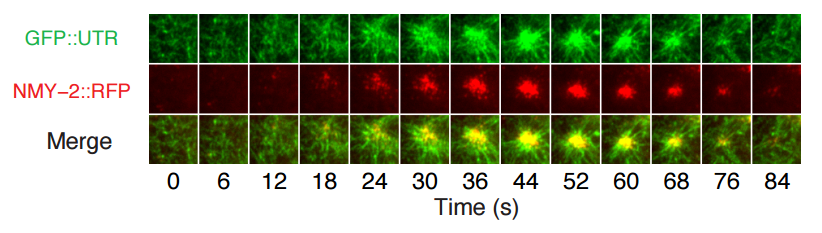
Excitable RhoA dynamics drive pulsed contractions in the early C. elegans embryo. François B Robin, Jonathan B Michaux, William M McFadden, Edwin M Munro
ATX-2, the C. elegans Ortholog of Human Ataxin-2, Regulates Centrosome Size and Microtubule Dynamics. Michael Stubenvoll, Jeffrey Medley, Miranda Irwin, Mi Hye Song
Synaptonemal complex components are required for meiotic checkpoint function in C. elegans. Tisha Bohr, Guinevere Ashley, Evan Eggleston, Kyra Firestone, Needhi Bhalla
Temporal regulation of epithelium formation. Stephen E Von Stetina, Jennifer J. Liang, Georgios Marnellos, Susan E Mango
Myosin II controls junction fluctuations to guide epithelial tissue ordering. Scott Curran, Charlotte Strandkvist, Jasper Bathmann, Marc de Gennes, Alexandre Kabla, Guillaume Salbreux, Buzz Baum
Geometry can provide long-range mechanical guidance for embryogenesis. Mahamar Dicko, Pierre Saramito, Jocelyn Etienne
Identification and Functional Characterization of Muscle Satellite Cells in Drosophila. Rajesh D. Gunage, Heinrich Reichert, K. VijayRaghavan
Translational control of gurken mRNA in Drosophila development. Christopher J Derrick, Timothy T Weil
Lipid metabolic perturbation is an early-onset phenotype in adult spin mutants: a Drosophila model for lysosomal storage disorders. Sarita hebbar, Avinash Khandelwal, Jayashree R, Samantha J Hindle, Yin Ning Chiang, Joanne Yew, Sean Sweeney, Dominik Schwudke
Regions of very low H3K27me3 partition the Drosophila genome into topological domains. Sherif El-Sharnouby, Bettina Fischer, Jose Paolo Magbanua, Benjamin Umans, Rosalyn Flower, Siew Woh Choo, Steven Russell, Rob White
Sexual dimorphism in the Drosophila metabolome increases throughout development. Fiona C Ingleby, Edward H Morrow
Cell Biology
Diverse roles of guanine nucleotide exchange factors in regulating collective cell migration. Assaf M Zaritsky, Yun-Yu Tseng, Angeles M. Rabadan, Michael Overholtzer, Gaudenz Danuser, Alan Hall
Laurdan and di-4-ANEPPDHQ probe different properties of the membrane. Mariana Amaro, Francesco Reina, Martin Hof, Christian Eggeling, Erdinc Sezgin
Heterogeneity of single-cell mechanical responses to tumorigenic factors. Aldo Leal-Egana, Gaelle Letort, Jean-Louis Martiel, Andreas Christ, Timothee Vignaud, Caroline Roelants, Odile Filhol, Manuel Thery
The FACT complex and cell cycle progression are essential to maintain asymmetric transcription factor partitioning during cell division. Eva Herrero, Sonia Stinus, Eleanor Bellows, Peter H Thorpe
Ankyrin G membrane partners drive the establishment and maintenance of the axon initial segment. Christophe Leterrier, Nadine Clerc, Fanny Rueda-Boroni, Audrey Montersino, Bénédicte Dargent, Francis Castets
Evolution etc.
