Shaping Snapdragons
Posted by XanaRebocho, on 4 April 2017
Have you ever wondered what makes the shapes in the animal and plant kingdom so different? We take for granted the diversity of natural shapes that surround us, from a simple pine leaf to complex orchid flower. However, they pose one of the most beautiful scientific challenges. For centuries, scientists have been fascinated by how a certain shape is generated and what drives such diversity of shapes. I am no different! From my university days, I always wanted to study evolutionary development, to go into the depth of why and what lies behind the diversity of shapes in nature.
After my PhD days using genetics, molecular biology and comparative development as tools to study the evolution of shape in plants, I was keen to explore the upcoming computational modeling field as a tool to tackle complex development problems. This is where this article’s journey started, when I joined the lab of Enrico (Rico) Coen at the John Innes Centre to study the evolution and development of the complex 3D Snapdragon flower shape. I still remember my first contact with modeling. I was immediately hooked by the colorful shapes in the computer screen which looked so much like a snapdragon flower. However, nothing could have prepared me for the thousand lines of computational code that sustained the generation of the virtual Snapdragon model. At that moment I thought to myself, maybe I have bitten more than I can chew by wanting to do both the biology and the computational modeling.
The personal story that goes with the published article is a roller coaster of wrong hypothesis, failed and impossible experiments, and frustrating models but also glimpses of successes. You might be thinking, oh no this is going to be a tale of woe but let me disclose the end already – this is no sad story, this is the actual necessary evolution of thought, knowledge and personal growth that sustains any challenging scientific problem. For my feeling the story behind this paper feels more like a Tolkien novel with lots of downs, that you almost don’t expect to overcome, but also amazing ups.
Our approach at the start of my post-doc was to look into the working hypothesis generated by a previous in-house paper where the growth of the Snapdragon petals was analyzed and a virtual flower corolla was produced. The main assumptions underlying this model related to the spatiotemporal changes in the growth pattern due a local cell polarity inversion event combined with a local boost in growth rate.
To visualize the inversion of polarity in cells we decide to use an accepted marker of plant cell polarity that generally shows an asymmetric cellular localization – the PIN1 protein. For two long years, I struggled with PIN1 antibodies that didn’t work or produced slight promising subcellular signals that held us in the false hope of results. After changing the antibody company and spreading the risks by designing several different epitopes to the different PIN1s, I finally had a good working antibody that produced the expected pattern of cellular polarity in emerging organs. Armed with this tool, I initiated a marathon of immunolocalization sections across the petals at different stages of development to show the local inversion of cellular polarity. From the beginning it was obvious that something special was happening to the expression of the PIN1 in the predicted region at the predicted time by the model. However, after a year of thorough sectioning experiments and analysis, I was struggling to reconstruct the 3D PIN1 pattern from the 2D sections. At that point we wanted to start writing the paper to show the peculiar upregulation of PIN1 in the predicted region by the model but unfortunately the precise pattern of the PIN1 was still undetermined and led to a lot of team discussions and wild ideas on how to distil the result. Emotions were running high, as we were failing to grasp the final proof of the inversion of polarity. After we decided to put the paper on hold and re-think our approach, I got desperate! In other models system and exposed organs such as roots, the first solution would be to do whole-mount immunolocalization, where the whole organ is hybridized and the 3D patterns can be imaged. However, our tissue of interested was the inner surface of tiny petals, in buds as small as 500 microns. Desperate times require desperate measures, and after a bad night of sleep and probably a delirious morning, I started dissecting incredibly tiny petals and sticking them to a glass slide using prosthetic glue. Then, to make sure the antibody could penetrate the tissue, I boiled these fragile tissues cringing at every little water bubble brushing through the stuck petals. To check the cell/tissue integrity and in anticipation of no PIN1 signal, I also stained the petal tissues with a cellulose fluorescent stain called calcofluor. After three agonizing days of hoping that the tissue was not turned into a pulp, I descended the stairs that separate our lab from the confocal microscopy room with a heavy heart; this is our last crazy chance! If this doesn’t work, I am out of ideas. As I started up the confocal and focused on the first set of 5 petals, I couldn’t believe what I was seeing, a beautiful ladder of PIN1 signal in the regions of interest -I couldn’t contain my joy.
