The evolution from mouse to human models of lung development
Posted by markonikolic, on 4 September 2017
The story behind our eLife paper: Nikolić, M. Z., Caritg, O., Jeng, Q., Johnson, J.-A., Sun, D., Howell, K. J., Brady, J. L., Laresgoiti, U., Allen, G., Butler, R., Zilbauer, M., Giangreco, A., Rawlins E.L. (2017). Human embryonic lung epithelial tips are multipotent progenitors that can be expanded in vitro as long-term self-renewing organoids. Elife 6, e26575.
As a medical doctor specialising in respiratory diseases, with limited experience in real lab-based research, I was disappointed how little could be done for patients with end-stage lung disease (ESLD). Although respiratory diseases account for the third highest mortality of non-infectious disease deaths worldwide and lung cancer is the most common cancer (World Health Organization, 2011), treatment is largely supportive and lung tissue destruction is irreversible.
I started my PhD with Emma Rawlins at the Gurdon Institute, University of Cambridge, as part of the Wellcome PhD Programme for Clinicians by looking at human induced pluripotent stem cells (hiPSCs). These seemed to provide an opportunity to apply the discoveries of developmental biology to treat disease (Takahashi and Yamanaka, 2006). Differentiating patient-specific iPSCs into any relevant cell type could potentially be used for disease modelling and drug screening, and also bring us a step closer to cell-based therapies including personalised organogenesis. Research on murine lung development as a model for human lung development has been crucial to current protocols for differentiating human iPSCs into lung (Firth et al., 2014; Firth et al., 2015; Ghaedi et al., 2013; Huang et al., 2015; Huang et al., 2013; Longmire et al., 2012; Mou et al., 2012; Wong et al., 2012; Wong et al., 2015). I initially started work on iPSC directed differentiation to lung and wanted to use human embryonic lungs simply for validation purposes. However, it became evident very quickly that little developmental biology had been done on human embryonic lungs and it was not clear how closely the published step-wise iPSC differentiation protocols actually reflect human embryonic and foetal lung development.
Characterising the different progenitor populations in both the developing mouse and human embryonic lung seemed to me an essential part of regenerative medicine. The molecular understanding of developing human lungs was inadequate, as human lung development had not been characterised using cells derived from the human embryonic lung itself, but only using iPS-derived lung cells.
Mouse lung development
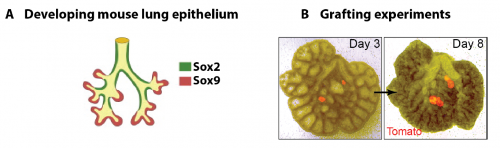
In parallel to my human work, I also worked on mouse lung development since it has been so extensively used as a model for human lung development. Considering the various anatomical and molecular differences already known between mouse and human lungs, I felt it was important to understand both mouse and human lung development more fully in order to ascertain to what extent the mouse findings could indeed be applied to human lungs. In both species, the lung develops as a blind-ending tube which branches multiple times, a process which results in the production of gas-delivering bronchioles and gas-exchanging alveoli. We know from lineage tracing experiments that the most important epithelial progenitor population in the developing mouse lung is found in the branching tips, giving rise to all the various epithelial cell types in the adult lung, both bronchiolar and alveolar cells (Rawlins et al., 2009). In mice, these epithelial tip cells are classically Sox9+ and initially give rise to the neighbouring Sox2+ stalk cells which are bronchiolar progenitors and give rise to the future airway (Fig.1 A). Using technically challenging grafting experiments I showed, among other things, that extrinsic signalling is the key in determining the tip and stalk progenitor cell fate, but that a significant degree of plasticity is present and that this gradually decreases throughout development (Fig.1 B) (Laresgoiti et al., 2016). I got a lot of practice in dissecting the epithelium free from any mesenchyme, which was very useful later on for the human work (Fig. 3B).
