The people behind the papers – Kayt Scott and Bruce Appel
Posted by the Node Interviews, on 26 December 2020
This interview, the 86th in our series, was published in Development earlier this year.
In the developing spinal cord, progenitor cells sequentially give rise to motor neurons and precursors of one of the major glial cell types: oligodendrocytes. A new paper in Development unpicks the molecular control of the neuron-glia switch and the differentiation of oligodendrocyte precursors in the zebrafish embryo. To find out more about the work, we met first author and graduate student Kayt Scott and her supervisor Bruce Appel, who holds the Diane G. Wallach Chair of Pediatric Stem Cell Biology and is Professor and Head of the Section of Developmental Biology at the Department of Pediatrics, University of Colorado School of Medicine in Aurora.

Bruce, can you give us your scientific biography and the questions your lab is trying to answer?
BA: I earned my PhD at the University of Utah under the mentorship of Shigeru Sakonju. My thesis work focused on transcriptional control mechanisms in the context of homeotic genes and developmental patterning, and it was during that time that I became interested in problems of cell fate specification. I then moved to the University of Oregon for postdoctoral training with Judith Eisen. Using zebrafish embryos, Judith had previously performed a brilliant set of experiments that led her to conclude that communication between newly born motor neurons determined their subtype identities. As I worked to try to figure out how those subtype identities were specified, other work in the field began to suggest that the same progenitor population that produces motor neurons also gives rise to oligodendrocytes: the myelinating glial cell type of the CNS. I initially thought oligodendrocytes and myelin were pretty boring but as we developed new tools and learned how to do time-lapse imaging, we discovered that these cells and their ability to form a myelin sheath on an axon is incredibly fascinating. This has spun off into many projects investigating the molecular and cellular mechanisms that regulate developmental myelination and how myelin changes in response to brain activity. However, I still consider the age-old question of cell fate specification to be the foundation of all that we do.
And Kayt – how did you come to work in Bruce’s lab and what drives your research today?
KS: Developmental biology has always been a passion of mine, especially in the context of early nervous system formation. So, before I had even applied to graduate school, I was aware of and intrigued by the work conducted in Dr Bruce Appel’s lab. In fact, I discussed my interest in his research in my application letter. Once I entered graduate school, I did a rotation in Dr Appel’s lab and began the study of Prdm8 function in the zebrafish spinal cord. By the end of my rotation, I was sure I wanted to be part of the lab and further my rotation project. Through my experiences in the lab, I have really become interested in understanding how early tissue structure is organized and how this organization is altered through time and these questions are what drives my research today.
How has your research been affected by the COVID-19 pandemic?
KS: At the time of the pandemic shut down I was working from home already, feverishly compiling this manuscript and working on a fellowship application, so the first couple of months weren’t too disruptive. However, after accomplishing these endeavours, I have been unable to get into the lab and further explore the mechanisms of how Prdm8 regulates Shh signalling response. Fortunately, this has provided me with a unique opportunity to delve into the single cell RNA-seq dataset we had generated and develop my bioinformatic skills. This has opened many new avenues of research in the Appel lab.
I am hopeful that the pandemic is making us more collaborative and communicative
BA: I am hopeful that the pandemic is making us more collaborative and communicative. Although we all miss interacting with our friends, colleagues and trainees, the pandemic is forcing us to find new ways to share information. I have been really impressed by how well the virtual scientific conferences are working. The conferences are now available to so many people who couldn’t afford them or had other barriers to travel. Additionally, distance no longer limits with whom we can share seminars or journal clubs or lab meetings. Consequently, I’m optimistic that our research will actually benefit because we will have better information and shared expertise.
In your efforts to understand the regulation of pMN development, what made Prdm8 a good candidate?
KS: Originally our interest in Prdm8 was motivated by understanding how oligodendrocyte precursor cells (OPCs) are maintained into adulthood. Prdm8 was identified in our bulk RNA-seq data as being highly expressed by OPCs and not expressed in oligodendrocytes, and because Prdm8 was a known transcriptional repressor that can complex with bHLH transcription factors, we predicted that Prdm8 was forming a complex with Olig2 to repress oligodendrocyte differentiation. As we began our investigation of Prdm8 function, we came across some interesting and unexpected phenotypes that transitioned our interests in Prdm8 function within pMN progenitors.
