Borders and communities: solving old puzzles with new tools
Posted by David Wilkinson, on 10 May 2018
An important question in developmental biology is how regions with distinct identity are established despite the intermingling of cells that occurs during growth and morphogenesis. Our recent work revisited some old studies of how the vertebrate hindbrain is patterned, and found that sharp and homogeneous segments are formed through a combination of identity switching and border control.
The story started in the late 1980s, in the early days of analysing developmental gene expression using in situ hybridisation. One of the genes we analysed was egr2 (a.k.a. krox20), a transcription factor which had been identified by Patrick Charnay as an early growth response gene in fibroblasts. To our amazement, we found that egr2 is expressed in stripes in the hindbrain, corresponding to two rhombomeres, r3 and r5. We then collaborated with Robb Krumlauf to show that hox genes have segmental expression in the hindbrain. egr2 and hox genes were later found to be components of a network that regulates segmental identity. A striking feature of their segmental expression is that they come to have razor sharp borders, and a clue to how these form came from the work of Scott Fraser and colleagues. They found that once morphological borders are seen in the chick hindbrain, cells do not intermingle between segments.
In another collaboration with the Charnay lab, we carried out a screen to identify kinases that are segmentally expressed in the hindbrain. One of these, a receptor tyrosine kinase subsequently named EphA4, is expressed in r3 and r5, and we later found that it is a direct transcriptional target of egr2. We went on to show that Eph receptors and their ephrin ligands underlie cell segregation that sharpens the segment borders. This turned out to be the first example of a general role of Ephs and ephrins in border formation during development.
These findings fit the familiar idea that cell segregation sharpens and stabilises tissue organisation. However, the lineage studies in chick had found that cells marked at early stages can contribute progeny to adjacent hindbrain segments. Furthermore, experiments by Trainor and Krumlauf in mouse, and by Schilling, Prince and Ingham in zebrafish, had shown that cells transplanted between segments change identity to match their new neighbours. Intriguingly, identity switching occurs for single cells but not when groups of cells are transplanted. These findings in the early 2000s suggested that some intermingling occurs, and identity switching ensures that segments nevertheless establish a homogeneous identity. However, this idea languished as intermingling between segments had not been directly visualised, and the mechanism of switching remained a mystery. This was the problem that Megan Addison took on as her PhD project in my lab.
We reasoned that intermingling and identity switching of cells would mainly occur at early stages, when egr2 is first expressed but EphA4 has yet to be upregulated to sufficient levels to drive cell segregation. The key question is whether any egr2-expressing cells intermingle from r3 and r5 into adjacent segments and then downregulate egr2 expression. To address this question, we used the newly-emerging techniques for genome manipulation in zebrafish to create an enhancer trap in which a stable reporter is expressed directly from the egr2 locus. During this work, another lab reported that intermingling does not occur between hindbrain segments in zebrafish, but used reporters expressed one step downstream of egr2, which might miss the time window in which mixing occurs. Indeed, using the early reporter line that we created we found that cell intermingling and identity switching does occur.
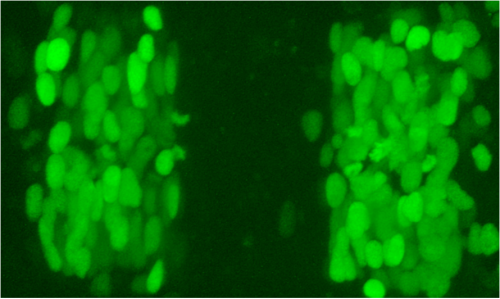
We started wondering what the mechanism of switching might be, and here some other old findings came into play. The Charnay lab had reported in 2001 that mosaic ectopic expression of egr2 in the chick hindbrain causes adjacent cells to upregulate egr2. This suggested that egr2 regulates a community effect, which in classical models would involve upregulation of a signal that non-autonomously induces egr2. Such community signaling leads to homogeneous gene expression within a field of cells, and might explain why groups of transplanted cells do not switch identity. However, the puzzle of how egr2 induces egr2 in adjacent cells had also languished in the literature.
Megan analysed whether ectopically expressed egr2 acts non-autonomously in the zebrafish hindbrain. We found that it does when the egr2-expressing cells have a scattered distribution, but not when they later segregate from cells with even-numbered identity. Since our previous work had shown that the segregation is driven by EphA4, we blocked it by simultaneous knockdown of EphA4 and found that non-autonomous induction was restored. Non-autonomous induction thus depends upon how many neighbours of the same or different type you have: egr2 is only induced in cells that are surrounded by egr2-expressing cells.
What might the mechanism of the community regulation of egr2 be, and does it account for identity switching? We started thinking about retinoic acid (RA) signaling as a candidate. An RA gradient establishes segmental identity in the hindbrain, and studies of Tom Schilling in zebrafish had shown that graded expression of an RA-degrading enzyme, cyp26a1, has a key role. The lab of Cecilia Moens found that two other family members, cyp26b1 and cyp26c1, have dynamic segmental expression that also contributes to A-P patterning. We wondered if this segmental expression is under the control of segment identity genes and thus acts in a feedback loop. We found that this is indeed the case: egr2 underlies the lower expression level of cyp26b1 and cyp26c1 in r3 and r5 compared with r2, r4 and r6. Since a high level of cyp26 enzymes can non-autonomously decrease RA levels in neighbouring cells, they could decrease RA signaling in single cells that have intermingled. Indeed, loss of cyp26b1 and cyp26c1 function disrupted the identity switching of egr2-expressing cells that have intermingled into adjacent segments. We showed that in r4 the switching involves upregulation of hoxb1, which in turn represses egr2 expression.
This work has revealed parallel mechanisms of identity switching and border control that establish sharp and homogeneous segments in the hindbrain. At early stages, some cells mix into adjacent segments and switch identity to match their new neighbours. This mediated by a community effect in which there is reciprocal feedback between RA levels and segmental identity. Subsequently, Eph receptors and ephrins are upregulated and they underlie segregation that prevents intermingling and sharpens the border.
These studies of hindbrain patterning raise the question of whether similar mechanisms operate in other tissues. Since mediators of cell segregation are often regulated downstream of regional identity genes, some intermingling between adjacent regions may occur at early stages. Furthermore, plasticity in cell identity is a common feature at early stages of development. Insights will come from the creation of further reporter lines to visualise cell intermingling and cell identity.
Cell Identity Switching Regulated by Retinoic Acid Signaling Maintains Homogeneous Segments in the Hindbrain. Megan Addison, Qiling Xu, Jordi Cayuso, David G. Wilkinson. Developmental Cell. DOI: https://doi.org/10.1016/j.devcel.2018.04.003


 (2 votes)
(2 votes)