Stem cell fate choice: determined in an instant
Posted by Jun Chen, on 6 March 2018
Jun Chen
National Institute of Biological Sciences, Beijing
A discussion of our recent paper: Chen J, Xu N, Wang C, Huang P, Huang H, Jin Z, Yu Z, Cai T, Jiao R, Xi R. Transient Scute activation via a self-stimulatory loop directs enteroendocrine cell pair specification from self-renewing intestinal stem cells. Nat Cell Biol. 2018 Feb;20(2):152-161. doi: 10.1038/s41556-017-0020-0. Epub 2018 Jan 15.
Behind the story
On 2016-5-18, the second day after my first research paper 1 was published online at my third year of PhD courses, my mentor Rongwen Xi told me to take over the “EEP project”. This project had begun long before I started my PhD courses, and until my participation, has been passed along by three researchers in turn: Na Xu, Pin Huang, and Chenhui Wang; each of them subsequently graduated and moved on with their own academic or industrial paths. Encouraged by my first successful publication, I quickly agreed to take this seemingly never-ending project. I told Dr. Xi a sentence that amused me afterwards, “I won’t give it up until you give up.” This is how this long and tough process begins, and this is also the instant that determines the end of the story.
Introduction
Even as adults, we have stem cells throughout our bodies that are responsible for maintaining many of our tissues. These adult stem cells constantly divide and produce daughter cells, which, through a process called differentiation, become multiple types of mature cells. The fate of the daughter cells can be actively specified by asymmetric cell division, in which cell fate determinants are specifically segregated into one of two stem cell daughter cells 2. Alternatively, cell fate can be specified passively; in this case, cells physically depart from the self-renewal niche environment, as with the specification of cystoblasts from Drosophila germline stem cells, and the initiation of differentiation of stem cells in the mouse small intestine upon their departure from the Paneth cell niche 3,4. Despite several implications from these “renew or differentiate” fate determination events, very little is known about the molecular mechanisms by which distinct, lineage-restricted progenitor cells are generated from a common stem cell pool.
To study this question, we investigated cell fate in a multipotent intestinal stem cell (ISC) experimental model from adult fruit flies. The default mode for cell fate is that ISCs differentiate into enterocytes (EC), which have been shown to occur from approximately 90% of ISC divisions 5. However, there is a less-well-understood mode in which ISCs differentiate into pairs of enteroendocrine cells (EEs), which occur from approximately 10% of ISC divisions.
When I started to do this project, previous studies suggested that EEs are directly differentiated from ISCs, implying that the decision of EE specification may occur at the stem cell level in ISCs 6,7, but how this occurs remains unclear. It has also been revealed that the four-gene cluster acheate-scute complex (As-c) act as EE-fate-determination factors. Furthermore, one of the As-c genes, scute (sc), is both necessary and sufficient for EE specification. Nevertheless, important questions remain about both the molecular and cellular mechanisms through which Sc functions in EE fate decision, and we do not yet know how Sc is regulated in ISCs to control EE fate.
We finally answered these questions in our recent paper, in which we reported that transient activation of Sc determines both the type and number of committed progenitor cells from Drosophila ISCs.
