Turning lemons into lemonade…sick planarians reveal link between microbiome and regeneration
Posted by Christopher Arnold, on 30 September 2016
Commentary on
Pathogenic shifts in endogenous microbiota impede tissue regeneration via distinct activation of TAK1/MKK/p38
in eLife 2016; 5: e16793 DOI: http://dx.doi.org/10.7554/eLife.16793
During our research as biologists we endure numerous problems, failures, and setbacks in our efforts to advance scientific knowledge. But sometimes the very problems we encounter are opportunities in disguise. This was my experience as a postdoc studying regeneration in the planaria Schmidtea mediterranea in the lab of Alejandro Sánchez Alvarado at the Stowers Institute for Medical Research.
Planarians are free-living flatworms that have the remarkable capability of regenerating any and all of their tissues throughout their seemingly endless adult life (Elliott and Sánchez Alvarado 2013). The negligible senescence and regenerative capabilities of planarians can be traced to an abundant population of persistent pluripotent stem cells that constantly fulfill the tissue homeostasis and regeneration demands of these organisms. This population of cells has the additional feature of enabling amputated tissue fragments containing at least one pluripotent stem cell to regenerate into an entire clonal organism (Wagner, Wang et al. 2011).
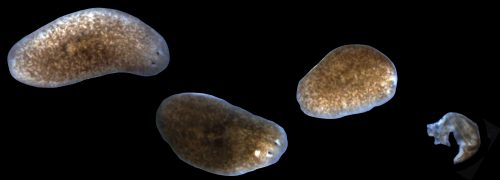
While these properties earned planaria the classic moniker of being “immortal under the edge of the knife” (Dalyell, 1814), our laboratory would receive a potent lesson in planarian mortality. In order to expand our planarian colony, animals were subjected to a vigorous amputation and feeding regimen. Initially this aggressive expansion strategy seemed to be working, but soon it was discovered that the stage had also been set for an equally aggressive bacterial infection. Visible tissue lesions formed and amputated fragments were lysing with increasing frequency (Figure 1). The outbreak transitioned from acute to chronic and in some cases entire boxes of worms amputated one day would be lysed by the next morning. At first, treatment with antibiotics stabilized the colony and kept death at bay, but the propensity for bacteria to evolve antibiotic resistance made this truce a fragile one.
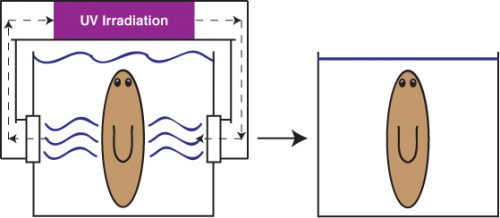
Alejandro sought the assistance of the Stowers Institute for Medical Research Aquatics Facility to develop a long-term solution that could allow us to indefinitely maintain planaria in a stable, low septic environment. The answer was a novel recirculation culture system that utilized constant UV irradiation of recirculating planaria water to prevent uncontrolled bacterial outgrowth (Figure 2). The system was able to rear biomass levels of healthy worms ripe for experimentation. I had just begun my postdoc in the Sánchez lab in the wake of this worm crisis and was excited to receive my first box of worms from this new system.
To my dismay, the very first planarians under my care quickly got sick. As the whole lab would discover, worms withdrawn from the recirculation system were highly susceptible to the development of tissue lesions and degeneration characteristic of sick worms (Figure 1). Just as before, these symptoms could be managed with reduced feedings, antibiotics, and time. Instead of allowing disappointment to gain the upper hand, we saw this as an opportunity to explore the potential role of bacteria and the immune system in animal regeneration.
The role of the immune system in regeneration is an area of intense and ongoing research. Significant evidence spanning a diverse array of organisms and tissues indicated that the immune response can play a central role in either promoting or hindering wound healing and tissue regeneration (Eming, Hammerschmidt et al. 2009, Karin and Clevers 2016). The precise determinants facilitating these diametrically opposed outcomes have important implications in the treatment of chronic non-healing wounds, scar formation, and inflammatory disorders. This reproducible and robust means of inducing tissue degeneration via bacterial infections in planarians struck us as a viable experimental vulnerability to decipher the relationship between microbiota, the immune system, and regeneration in planaria.