The finding that the environment is crucial in determining distal tip progenitor fate suggested that it would be possible to define in vitro culture conditions for human embryonic lung epithelium, both for the self-renewal of tip cells and their differentiation towards alveolar and bronchiolar lineages. This was an exciting realization as successful identification of culture conditions for developing human lungs would make molecular genetics possible.
Main challenges of working on human embryonic lungs
One of the main limitations of working with human tissue is the lack of continuous tissue supply. This changed quickly and unexpectedly when I went to one of the Cambridge pubs to discuss my research plans and get some advice from a post-doctoral friend. He mentioned the “TransEuro” trial going on at the University of Cambridge – a clinical trial in which brain tissue from terminations of pregnancies is used for neurosurgical transplantation into treatment-resistant Parkinson’s patients. This meant that developing lung tissue was regularly discarded without any scientific benefit. This seemed to be the ideal source of human lung tissue for validation of my attempts at iPS-derived directed differentiation, but it quickly became clear what a huge potential an in-depth molecular characterisation of human embryonic lungs would have for the entire lung developmental and stem cell biology community. Additionally, the Joint MRC/Wellcome Human Developmental Biology Resource (www.HDBR.org) coordinated tissue harvesting from all over the UK via London and Newcastle. This meant that we had lung tissue provided to us multiple times per month from all over the country. The developmental age ranged from 5 to 20 weeks’ developmental age (7-22 weeks’ gestational age) and the samples used had no known genetic abnormalities.
Another challenge was that tissue used for sequencing and in vitro culture had to be as fresh as possible. Initially, we did not want to take any chances and dissection usually occurred as soon as possible after tissue was obtained, which was often late in the day. Experiments frequently continued until the early hours of the morning. I must say I was grateful once it became clear that overnight storage in specifically designed embryonic tissue storage medium did not affect gene expression, reproducibility and viability!
Since this was an exciting new area with huge potential, I was very grateful for the help I received from two amazing undergraduate/masters students, Oriol Caritg and Dawei Sun, who assisted me with my human work, as well as another PhD student, Jo Johnson.
Main achievements
Molecular comparison of human and mouse embryonic lungs
Molecular characterisation of human embryonic and foetal lungs using immunostaining, qRT-PCR and RNAseq transcriptome analysis revealed both major similarities and differences between mouse and human lung epithelial progenitors (Nikolić et al., 2017). Interestingly, 96% of orthologous genes that were expressed in human tips were also present in mouse.
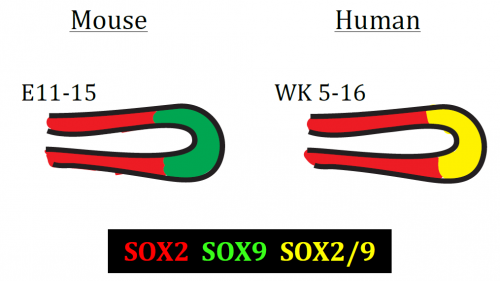
The most important difference was the discovery that the classical stalk marker SOX2 is expressed in the human SOX9+ distal tip progenitors (Fig. 2), in stark contrast to the expression pattern in mice. We showed that this is a true interspecies difference and cannot be explained by relative age. This is an exciting finding, since Sox9 has been used as a classical mouse tip marker and Sox2 as a classical mouse stalk marker, with a clear demarcation between them (Chang et al., 2013; Liu and Hogan, 2002). My work suggests that SOX2 cannot be used as a unique stalk marker in human embryonic lungs and for validation during iPS-derived directed differentiation. Furthermore, there were more subtle differences in tip signalling pathways between mouse and human (Figure 3). The same signalling pathways were present, but these were wired in fundamentally different ways. For example, BMP2 and BMP7 were highly enriched in human tips, whereas Bmp4 was enriched in the mouse (Bellusci et al., 1996); IHH was expressed in human lungs whereas mouse lungs express Shh (Bellusci et al., 1997).
On the one hand these data provide support for the use of mouse as a model for human lung development, but on the other hand they question its applicability in view of the many genetic differences, the functional consequences of which are not yet known. Therefore, to study the functional consequences of manipulating these genes, a genetically modifiable system for studying human lung development is required.