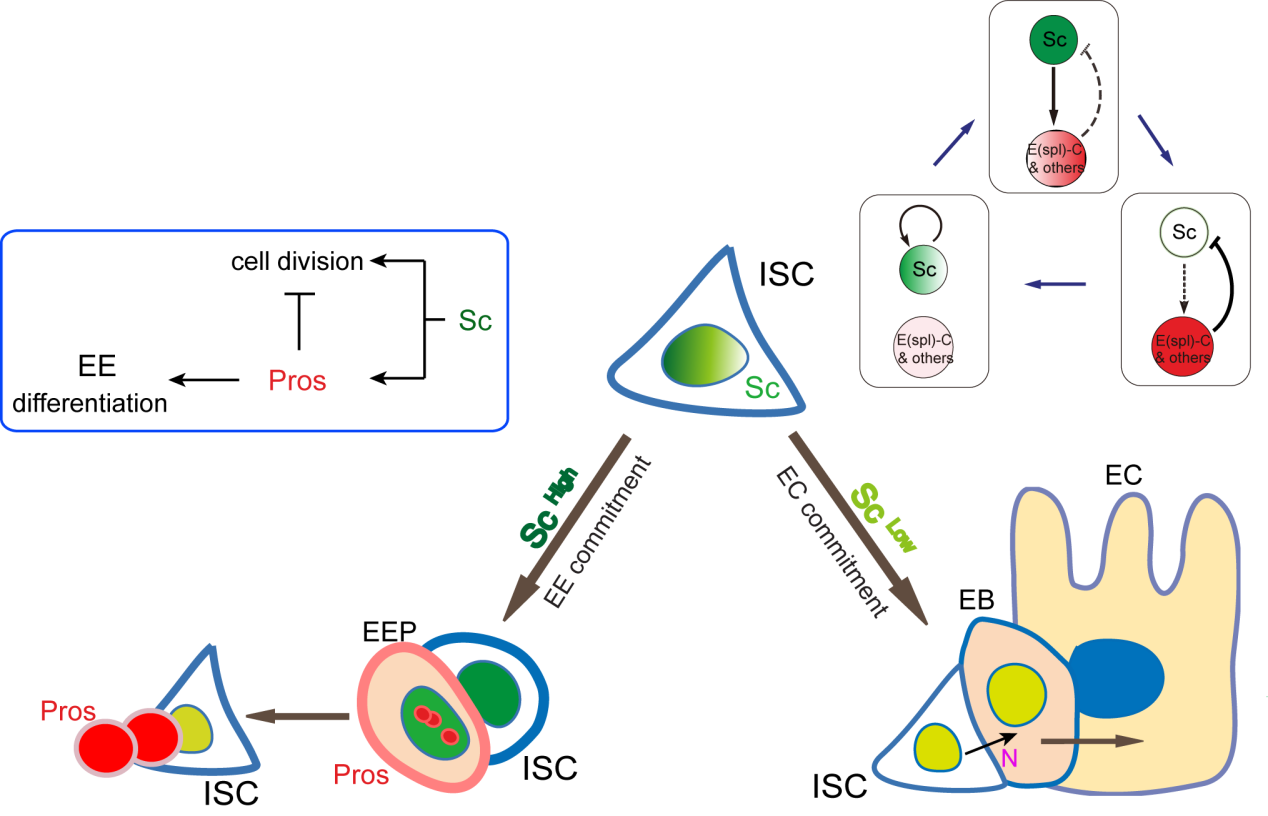
A cell fate is determined by a transiently expressed protein
To better understand the process of EE specification in ISCs, we set up an EE regeneration assay and examined de novo EE regeneration. This assay was first beautifully set up by Na Xu and Pin Huang. Based on the finding that Sc is required for EE generation from ISCs, we temporally knocked down sc starting from the pupal stage, and this process produced flies with midguts lacking EE cells. We then used these EE-less midguts to examine the process of EE production by using temperature shift to re-introduce Sc expression in the midgut. With this assay, we discovered that (i) ISCs actually undergo an initial division to generate a new EE progenitor cell (EEP), and (ii) the EEP then undergoes one final round of cell division to produce a pair of EEs (Figure 2).