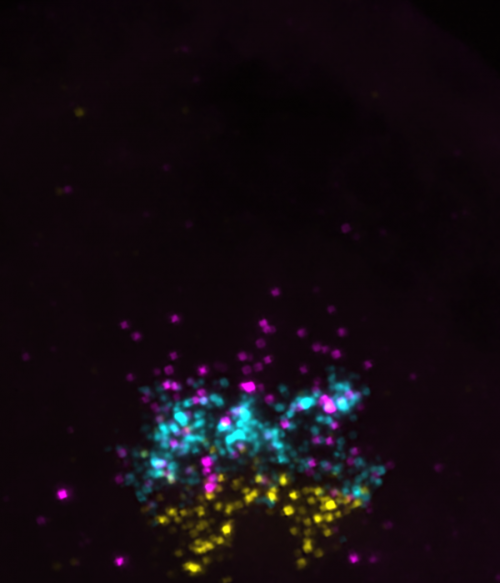
BA: Whereas much of the myelin field focuses on how oligodendrocytes make myelin, we have been curious about why so many OPCs persist without differentiating. Prdm8 jumped out of our RNA-seq data because OPCs expressed it at very high levels compared with myelinating oligodendrocytes. Because Prdm8 can function as a transcriptional repressor, the initial hypothesis was obvious. As Kayt notes, however, her work revealed that Prdm8 has an earlier function in regulating progenitor fate specification.
Can you give us the key results of the paper in a paragraph?
KS: We set out to understand the potential function Prdm8 plays in pMN cell fate. Detailed expression analysis revealed that pMN progenitors express prdm8 through development and that differentiating oligodendrocytes downregulate prdm8 expression, whereas OPCs maintain expression. We then learned that prdm8 mutant embryos have an excess of oligodendrocytes and a deficit in motor neurons as a result of a premature motor neuron-oligodendrocyte fate switch. This premature switch in cell fate was accompanied by a premature increase in nkx2.2a expression and ventral neural tube Shh activity with Prdm8 loss of function. By suppressing Shh signalling in prdm8 mutants, we were able to restore the motor neuron population but were not able to reduce the excess oligodendrocyte phenotype observed in mutants. In all, these data support the hypothesis that Prdm8 prevents the neuron-glia switch in pMN progenitors by suppressing Shh signalling activity, and, independently of Shh activity, Prdm8 regulates the balance of OPCs and oligodendrocytes.
Do you have any clues as to how, at the molecular level, Prdm8 inhibits Shh signalling?
KS & BA: One possibility we have entertained is that Prdm8 might form a complex with Olig2 to suppress transcription of genes that promote cellular response to Shh. Figuring this problem out will be the next big push for the Prdm8 project.
When doing the research, did you have any particular result or eureka moment that has stuck with you?
KS: I would have to say the most striking result I came across in this project was that Shh repression was only able to rescue motor neuron populations, but resulted in a total oligodendrocyte lineage deficit while maintaining excess oligodendrocytes in mutant embryos. This was the point where it really sunk in that Prdm8 function was more complex than I imagined, and appeared to have temporal and spatially distinct functions even within a single lineage.
And what about the flipside: any moments of frustration or despair?
KS: The most frustrating part of this project was that I had started it as a rotation student and hadn’t quite honed my organizational skills. This resulted in having to redo several of the first experiments, because I had not kept good enough documentation of my methods or results. A lesson that will never leave me!
What’s next for you after this paper?
KS: Moving forward, I have been working on a scRNA-seq paper, characterizing pMN cell populations through time and have plans to graduate in the summer of 2021. Following graduation, I hope to obtain an IRACDA fellowship to receive comprehensive training in undergraduate education and develop my own research projects in the field of developmental biology.
Where will this story take the Appel lab?
BA: There is an old joke that, in the absence of good ideas, do a genetic screen. A modern-day version of that is to do single cell RNA-seq. We have been stumped about the earliest molecular events that specify neural progenitors for oligodendrocyte fate, so we carried out a series of single cell RNA sequencing experiments. The data are incredibly exciting and appear to match our previous fate-mapping studies suggesting that motor neurons and oligodendrocytes arise from distinct progenitors. We now have many new ideas to pursue in our attempt to gain a comprehensive understanding of the gene regulatory network that specifies oligodendrocyte fate.
Finally, let’s move outside the lab – what do you like to do in your spare time in Colorado?
KS: Outside of the lab, I never waste an opportunity to enjoy the beautiful state we live in. You will find me camping, hiking and fishing almost every weekend during the warmer months; in the winter, I am hitting the slopes on my snowboard or gliding through the woods on some cross-country skis.
BA: Now that our daughters are pretty much on their own, my wife and I try to spend as much time as possible at our place in the mountains. It’s off-grid and fairly remote, and a great jumping-off spot for hiking, backpacking, skiing and snowshoeing. I am also very slowly and ineptly trying to restore a 1942 Ford tractor that my parents had on their farm.


 (No Ratings Yet)
(No Ratings Yet)