Self-renewing 3D organoid culture of human tip progenitors
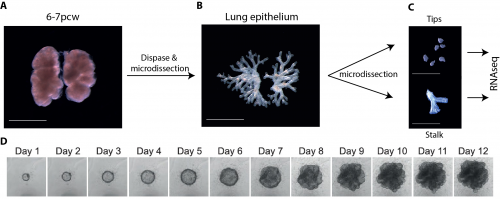
We therefore developed culture conditions which have the ability to expand human embryonic lung epithelial stem cells as an indefinitely self-renewing, expanding population of undifferentiated cells with a organoid forming efficiency of 100% (Figure 3). The concept is similar to capturing the inner cells mass of a blastocyst as a self-renewing embryonic stem cell line. Our lung organoids could also be frozen and thawed, and the technical advice obtained from Meritxell Huch’s lab within the Gurdon Institute was immensely helpful. This now allows the in vitro study of human embryonic lung development without the need to obtain fresh tissue for every experiment.
However, in mouse, the same culture conditions were not sufficient to maintain mouse lung epithelial progenitor cells after several passages. This suggests that acquiring a self-renewing state is possible with further optimisation of the culture conditions, but the required signalling mechanisms are likely to be different. This was not surprising based on the transcriptome comparison between mouse and human tip progenitors.
RNAseq of cultured tips and stalks further validated my self-renewing culture conditions, since key genes are shared by cultured tip, cultured stalk and fresh tip cells without expression of differentiation markers. This suggests that the in vitro conditions mimic the in vivo conditions very closely.
Furthermore, single cell efficiency was high at about 35% and unpublished results showed that we were able to genetically manipulate human epithelial tip progenitors using CRISPR-Cas9. Knocking out SOX9 led to loss-of-self-renewing state phenotype, showing that SOX9 expression is essential for organoid maintenance. What this means is that we can now study human lung progenitor self-renewal in vitro in a genetically modifiable system which is extremely efficient without introducing chromosomal abnormalities, and which reproduces what is happening in vivo. This opens up a whole new research field of studying the mechanism of self-renewal and differentiation.
In vivo and in vitro differentiation as proof for a self-renewing state
One of the most important proofs that we had actually achieved a self-renewing state was the ability to differentiate our expanded organoid cells towards both bronchiolar and alveolar lineages. If real stem cells do it, then our expanded ones should do it do. Indeed, human embryonic epithelial lung progenitors, which had been expanded as 3D organoids, could be differentiated in vitro towards both alveolar and bronchiolar lineages. Similarly, xenotransplantation of organoid cells mixed with E13.5 dissociated mouse lungs, into the mouse kidney capsule led to efficient bronchiolar differentiation and the appearance of ciliated cells, mucus cells and basal cells, although alveolar differentiation was much less efficient. This strongly supported our hypothesis that a self-renewing state had been achieved.
We have also performed xenotransplantation into bleomycin-injured adult mouse lungs which showed highly efficient integration and proliferation of intratracheally administered single organoid cells. It also showed that the organoid cells can differentiate into bronchiolar cells, suggesting that lung identity is retained within the organoids. Such efficient engraftment has not been achieved previously. This is an exciting new model with many future possibilities.
Impact and future directions
In conclusion, the platform that we have developed for in vitro analysis of the embryonic human lung will be the first system available for functional experiments on human embryonic lungs. It will allow human embryonic lung development to be studied using modern molecular genetic techniques. The fact that we can genetically modify our organoid system also means that we can use CRISPR-Cas9 to identify novel genes required for alveolar and bronchiolar differentiation. This has been done in pancreatic development using iPS-derived cells (McGrath et al., 2015). However, our approach would be more reliable, as we would be doing this with real lung progenitors, albeit expanded ones, rather than iPS-derived ones. This research has the long-term potential to be transformative for lung regeneration by disease modelling, specifically by determining the role of genetic variants associated with lung diseases, by developing improved therapies for premature neonatal lung maturation, and also for in vitro differentiation of iPS cells. It has been claimed that iPSCs offer unparalleled opportunities to model lung development and disease using human cells. I would argue that our culture system using real human embryonic lung cells provides a more authentic and reliable platform to model lung development and disease.