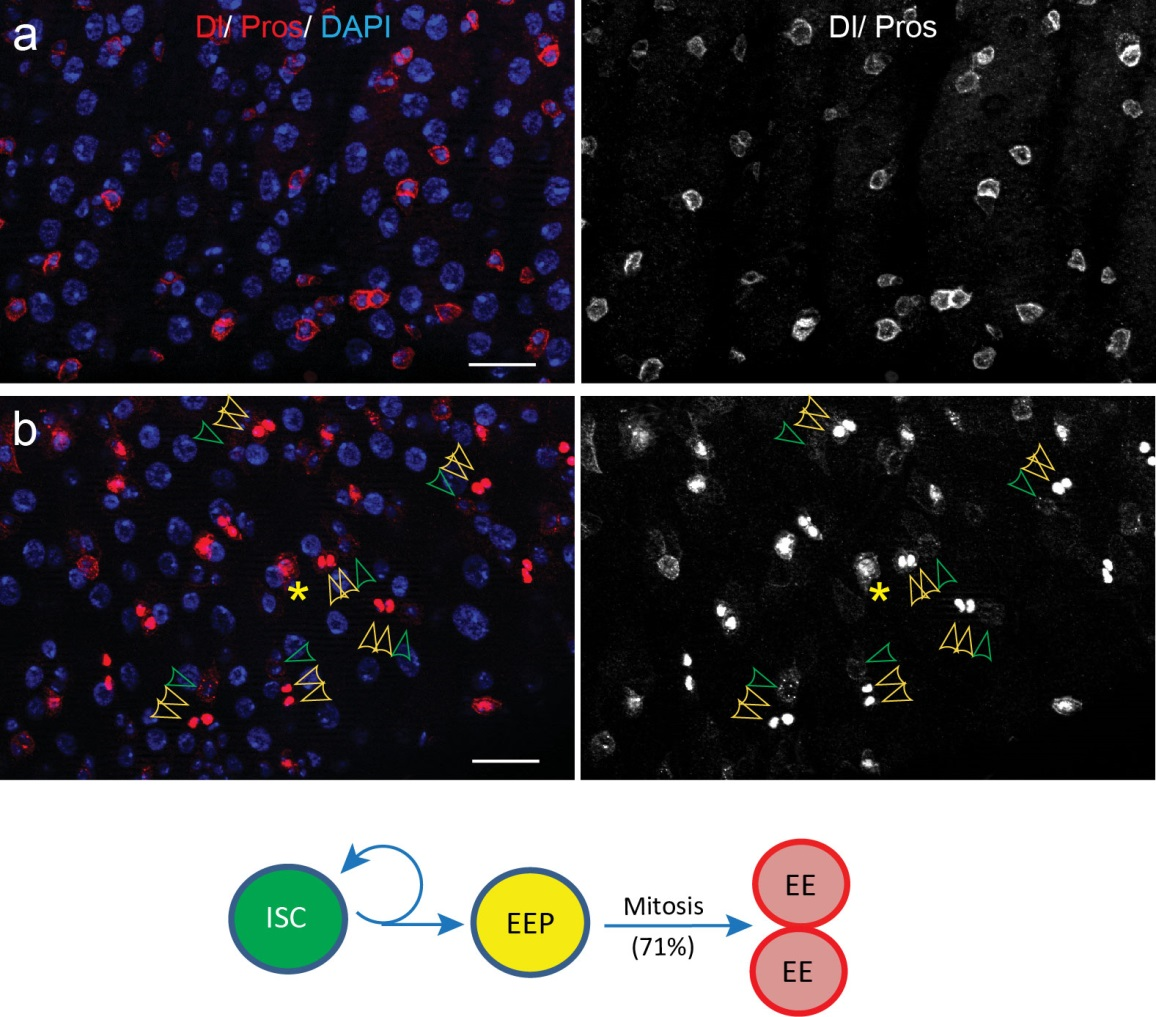
To further analyze this two-step cell division process, Chenhui Wang genetically overexpressed sc in ISCs and monitored the cellular events in a time-course experiment. Chenhui found that transient sc expression caused a rapid cell division response, and also induced expression of the EE-marker gene Pros, which is known as a potent cell-cycle inhibitor. These findings and subsequent experiments enable us to precisely define the regulatory circuitry that directs the formation of a pair of EEs from each ISC (Figure 1). Here a concern still exists that we have not given a “seeing is believing” results for cellular events of EE generation because we have not established long-term live imaging technique for fly midgut yet. To solve this problem, I expressed a UAS-RedStinger reporter in sc overexpression system. RedStinger is relatively stable and can serve as a lineage marker to trace the progeny of the originally marked ISCs. The number of cell divisions of the initially labeled ISCs could be deduced based on the mitotic marker PH3 and the number of RedStinger+ cells in a single cluster. In this experiment, I observed a tightly ordered process: The first cell division following sc overexpression occurred in ISCs (PH3+ in a one-cell clone), and at telophase of the first cell division, one of the two daughter cells began to show cytoplasmic Pros accumulation; the second cell division (PH3+ in a two-cell clone) always occurred in the Pros+ daughter cell, that is EEP; the third cell division (PH3+ in a three-cell clone) occurred again in ISCs. These observations suggest that EEs are generated from ISCs via two rounds of cell divisions: an asymmetric division of ISC to generate an EEP, and then the EEP division to produce an EE pair (Figure 3).