I picked up the phone and called some lab mates to be sure they could see what I was seeing (I couldn’t exclude delirious mirages at this point with my nerves on the fringe). After they confirmed what I was seeing and shared a happy dance with me, I went to get Rico from his office luring him with promises of something special without giving too much away. We still have photos of this moment that we shared with the four of us in a confocal room. There are moments that you don’t forget in your life and this one is engrained in my memory.
I finally could extract full epidermal patterns of PIN1 in wild-type and mutant petals at different stages of development, and guess what? I never saw an inversion or convergence of polarity in the region predicted by the model. I did see an upregulation of PIN1 at the predicted time and space but the pattern of PIN1 was mainly proximodistal (along one axis). This might feel like a bitter sweet result as it contradicted our model but actually this was quite an unexpected and exciting result. In my opinion, one of the beautiful givens in science is that it will surprise you. You make a hypothesis, your results prove it wrong and you need to re-evaluate your thinking. This was indeed what happened to us but as a result we realized that we didn’t need an inversion of polarity, as the key information required for growth lay in the orientation of the arrows (axiality), not in the direction in which they are pointing. So we came up with an alternative model that still involved a switch in growth orientations but without a polarity inversion (I can’t imagine I just summarized months of modeling with a sentence but there you have it!).
We then tested this model by correlating cell wall patterns with growth orientations and showing the switch in growth orientation at the predicted time and space.
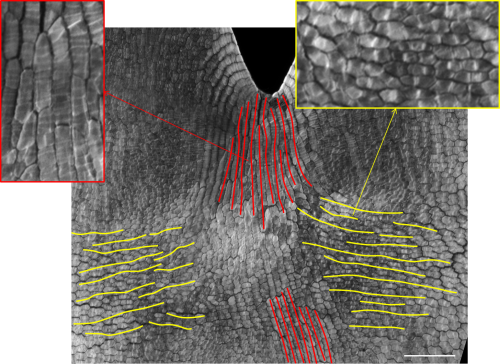
We were finally ready to write our results, publish our new model and make a contribution to the scientific understanding of 3D morphogenesis. However, one of our biggest challenges was still to surface – conveying the ideas developed over the years in words, while connecting the static 2D images of cell polarity and growth orientations with the 3D flower shape. When you read the paper now, the logic seems to fall into place, with the increasing complexity of the models and the experimental data driving a new model hypothesis. But the reality was a long winding road, with several versions of the paper; many rejected by ourselves, others by our critical colleagues, and others by peer-reviewed journals. This was a very frustrating time for the team and I am not really sure when we started to see the light, as this was an incremental and slow process, but at a certain point we felt that we were finally are on track. By deconstructing the model complexity (from square piece of tissue to wedge model to full corolla model and to mutant models) and conveying the idea of how differential growth behaviors can lead to tissue conflict resolutions, we were finally bringing the disjointed paper together. Our paper distilled three tissue conflict resolution behaviors leading to an out-of-plane deformation: surface conflict (differential growth/contraction rates between two surfaces), areal conflict (differential growth/contraction rates across a tissue) and directional conflict (differential orientation growth/contraction across a tissue). These three behaviors are the basis of many of the described morphogenetic events in plants and animals, and can be combined to produce the panoply of 3D shape complexity that surrounds us.
For me this journey made me a more complete scientist. I have explored new disciplines and learned to think in a different way. I have developed extra respect for the power of modeling and became humbler when it comes to tackling complex biological problems. Lastly, I realized that the essence of a piece of research lies in the evolution of thought of the researchers, their failures and frustrations, their stubbornness and their hunches, their commitment and their successes, and that is what makes science so amazing.
If you want to read the paper that resulted from this emotional rollercoaster – https://elifesciences.org/content/6/e20156.


 (8 votes)
(8 votes)