The story behind the screen – flashbacks from the first RNAi screen in a whole vertebrate
Posted by Raman Das, on 9 May 2011
The story of our recently released Development paper ‘FatJ acts via the Hippo mediator Yap1 to restrict the size of neural progenitor cell pools’ involves hundreds of dozens of fresh free-range eggs and not trivial amounts of time spent peering down a microscope. I have written this with Nick van Hateren, who is the joint first author of this paper along with me.
We had recently developed a short hairpin based system for carrying out RNA interference in the chicken called the pRFPRNAi system. This was an exciting time in the lab, as there had previously been no such system to carry out functional genetics in our favourite model system, and we were looking forward to make full use of it. This was also a great chance for us to demonstrate to the community that the chicken really is an excellent model system to do RNAi screens.
During this time, RNA interference screens were all the rage, and several screens had already been carried out in Drosophila cell lines and the worm, but never in a whole vertebrate. Even though shRNA was possible in the mouse, introducing constructs into mouse embryonic tissue was not a trivial matter, and carrying out a screen even on a small scale would present significant challenges, and indeed still would. The main obstacle here was the inaccessibility of the mouse embryos as they developed within the mother. The chick embryo, on the other hand came conveniently packaged inside an egg, and transfecting tissues by electroporation is a well-established and efficient technique. The spinal cord, in particular, was ideal for our studies. It is shaped like a tube, making it easy to inject it with a DNA solution. The DNA can then be transfected to only one side of the spinal cord by electroporation, while the other side would remain as a convenient internal control.
Armed with these reassuring facts, we began to search for suitable candidates for an RNAi screen. We had previously carried out a microarray analysis of the chick spinal cord, and amongst the thousands of genes expressed there, there were 40 genes that contained cadherin domains. These appeared to be the perfect choice, since they were a reasonable number and also because the large size of the cadherins makes it difficult to carry out overexpression studies.
We decided it would be prudent to target three different regions of each gene, which meant that we would need to sub-clone 120 shRNA sequences. Even though this sounded like a daunting task, the reality was far from it. Our cloning strategy was already well optimised, and we were done sooner than we expected. It was time to get down to the interesting work…
The screen itself was carried out very systematically. We had planned out the whole week so we could get in two rounds of electroporations and end up with a batch of fixed and frozen embryos ready for sectioning and analysis. The longest part was actually sectioning the vast numbers of embryos we generated, but we were lucky that our friendly lab technician Vicky was happy to give us a hand with this.
The screen was a rollercoaster ride of emotions – ranging from euphoria to dejection when a whole batch of our antibodies went bad. By the end of it we had many more positive hits than we had expected, and the range of phenotypes was also reassuringly diverse (Examples of two are in the image below). We were very excited that this would be a wonderful showcase for the feasibility of RNAi screens in the chick embryo.
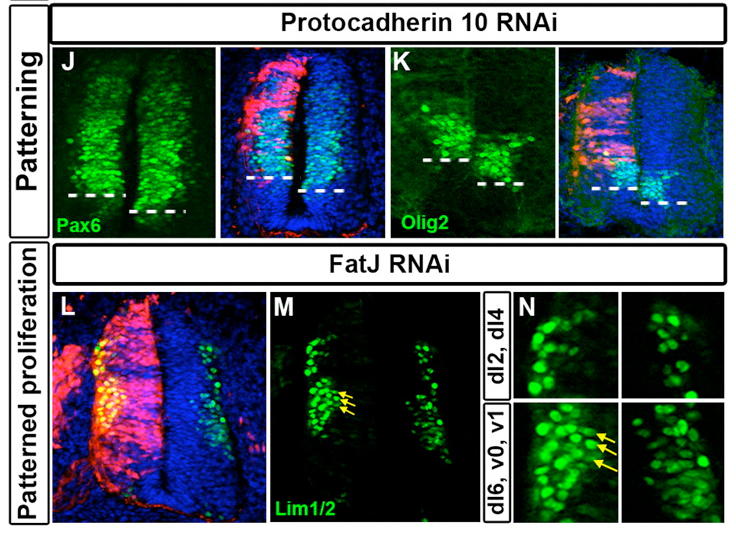
One of our most intriguing hits was FatJ, a cadherin that appeared to be important for controlling the number of a small sub-population of interneurons. Loss of FatJ caused a small but robustly reproducible increase in the number of these interneurons, and we were intrigued to understand more about this phenotype.