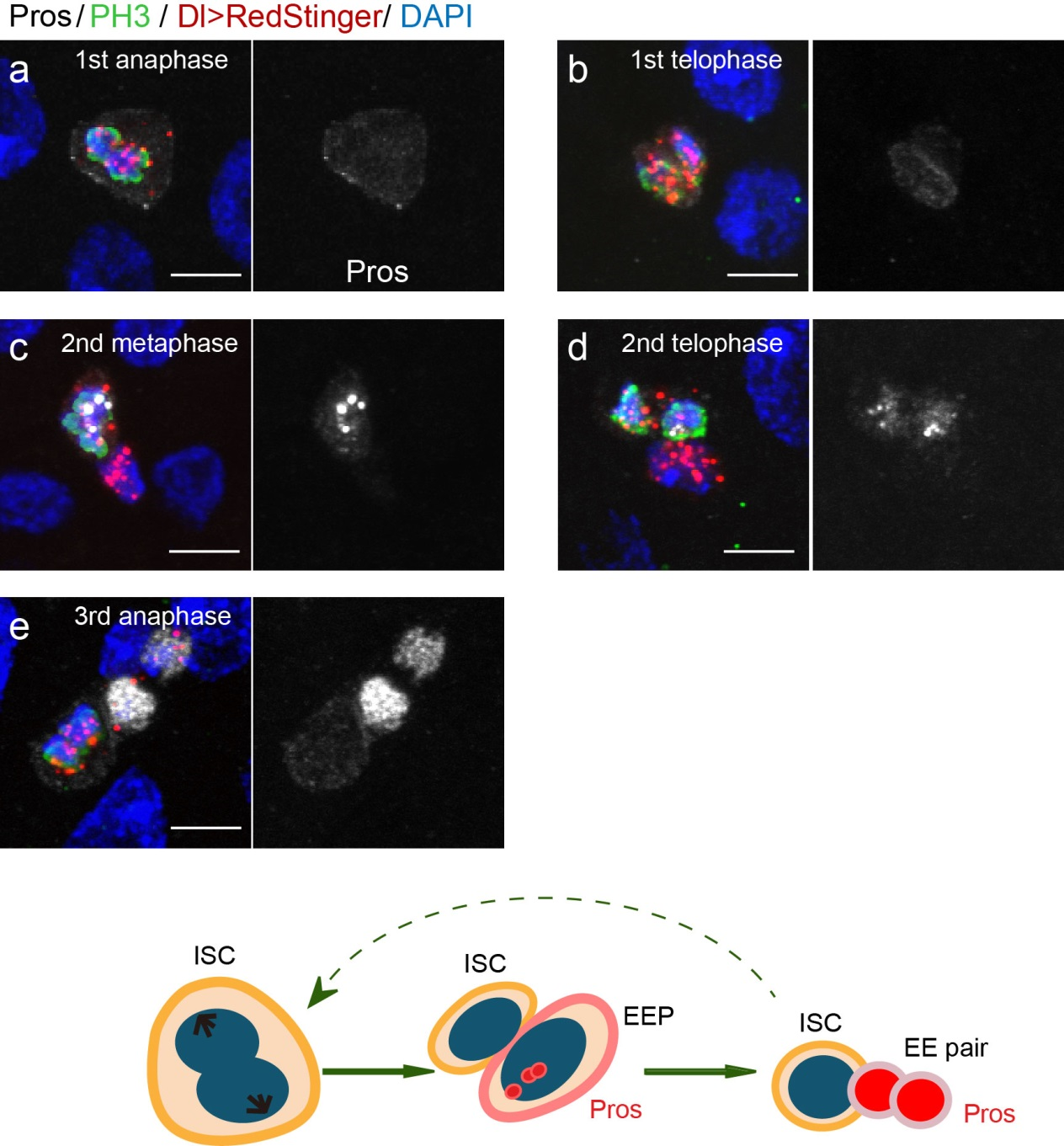
Next, to visualize the expression of Sc in midgut, Pin generated a green fluorescent protein (GFP) tagged line for Sc in collaboration with Zhongsheng Yu and Renjie Jiao from the Institute of Biophysics of the Chinese Academy of Sciences. In this line, the GFP was fused to 3’ of the Sc coding region. Initially we were a little bit disappointed as the GFP signal was too weak to visualize and all the researchers had to immunostain with anti-GFP antibody, which effectively amplified the Sc-GFP signal. Immunostaining results revealed that Sc-GFP could be observed in virtually all ISCs but the expression level is largely indistinguishable among ISCs. With improved microscopy technology, I managed to capture GFP signal in unstained samples and found that the Sc-GFP fusion protein is expressed at higher levels in ~15% of the ISCs (Figure 4). This result was exciting because it indicated that Sc may be expressed in a dynamic manner in ISCs, and in a snap shot, you may see a weak expression level in most ISCs, and increased expression levels in a small subset of ISCs.