We used a two-pronged strategy of bacterial 16s deep sequencing and plating of planaria homogenate to identify bacterial changes after removal of planarians from the recirculation system. Full analyses of the 16s sequencing was not completed until much later in the project. While the deep sequencing strategy was ongoing, culture of planarian homogenate on LB media plates yielded a strain of Pseudomonas that expanded in worms upon removal from the recirculation system. Infection of healthy worms with Pseudomonas was sufficient to induce the tissue degeneration we observed in sick animals. Furthermore, worms infected with Pseudomonas failed to properly regenerate amputated tissues (Figure 3).
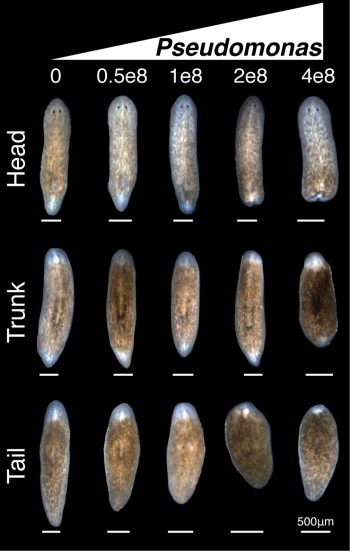
We now had an inducible, highly penetrant, and visibly score-able tissue degeneration phenotype, which combined with the amenability of planaria to RNAi afforded us with the opportunity to dissect genes involved in modulating infection, and to uncover any roles such genes may play in regulating tissue lesions and/or regeneration. Healthy worms directly isolated from the recirculation system were the ideal source to study this process. Unfortunately, our initial attempts were unsuccessful as worms fresh from the recirculation system subjected to the 10 day RNAi feeding protocol developed tissue lesions and degeneration prior to Pseudomonas infection. This presented us with a new challenge: preserving the reduced septic state of the recirculation culture system in a means conducive to the RNAi feeding protocol. To solve this problem, we once again worked with the Aquatics Core of the Stowers. We developed a flow culture system that allows us to house small groups of planaria in individual vessels supplied with a continuous flow of fresh planaria water (Figure 4). This permitted worms removed from the recirculation system to be subjected to the RNAi feeding protocol while preventing the pathogenic bacterial bloom. Worms could then be infected and the results of the gene knockdown could be assayed.
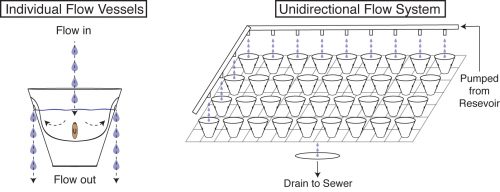
With this new methodology, I performed a candidate screen that quickly identified members of the TAK1/MKK/p38 pathway, a conserved innate immune/inflammatory-signaling module, as the primary mediator of tissue degeneration in response to infection. RNAi of activators of TAK1/MKK/p38 signaling prevented bacterial induced tissue degeneration while RNAi of inhibitors of this signaling pathway accelerated tissue degeneration. Additionally, this innate immune signaling impeded the regeneration of worms during infection, but had no discernible effect on the phenotypic regeneration of uninfected worms (Figure 5).
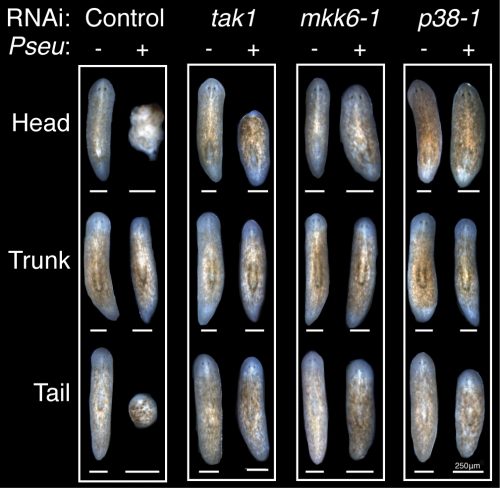
Further molecular analyses revealed a role for TAK1/MKK/p38 in promoting apoptosis in the anterior region of the worm following infection and preceding tissue degeneration. Unexpectedly TAK1/MKK/p38 signaling repressed apoptosis normally coincident with tissue remodeling in un-infected amputated worms. The opposing regulation of apoptosis mediated by TAK1/MKK/p38 in infection versus normal regeneration mirrored the duality of the role of inflammatory signaling in regeneration in other experimental paradigms, suggesting potential cell type specific or context dependent activation underlying its seemingly contradictory roles.