We found that FatJ expression is restricted to the intermediate region of the neural tube, and we were very encouraged to find that this domain corresponded to the progenitor pools for the interneurons whose numbers were increased following FatJ knockdown. We then examined the number of cells in different progenitor pools within the FatJ expression domain. After a great many cell counts and many hours of confocal microscope time, we determined there was a corresponding increase in the number of progenitor cells within the FatJ expression domain. This gave us a valuable clue to the mechanism of FatJ action: the loss of FatJ causes an increase in the number of progenitors which then differentiate normally to produce a corresponding increase in the number of interneurons. We confirmed this by double labelling with progenitor and differentiated interneuron markers and ensuring no cells expressed both markers simultaneously.
At that time, there were relatively few studies of FatJ reported in the literature; however we noticed that FatJ was the closest vertebrate orthologue of Drosophila Fat (dFat) which was known to be involved in planar cell polarity and was upstream of the newly-discovered Hippo pathway that controls tissue size in Drosophila. Many components of this pathway are highly conserved in vertebrates so we reasoned that FatJ might act through the Hippo pathway to regulate proliferation of neural progenitors. The Hippo pathway is a MAP Kinase cascade that phosphorylates the transcriptional regulator Yorkie (Yap in vertebrates) and this prevents the expression of proliferative and anti-apoptosis genes. Our hypothesis was that, in the absence of FatJ, there was no signalling through the Hippo pathway so Yap1 was not phosphorylated and proliferative genes continued to be expressed. This would lead to an increase in the number of cells within the progenitor pool. To test this theory, we designed shRNAs to target Yap1 and Tead4 (the transcription factor partner of Yap1) and electroporated these at the same time as FatJ shRNAs. We found that loss of Yap1 or Tead4 at the same time as loss of FatJ produced a normal number of interneurons and therefore rescued the FatJ phenotype.
Around this time, a paper by Cao et al (Genes Dev. 2008 Dec 1;22(23):3320-34) was published reporting the regulation of neural progenitor pools by the Hippo pathway and that dominant repressor forms of Yap1 and Tead produce an increase in the number of Lim1/2 positive cells – the same phenotype we observed after FatJ knockdown! Crucially, the authors did not focus on the upstream signal controlling the hippo pathway, which we believed to be FatJ. To address this, we attempted to determine more directly if loss of FatJ caused a change in the phosphorylation state of Hippo pathway components. This was a time-consuming process involving many electroporations followed by sub-dissection of transfected cells and then western blot analysis with phospho-specific antibodies. Unfortunately, the anti-phosphoMst antibody (the Hippo orthologue) did not work well enough to detect a change in activity of the Hippo pathway. However, we did detect a decrease in the level of phospho-Yap1 after FatJ knockdown and this decrease was also evident by immunohistochemistry of neural tube sections. Therefore, we had confirmation that loss of FatJ causes a decrease in phosphorylation of a downstream hippo component.
This gave us a mechanism for the observed phenotype; FatJ normally acts via downstream Hippo pathway components to limit the size of specific progenitor pools in the neural tube. In the absence of FatJ, these progenitors continue to proliferate resulting in a corresponding increase in the number of the interneurons. Intriguingly, the FatJ-/- mutant mouse phenotype displays a wider neural tube than wild-type littermates suggesting that longer-term loss of FatJ expression could lead to significant tissue overgrowth.
This brought us to the end of a long journey; starting from an RNAi screen and ending with a mechanism. Even though our screen focused on a specific group of genes, we ended with a range of phenotypes – this really highlights the usefulness of the chick as a model system and has proven that RNAi screens are indeed feasible in this system, opening up new possibilities for functional genomics in higher vertebrates.
Van Hateren, N., Das, R., Hautbergue, G., Borycki, A., Placzek, M., & Wilson, S. (2011). FatJ acts via the Hippo mediator Yap1 to restrict the size of neural progenitor cell pools Development, 138 (10), 1893-1902 DOI: 10.1242/dev.064204


 (5 votes)
(5 votes)