Nuclear pore-like structures in a compartmentalized bacterium. Evgeny Sagulenko, Amanda Nouwens, Richard I Webb, Kathryn Green, Benjamin Yee, Gary Morgan, Andrew Leis, Kuo-Chang Lee, Margaret K Butler, Nicholas Chia, Uyen Thi Phuong Pham, Stinus Lindgreen, Ryan Catchpole, Anthony M Poole, John A. Fuerst
A structural variant in the 5′-flanking region of the TWIST2 gene affects melanocyte development in belted cattle. Niveditha Awasthi Mishra, Cord Drögemüller, Vidhya Jagannathan, Rémy Bruggmann, Julia Beck, Ekkehard Schütz, Bertram Brenig, Steffi Demmel, Simon Moser, Heidi Signer-Hasler, Aldona Pieńkowska-Schelling, Claude Schelling, Ronald Rongen, Stefan Rieder, Robert N. Kelsh, Nadia Mercader, Tosso Leeb
Contrasting genome dynamics between domesticated and wild yeasts. Jia-Xing Yue, Jing Li, Louise Aigrain, Johan Hallin, Karl Persson, Karen Oliver, Anders Bergström, Paul Coupland, Jonas Warringer, View ORCID ProfileMarco Cosentino Lagomarsino, Gilles Fischer, Richard Durbin, Gianni Liti
The genetic basis and fitness consequences of sperm midpiece size in deer mice. Heidi S Fisher, Emily Jacobs-Palmer, Jean-Marc Lassance, Hopi E Hoekstra
Mito-nuclear Incompatibilities and Mitochondrial Replacement Therapy. Adam Eyre-Walker
De novo transcriptome assembly for the spiny mouse (Acomys cahirinus). Jared Mamrot, Roxane Legaie, Stacey J Ellery, Trevor Wilson, David Gardner, David W Walker, Peter Temple-Smith, Anthony T Papenfuss, Hayley Dickinson
Indomethacin reproducibly induces metamorphosis in Cassiopea xamachana scyphistomae. Patricia Cabrales-Arellano, Tania Islas-Flores, Patricia E. Thomé, Marco A. Villanueva
Tools, techniques, resources, modelling
Zipper plot: visualizing transcriptional activity of genomic regions. Francisco Avila Cobos, Jasper Anckaert, Pieter-Jan Volders, Dries Rombaut, Jo Vandesompele, Katleen De Preter, Pieter Mestdagh
Mapping protein function with CRISPR/Cas9-mediated mutagenesis. Katherine F Donovan, Mudra Hegde, Meagan Sullender, Emma W Vaimberg, Cory M Johannessen, David E Root, View ORCID ProfileJohn G Doench
Glutton: large-scale integration of non-model organism transcriptome data for comparative analysis. Alan Medlar, Laura Laakso, Andreia Miraldo, Ari Löytynoja
Nanopore DNA Sequencing and Genome Assembly on the International Space Station. Sarah L Castro-Wallace, Charles Y Chiu, Kristen K John, Sarah E Stahl, Kathleen H Rubins, Alexa B. R. McIntyre, Jason P Dworkin, Mark L Lupisella, David J Smith, Douglas J Botkin, Timothy A Stephenson, Sissel Juul, Daniel J Turner, Fernando Izquierdo, Scot Federman, Doug Stryke, Sneha Somasekar, Noah Alexander, Guixia Yu, Christopher Mason, Aaron S Burton
PCR artifact in testing for homologous recombination in genomic editing in zebrafish. Minho Won, Igor B Dawid
Integrating tissue specific mechanisms into GWAS summary results. Alvaro Barbeira, Scott P Dickinson, Jason M Torres, Eric S Torstenson, Jiamao Zheng, Heather E Wheeler, Kaanan P Shah, Todd Edwards, GTEx Consortium, Dan Nicolae, Nancy J Cox, Hae Kyung Im
Comparing individual-based approaches to modelling the self-organization of multicellular tissues. James M. Osborne, Alexander G. Fletcher, Joseph M. Pitt-Francis, Philip K. Maini, David J. Gavaghan
The Drosophila Gene Expression Tool (DGET) for expression analyses. Yanhui Hu, Aram Comjean, Norbert Perrimon, Stephanie Mohr
Noise-processing by signaling networks. Styliani Kontogeorgaki, Ruben Sanchez-Garcia, Rob Ewing, Konstantinos Zygalakis, Ben D. MacArthur
Robust cell tracking in epithelial tissues through identification of maximum common subgraphs. Jochen Kursawe, Rémi Bardenet, Jeremiah J Zartman, Ruth E Baker, Alexander George Fletcher


 (1 votes)
(1 votes)