From a clinical point of view, I hope that the work will make a difference to at least three groups of patients:
- Patients with end-stage lung disease: our work will guide iPS-derived directed differentiation, which could then be used to recellularise a decellularised lung or lobar scaffold to be used for lung transplantation without the need for immunosuppression.
- Premature neonates with lung failure due to immaturity: the 3D organoid culture system we developed will allow the study of late foetal development using real lung progenitors rather than iPS-derived ones. Understanding how the lung matures in these final stages of lung development will be crucial to improve survival for babies born prematurely.
- Patients with rare congenital lung conditions. Gene editing in our 3D culture system will enable disease modelling of conditions such as surfactant protein B and C deficiencies.
Publishing first and fast versus publishing “big”
Every scientist would like their work to be recognised as important and groundbreaking by those in their field. However, as far as career progression is concerned, the journal chosen often appears to be far more important than the actual quality of the work. It seems to me that journal impact factor is used as a substitute for quality by those who are not experts in a particular field, or who do not understand the impact of the work. In my experience, important factors in choosing a journal include the speed of the review process, the availability of person-power to do the revision work, the awareness of competitors, and related guesswork of what stage those competitors have reached and whether their work is yet of a standard suitable for publication. During our paper revision period, I was in full-time clinical training and I was reliant on the goodwill of my colleagues for the revision experiments, especially Oriol Caritg and Quitz Jeng. On reviewing all these aspects, we chose a journal known for its efficient and fair review process rather than focusing simply on journal impact factor. Put differently, we chose to publish first and fast, rather than attempting to publish “big”. I remain hopeful that our work (as well as that of others) will be judged purely by its contents and the impact it has on the field, rather than by the impact factor of the journal in which it was published.
Marko Nikolić
Wellcome Trust/CRUK Gurdon Institute & Wellcome Trust/MRC Stem Cell Institute,
University of Cambridge, Cambridge, UK
REFERENCES
Bellusci, S., Furuta, Y., Rush, M. G., Henderson, R., Winnier, G. and Hogan, B. L. (1997). Involvement of Sonic hedgehog (Shh) in mouse embryonic lung growth and morphogenesis. Development 124, 53–63.
Bellusci, S., Henderson, R., Winnier, G., Oikawa, T. and Hogan, B. L. (1996). Evidence from normal expression and targeted misexpression that bone morphogenetic protein (Bmp-4) plays a role in mouse embryonic lung morphogenesis. Development 122, 1693–1702.
Chang, D. R., Martinez Alanis, D., Miller, R. K., Ji, H., Akiyama, H., McCrea, P. D. and Chen, J. (2013). Lung epithelial branching program antagonizes alveolar differentiation. Proc. Natl. Acad. Sci. U.S.A. 110, 18042–18051.
Firth, A. L., Dargitz, C. T., Qualls, S. J., Menon, T., Wright, R., Singer, O., Gage, F. H., Khanna, A. and Verma, I. M. (2014). Generation of multiciliated cells in functional airway epithelia from human induced pluripotent stem cells. Proc. Natl. Acad. Sci. U.S.A.
Firth, A. L., Menon, T., Parker, G. S., Qualls, S. J., Lewis, B. M., Ke, E., Dargitz, C. T., Wright, R., Khanna, A., Gage, F. H., et al. (2015). Functional Gene Correction for Cystic Fibrosis in Lung Epithelial Cells Generated from Patient iPSCs. Cell Rep 12, 1385–1390.
Ghaedi, M., Calle, E. A., Mendez, J. J., Gard, A. L., Balestrini, J., Booth, A., Bove, P. F., Gui, L., White, E. S. and Niklason, L. E. (2013). Human iPS cell-derived alveolar epithelium repopulates lung extracellular matrix. J. Clin. Invest. 123, 4950–4962.
