Notch awakens: transitioning to the first differentiation step
Posted by Sergio Menchero, on 3 May 2019
The story behind our recent paper in eLife.
In the mid-1900s, Conrad Hal Waddington introduced the idea of development as a series of branching decisions taken under the control of genes1. In mammals, the first of these decisions takes place before the implantation of the embryo in the maternal uterus and leads to the distinction between the trophectoderm (TE, future placenta) and the inner cell mass (ICM, future embryo and yolk sac). In our recent study, we dissected the role of the Notch signalling pathway during early preimplantation development and found that it promotes the gradual loss of potency prior to the first lineage choice.
This project started back in 2014, after our previous work led by Teresa Rayon in which we characterised a regulatory element upstream of Cdx2, key gene in the specification of the trophectoderm2. Cdx2 was known to act downstream of the Hippo pathway, but we found that this enhancer was activated by the convergence of two pathways: Hippo and Notch. I had recently started my PhD in Miguel Manzanares’ lab at the Centro Nacional de Investigaciones Cardiovasculares (CNIC) in Madrid, and we decided to explore in more detail what Notch was doing in these early stages of mouse development.
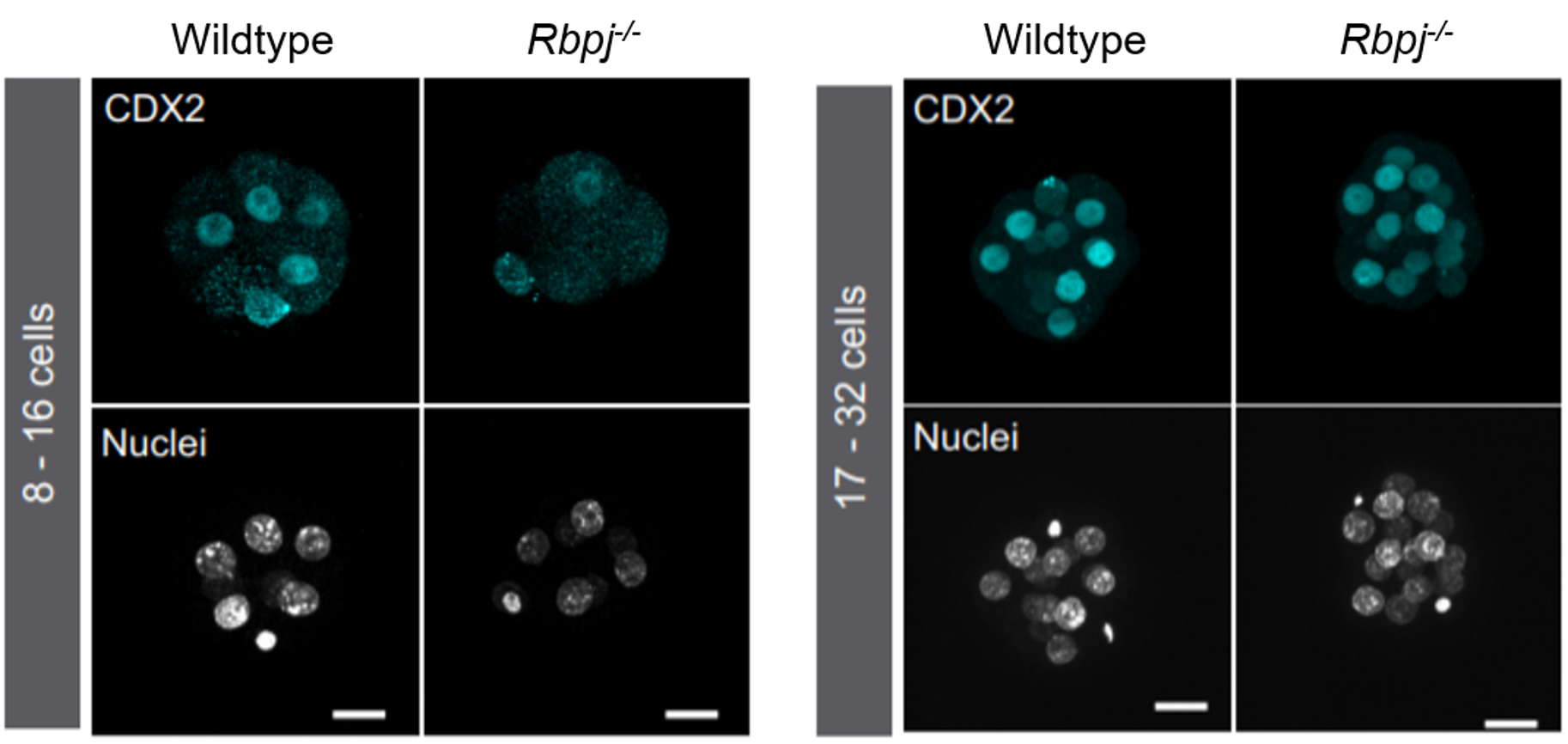
Different labs had been studying the role of the Hippo pathway during preimplantation development for a few years. So, we recapitulated all the information to try to find a cue of how Notch could be cooperating with it. Hippo is known to act as a readout of cell polarity and thus, only polarised cells (facing the outside) allow YAP to bind the transcription factor TEAD4 and activate Cdx2. However, initial expression of Cdx2 in the early compacted morula occurs in both inner and outer cells. Also, in the original work by Hiroshi Sasaki’s lab3, the authors claimed that the drop of Cdx2 expression in Tead4 mutant embryos was more dramatic in the blastocyst, when Cdx2 is only in the polarised TE, than in the morula. Therefore, our first hypothesis was that Notch could be acting in the early phases of Cdx2 expression, when this expression could not be completely explained by the role of YAP in polarised cells. We checked CDX2 in Rbpj and Notch1 mutant morulae (transcription factor and receptor of the Notch pathway respectively) and interestingly, we saw that CDX2 was strongly diminished specifically in the early morulae (<16 cells). After that, CDX2 levels were recovered, presumably because of the action of the Hippo pathway. Only double mutants for both Rbpj and Tead4 completely lacked CDX2 expression in the morula (and they did not reach the blastocyst