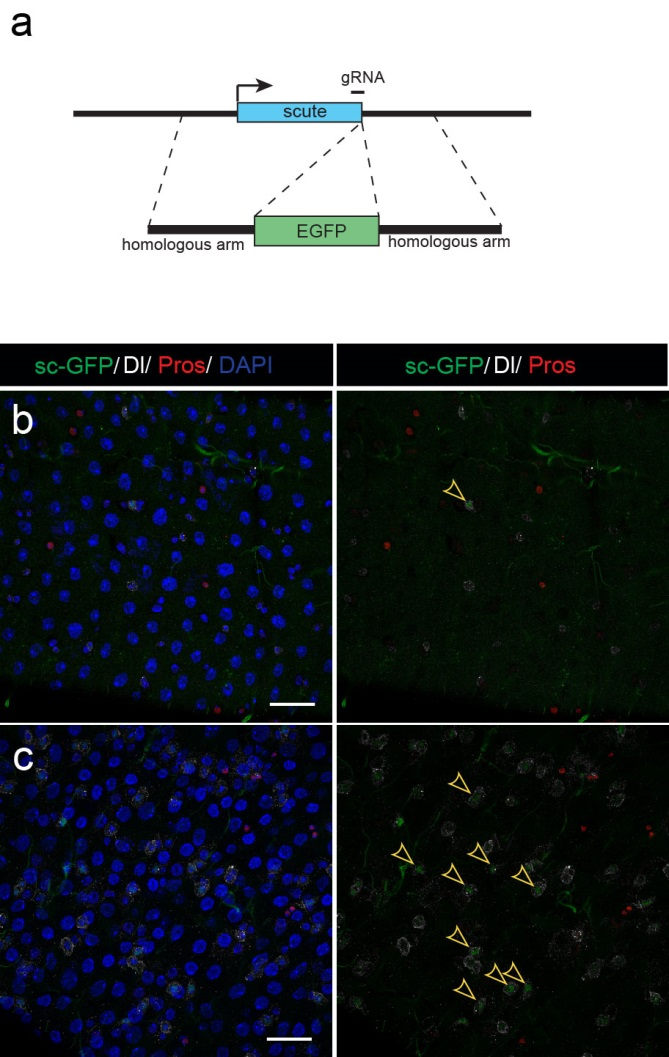
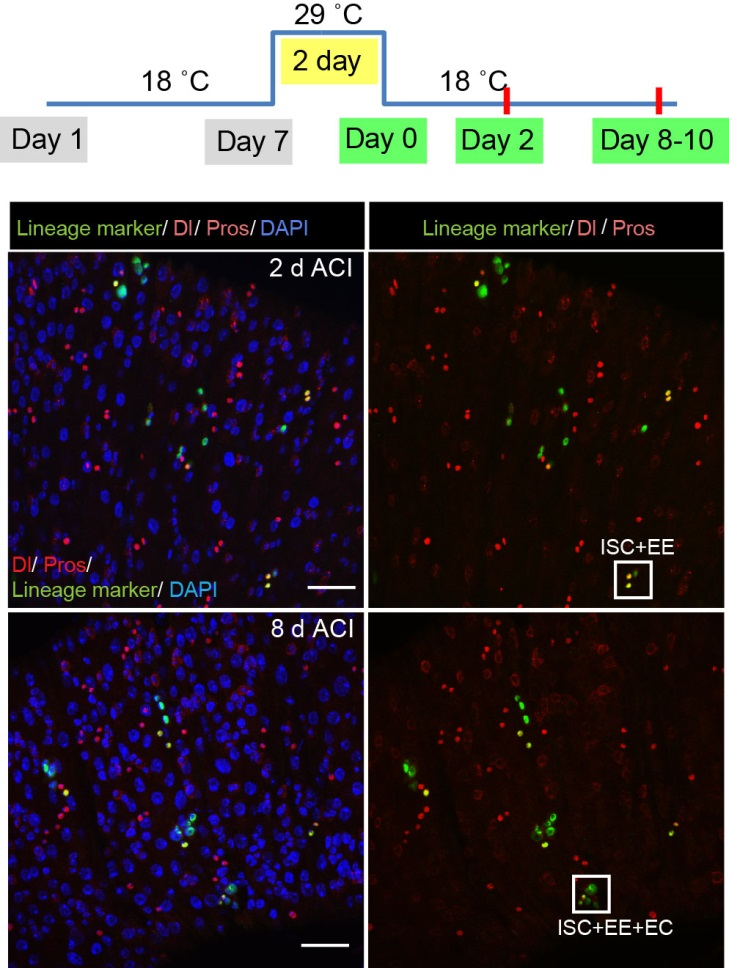
The next step was to test the cell lineage fate of these Schigh ISCs. The follow-up cell lineage tracing studies with a Sc-Gal4 line will help to do that, but there was no available Sc-Gal4 line at that time. Fortunately, from 23 (upstream and downstream) glass multiple reporter (GMR) enhancer-GAL4 lines generated for sc, I identified one GAL4 line that drove UAS-GFP expression in some diploid cells in the midgut epithelium. The density (also in ~15% ISCs), distribution, and individual variability of the GFP+ cells were largely similar to those of Sc-GFP+ cells, and about half of RFP reporter driven by this GAL4 line recapitulates Sc-GFP expression, suggesting that this GAL4 line is driven by the enhancer element for sc expression in the midgut. Cell lineage tracing studies with this GAL4 line revealed that the immediate daughter cells of Sc-GAL4+ ISCs were mainly EEs; however, these ISCs re-assume their default EC-producing fate once Sc expression is downregulated (Figures 1&5).
The knotty problem
With these exciting new observations, we inevitably faced a mechanistic question, “How does such transient upregulation of Sc in ISCs occur?” This question comes like a boss in video games, and has always been difficult to tackle. Studies over the decades on proneural genes have revealed that the AS-C genes in the neural cell lineages are regulated by highly-complex-cis-regulatory regions, and these regulatory regions are considered to constitute an integrating device for multiple signaling regulators and chromatin factors. Firstly came to our minds was to avoid such “net” and to set out from the reported signals that regulate EE specification in Drosophila midgut. Previous studies suggest that the Slit molecules secreted from EEs activate the Robo2 receptors of ISCs to prevent EE generation, thereby establishing a negative feedback to coordinate EE production with tissue demand. However, Sc expression pattern was unaltered in Robo2 mutants, in which the excessive EE phenotype was prominent. Considering Robo2 activation in ISCs is not sufficient to prevent EE production from ISCs, this mechanism appears to be a modulator rather than a key component in the EE fate decision process. Thus, I had to go back to hit the core of the question, the transcriptional control of As-c genes.