Huang, S. X. L., Green, M. D., de Carvalho, A. T., Mumau, M., Chen, Y.-W., d’Souza, S. L. and Snoeck, H.-W. (2015). The in vitro generation of lung and airway progenitor cells from human pluripotent stem cells. Nat Protoc 10, 413–425.
Huang, S. X. L., Islam, M. N., O’Neill, J., Hu, Z., Yang, Y.-G., Chen, Y.-W., Mumau, M., Green, M. D., Vunjak-Novakovic, G., Bhattacharya, J., et al. (2013). Efficient generation of lung and airway epithelial cells from human pluripotent stem cells. Nat. Biotechnol.
Laresgoiti, U., Nikolić, M. Z., Rao, C., Brady, J. L., Richardson, R. V., Batchen, E. J., Chapman, K. E. and Rawlins, E. L. (2016). Lung epithelial tip progenitors integrate Glucocorticoid and STAT3-mediated signals to control progeny fate. Development 143, 3686–3699.
Liu, Y. and Hogan, B. L. M. (2002). Differential gene expression in the distal tip endoderm of the embryonic mouse lung. Gene Expr. Patterns 2, 229–233.
Longmire, T. A., Ikonomou, L., Hawkins, F., Christodoulou, C., Cao, Y., Jean, J. C., Kwok, L. W., Mou, H., Rajagopal, J., Shen, S. S., et al. (2012). Efficient derivation of purified lung and thyroid progenitors from embryonic stem cells. Cell Stem Cell 10, 398–411.
McGrath, P. S., Watson, C. L., Ingram, C., Helmrath, M. A. and Wells, J. M. (2015). The Basic Helix-Loop-Helix Transcription Factor NEUROG3 Is Required for Development of the Human Endocrine Pancreas. Diabetes 64, 2497–2505.
Mou, H., Zhao, R., Sherwood, R., Ahfeldt, T., Lapey, A., Wain, J., Sicilian, L., Izvolsky, K., Musunuru, K., Cowan, C., et al. (2012). Generation of multipotent lung and airway progenitors from mouse ESCs and patient-specific cystic fibrosis iPSCs. Cell Stem Cell 10, 385–397.
Nikolić, M. Z., Caritg, O., Jeng, Q., Johnson, J.-A., Sun, D., Howell, K. J., Brady, J. L., Laresgoiti, U., Allen, G., Butler, R., et al. (2017). Human embryonic lung epithelial tips are multipotent progenitors that can be expanded in vitro as long-term self-renewing organoids. Elife 6, e26575.
Rawlins, E. L., Clark, C. P., Xue, Y. and Hogan, B. L. M. (2009). The Id2+ distal tip lung epithelium contains individual multipotent embryonic progenitor cells. Development 136, 3741–3745.
Takahashi, K. and Yamanaka, S. (2006). Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676.
Wang, Y., Tian, Y., Morley, M. P., Lu, M. M., Demayo, F. J., Olson, E. N. and Morrisey, E. E. (2013). Development and regeneration of Sox2+ endoderm progenitors are regulated by a Hdac1/2-Bmp4/Rb1 regulatory pathway. Dev. Cell 24, 345–358.
Wong, A. P., Bear, C. E., Chin, S., Pasceri, P., Thompson, T. O., Huan, L.-J., Ratjen, F., Ellis, J. and Rossant, J. (2012). Directed differentiation of human pluripotent stem cells into mature airway epithelia expressing functional CFTR protein. Nat. Biotechnol. 30, 876–882.
Wong, A. P., Chin, S., Xia, S., Garner, J., Bear, C. E. and Rossant, J. (2015). Efficient generation of functional CFTR-expressing airway epithelial cells from human pluripotent stem cells. Nat Protoc 10, 363–381.
World Health Organization (2011). Global Status Report on Noncommunicable Diseases 2010.


 (1 votes)
(1 votes)