stage). Cdx2 was responding to both pathways, but it seemed to behave differently depending on the stage. To be able to modulate the action of each pathway and verify if they were acting differently in these time windows, we used pharmacological inhibitors that allowed us to block them in a time-controlled manner. In the morula stage, only the blockade of the Notch pathway affected the expression of Cdx2. In contrast, the inhibition of TEAD/YAP was the one reducing Cdx2 from morula to blastocyst. Thus, we confirmed that although both pathways converged to regulate the same target gene, they did not act the same way.
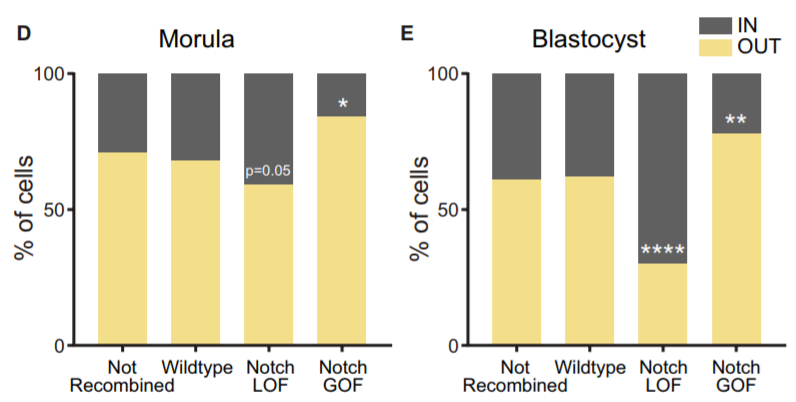
Another aspect we wanted to study was how Notch behaved during the morula to blastocyst transition, when its activity gets gradually restricted to the TE. In 2015, I was awarded a short stay fellowship from the Spanish Government that allowed me to visit the lab of Kat Hadjantonakis at the Sloan-Kettering Institute in New York for a few months, to continue with our collaboration and perform live imaging in embryos from the Notch activity reporter line (CBF1-VENUS)4. After some struggle, and with the help of Min Kang, we finally got movies where the embryos stayed in focus during the whole time-lapse (~24 hours). But the tougher part appeared later, back in Madrid. How could we manage all the information from those movies? The cell tracking for each embryo took a while… a long while. And once it was done, we still needed to consider many aspects. Fortunately, Antonio Lopez-Izquierdo, a biomedical engineer student, joined the lab to carry out the final project for his degree. He could programme in Matlab, so he developed a tool to 3D-reconstruct the embryos in each time frame and analyse the behaviour of the intensity of the reporter according to the position of the blastomeres within the embryo. The results indicated that there were already differences in the reporter intensity levels between outer and inner blastomeres in the morula, and that there was some correlation between these intensity levels and the position that cells occupied within the embryo which required further study. By then, the laboratory of Rui Benedito (our neighbours next door) had generated a transgenic line to produce mosaic cell populations with different Notch activity levels upon specific LoxP/CRE recombination5. We used this system to confront wildtype blastomeres with Notch loss of function (LOF) and gain of function (GOF) blastomeres. Beautifully, we saw how these differences affected the positioning of the cells within the embryo: Notch GOF blastomeres were more prone to occupy outer positions at the expenses of Notch LOF blastomeres which preferentially occupied inner locations.