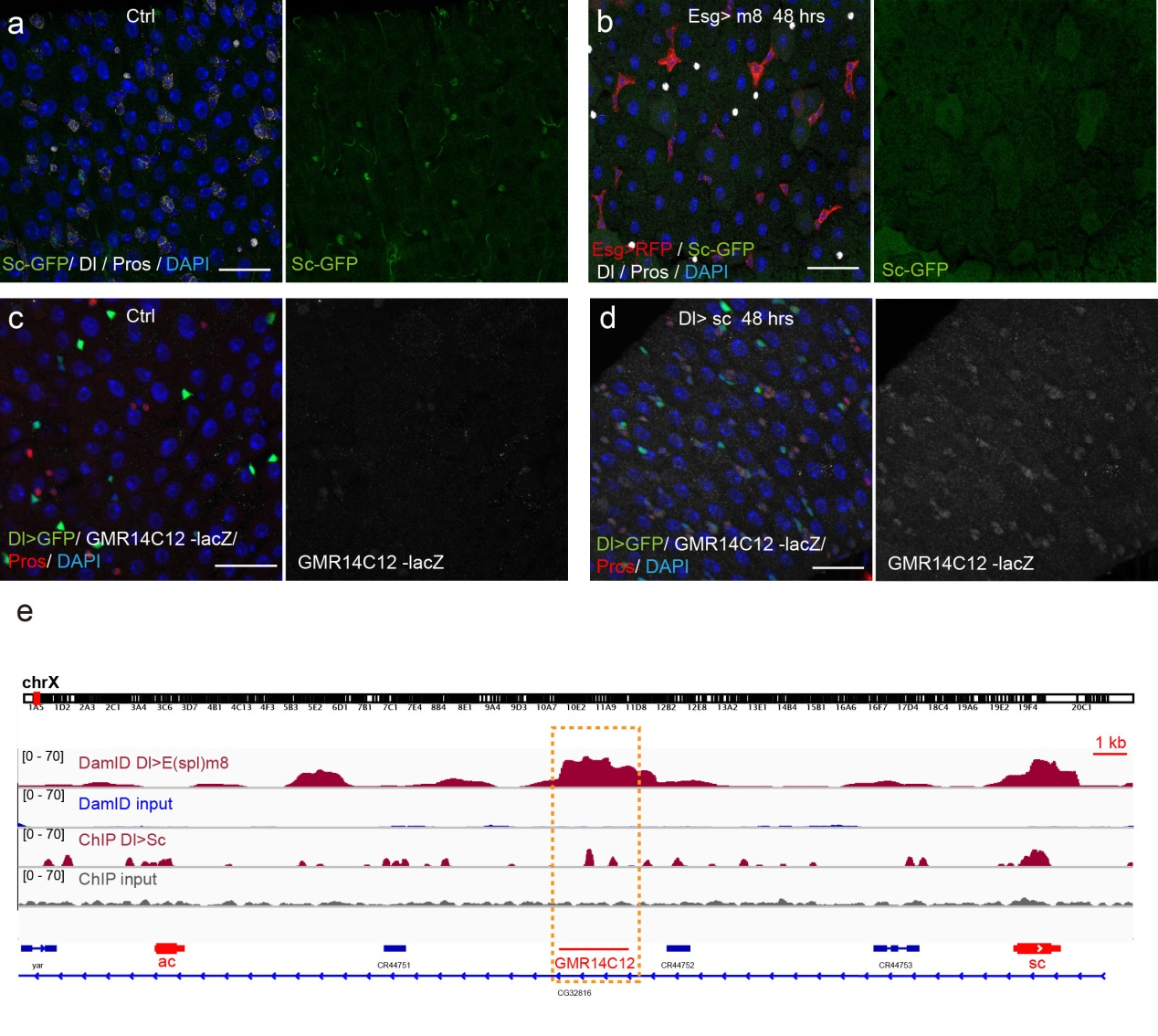
Previous studies on early Drosophila development have suggested reciprocal regulatory relationships between AS-C genes and the enhancer of split complex (E(spl)) genes, which are known as the Notch target genes. Inspired by these reports, I screened a number of candidate reporters for individual E(spl) genes, and identified a single reporter, m8-lacZ, which showed a weak, but similar expression pattern to Sc in wild type guts. To characterize the regulatory relationship between Sc and E(spl)m8, I transiently overexpressed sc in ISCs, and surprisingly saw robust upregulation of m8-lacZ expression in all ISCs. Notably, co-expressing Notch-RNAi did not prevent the upregulation of m8-lacZ expression caused by sc overexpression, suggesting that E(spl)m8 expression is independent of Notch activity in ISCs. To test whether such regulatory relationship similarly applies to other E(spl) genes, I sorted out sc-overexpressed ISCs for mRNA profiling by RNA-seq analysis. Strikingly, in addition to m8, many other E(spl) genes, including m4, m6, m7, mγ, and mδ were strongly upregulated upon sc overexpression. By combining genetic assays and ChIP-seq analysis, we showed Sc could bind to the enhancer regions of many E(spl) genes, and directly upregulate these E(spl) genes in ISCs (Figure 6).
It’s then instinctive to consider whether these E(spl) genes, also known as neural fate repressors, would in turn negatively regulate Sc expression. By combining genetic assays and targeted DamID analysis using a E(spl)m8-Dam fusion line, we showed that E(spl)m8 suppresses sc expression by directly binding to the enhancer region of sc. The direct two-way regulation between Sc and E(spl)m8 form a typical negative feedback regulatory loop, which may explain the transient activation pattern of Sc in ISCs (Figure 6).
The question still has half part unanswered, “how does sc initially build up?” Searching for other transcriptional activators, like other bHLH activators as reported, would make this question a “chick and egg” issue. Interestingly, Sc has been reported to transcriptionally self-stimulate itself, which acts as an essential mechanism for proneural protein accumulation during sensory organ development. To test whether self-stimulation of Sc also occurs in ISCs, we constructed LacZ transcriptional reporter for sc using the Sc-Gal4 enhancer fragment that we had identified. This lacZ reporter was barely detectable in WT midgut epithelium, but effectively induced in ISCs when sc was transiently induced. ChIP-seq data analysis also revealed two Sc binding peaks within this Sc-Gal4 enhancer region (Figure 6). Thus, Sc is able to stimulate its own transcription directly by binding to sc enhancer. Together, our results suggest that two feedback regulatory loops control the transient upregulation of Sc in ISCs prior to EE fate commitment. There is a transcriptional self-stimulation loop that allows Sc to gradually build up and eventually reach a high level to induce EEP specification, and there is a negative feedback regulation loop between Sc and E(spl) genes that returns sc expression back to the baseline level (Figure 1).
The beginning of the end
Given that negative feedback is a common mechanism underlying biochemical oscillations in virtually all organisms, the feedback loops between Sc and E(spl) genes could plausibly be the driver of an oscillatory expression pattern for Sc in ISCs; in theory such oscillatory expression could potentially serve as an internal timer for periodic production of EEs from ISCs. This clock mechanism would be similar to what is known about the circadian clocks, a biological research field that was recently honored with the 2017 Nobel Prize for Physiology or Medicine. We are obviously very excited about the findings and potential implications. However, this is just a tip of iceberg, future cellular and molecular analysis, likely in combination with in vivo live imaging work will allow further testing and refining of the oscillation model proposed in our study, and such experiments will determined whether and how any internal timer is regulated by certain endogenous and/or environmental cues, and whether the oscillation model is generally applicable in other tissue stem cells, including that in humans.
Finally, I want to say that I am very fortunate and grateful to be a part of such a wonderful research team and work on such an exciting project. This work would not be possible without the contribution and help from our past and current lab members, especially Na Xu, Chenhui Wang, and Pin Huang, as well as informaticians Huanwei Huang and Tao Cai at NIBS. I especially want to thank my mentor Dr. Xi for his great guidance and trust, as well as his helpful advice on the writing of this article. As you can imagine, in addition to the “high” moments when the exciting results were first observed, I also had many upset and head-scratching moments during the course of this study. These experiences have endowed me a lot on how to explore, to observe, to cooperate, to write, and to persevere. I believe that no matter how hard it seems like, if you continue to stay focused and think hard, great things may eventually happen, in an instant.
References
1 Chen, J., Xu, N., Huang, H., Cai, T. & Xi, R. A feedback amplification loop between stem cells and their progeny promotes tissue regeneration and tumorigenesis. eLife 5, doi:10.7554/eLife.14330 (2016).
2 Neumuller, R. A. & Knoblich, J. A. Dividing cellular asymmetry: asymmetric cell division and its implications for stem cells and cancer. Genes Dev 23, 2675-2699 (2009).
3 Morrison, S. J. & Spradling, A. C. Stem cells and niches: mechanisms that promote stem cell maintenance throughout life. Cell 132, 598-611 (2008).
4 Li, L. & Xie, T. Stem cell niche: structure and function. Annual review of cell and developmental biology 21, 605-631 (2005).
5 Ohlstein, B. & Spradling, A. Multipotent Drosophila intestinal stem cells specify daughter cell fates by differential notch signaling. Science 315, 988-992 (2007).
6 Zeng, X. & Hou, S. X. Enteroendocrine cells are generated from stem cells through a distinct progenitor in the adult Drosophila posterior midgut. Development 142, 644-653, doi:10.1242/dev.113357 (2015).
7 Biteau, B. & Jasper, H. Slit/Robo signaling regulates cell fate decisions in the intestinal stem cell lineage of Drosophila. Cell reports 7, 1867-1875, doi:10.1016/j.celrep.2014.05.024 (2014).


 (4 votes)
(4 votes)