At that point, we knew more details about the role of Notch regulating Cdx2 and favouring the positioning of blastomeres in the embryo. Nevertheless, we had the feeling that Notch was doing something else and we wished to gain a broader view on how it was working. Transcriptome profiling using low amounts of RNA was emerging, so that was the way to go. We decided to carry out single embryo RNA-seq in wildtype and Rbpj mutant morulae. Two issues arose: each sample consisted of only a dozen cells so we had to test and fine-tune the protocol to make sure that it would work with the limited material; and we could only genotype after the sequencing so we did not know how many samples of each genotype we had. Once we had it, the analysis showed that most of the genes (~70%) were downregulated; these were not only TE-related genes, but also some pluripotency genes. So, the role of Notch did not seem to be just the specification of the TE.
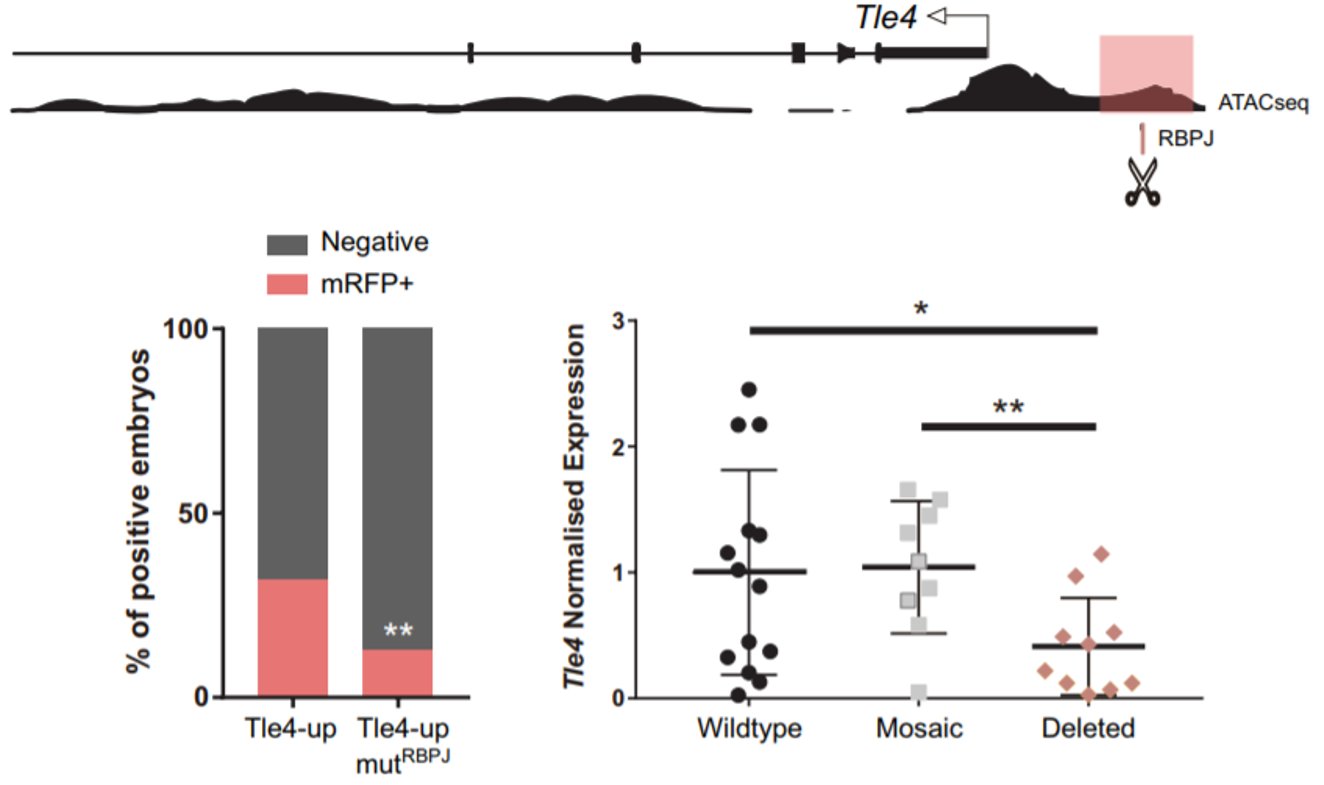
Remarkably, among the genes that were upregulated, we found two naïve pluripotency genes: Prdm14 and Dppa3. When we obtained this result, we had already seen that Notch was active very early, when the embryo consists of only four cells. Interestingly, Prdm14 is known to be heterogeneously expressed at the 4-cell stage and its expression fades away to be re-expressed in the ICM of the blastocyst6. That is the opposite pattern of Notch activity. Hence, we hypothesised that Notch could be blocking the expression of those naïve pluripotency genes in the early embryo, to boost the first differentiation programs. We did not think that this could be a direct target of Notch, given that when Notch is active it mainly activates gene expression. It is when Notch is not active that RBPJ can act as a repressor. We wondered if we could identify any gene that could be activated by Notch to repress the naïve pluripotency genes. To do that, we combined the analysis of our RNA-seq with predicted RBPJ binding motifs and data of ATAC-seq in 8-cell embryos7. In the resulting list (which was much more bearable), we found two candidates that had been described to block naïve pluripotency markers in mouse ES cells: Tle4 and Tbx3; and we decided to study them in parallel. With the help of Isabel Rollan in the lab, we identified two enhancers in the genomic landscapes of those genes that included an RBPJ motif in their sequences (Tle4-up and Tbx3-i7). To see if these sites were important for the regulation of the enhancers, we mutated these motifs and only the activity of the Tle4 enhancer was reduced. Finally, in order to see if these sites also affected the endogenous expression of Tle4 and Tbx3 respectively, we deleted them using the CRSPR/Cas9 system and we observed that Tle4 expression was affected in the edited embryos, but Tbx3 levels remained normal. Thus, RBPJ was important for the activity of the Tle4-up enhancer and for proper Tle4 expression in the morula.
We concluded that Notch was regulating the transition in the embryo towards the first lineage decision. First, promoting a differentiating scenario by blocking naïve pluripotency genes (possibly through TLE4) and then, inducing the expression of Cdx2 to specify the trophectoderm in cooperation with Hippo.
It is curious to think how the role of Notch during preimplantation development was ruled out more than a decade ago because single mutants were not lethal until postimplantation stages, and how convergence with other inputs can give robustness to embryonic development albeit masking important roles.
I want to finish thanking other lab members who were also involved in this exciting story: Mariajo Andreu, Julio Sainz de Aja and Javier Adan.
Read the full story:
Transitions in cell potency during early mouse development are driven by Notch
Sergio Menchero, Isabel Rollan, Antonio Lopez-Izquierdo, Maria Jose Andreu, Julio Sainz de Aja, Minjung Kang, Javier Adan, Rui Benedito, Teresa Rayon, Anna-Katerina Hadjantonakis, Miguel Manzanares
eLife 2019;8:e42930 DOI: 10.7554/eLife.42930
References:
- Slack, J. M. W. Conrad Hal Waddington: the last Renaissance biologist? Nat. Rev. Genet. 3, 889–895 (2002).
- Rayon, T. et al. Notch and hippo converge on Cdx2 to specify the trophectoderm lineage in the mouse blastocyst. Dev Cell 30, 410–422 (2014).
- Nishioka, N. et al. Tead4 is required for specification of trophectoderm in pre-implantation mouse embryos. Mech Dev 125, 270–283 (2008).
- Nowotschin, S., Xenopoulos, P., Schrode, N. & Hadjantonakis, A. K. A bright single-cell resolution live imaging reporter of Notch signaling in the mouse. BMC Dev Biol 13, 15 (2013).
- Pontes-Quero, S. et al. Dual ifgMosaic: A Versatile Method for Multispectral and Combinatorial Mosaic Gene-Function Analysis. Cell 170, 800–814.e18 (2017).
- Burton, A. et al. Single-Cell Profiling of Epigenetic Modifiers Identifies PRDM14 as an Inducer of Cell Fate in the Mammalian Embryo. Cell Rep. 5, 687–701 (2013).
- Wu, J. et al. The landscape of accessible chromatin in mammalian preimplantation embryos. Nature 534, 652–657 (2016).


 (1 votes)
(1 votes)