Analyses of the 16s sequencing initiated earlier on in the project was completed and revealed that a high Bacteroidetes to Proteobacteria ratio within the planaria from the recirculation culture system mirrored the distribution of these phyla within the human lower intestinal tract (Human Microbiome Project 2012). This phyla composition is not conserved in other invertebrate (e.g., C. elegans and D. melanogaster) or even vertebrate (e.g., D. rerio) model systems (Lee and Brey 2013, Berg, Stenuit et al. 2016). This was the icing on the cake. The enrichment in Bacteroidetes in planaria was significant as this phylum represents a diverse array of key symbionts known to support proper tissue homeostasis, the development of the immune system, as well as the prevention of inflammatory diseases in mammals, and, intriguingly, have even been linked to the early evolution of multicellular organisms (Rakoff-Nahoum, Paglino et al. 2004, Mazmanian, Liu et al. 2005, Mazmanian, Round et al. 2008, Alegado, Brown et al. 2012). Removal from the recirculation system triggered an expansion in Proteobacteria, including Pseudomonas. Proteobacteria expansion has been linked to a myriad of human inflammatory disorders and coincided with the susceptibility of worms to tissue degeneration (Frank, St Amand et al. 2007, Shin, Whon et al. 2015) (Figure 6). This discovery provided our work with a unique parallel to poorly understood disease-relevant shifts in the human microbiome.
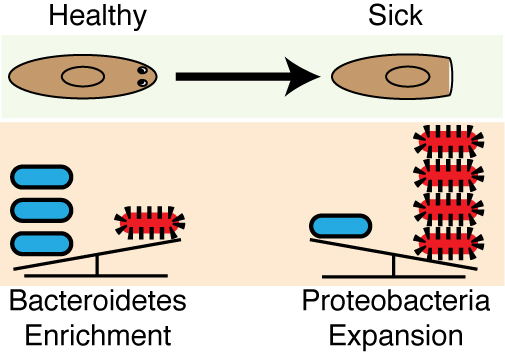
This work began as a systemic problem of worms becoming sick and evolved into a study linking shifts in endogenous microbiota, the innate immune system, and regeneration. Maybe there are very few problems that we as biologist encounter in our research that can become opportunities. Maybe there are many more. But in the end, it is our attitude as curious scientists that will always help us to ultimately turn lemons into lemonade.
Contributors: Christopher Arnold and Alejandro Sánchez Alvarado
Alegado, R. A., L. W. Brown, S. Cao, R. K. Dermenjian, R. Zuzow, S. R. Fairclough, J. Clardy and N. King (2012). “A bacterial sulfonolipid triggers multicellular development in the closest living relatives of animals.” Elife 1: e00013.
Berg, M., B. Stenuit, J. Ho, A. Wang, C. Parke, M. Knight, L. Alvarez-Cohen and M. Shapira (2016). “Assembly of the Caenorhabditis elegans gut microbiota from diverse soil microbial environments.” ISME J.
Elliott, S. A. and A. Sánchez Alvarado (2013). “The history and enduring contributions of planarians to the study of animal regeneration.” Wiley Interdiscip Rev Dev Biol 2(3): 301-326.
Eming, S. A., M. Hammerschmidt, T. Krieg and A. Roers (2009). “Interrelation of immunity and tissue repair or regeneration.” Semin Cell Dev Biol 20(5): 517-527.
Frank, D. N., A. L. St Amand, R. A. Feldman, E. C. Boedeker, N. Harpaz and N. R. Pace (2007). “Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases.” Proc Natl Acad Sci U S A 104(34): 13780-13785.
Human Microbiome Project, C. (2012). “Structure, function and diversity of the healthy human microbiome.” Nature 486(7402): 207-214.
Karin, M. and H. Clevers (2016). “Reparative inflammation takes charge of tissue regeneration.” Nature 529(7586): 307-315.
Lee, W. J. and P. T. Brey (2013). “How microbiomes influence metazoan development: insights from history and Drosophila modeling of gut-microbe interactions.” Annu Rev Cell Dev Biol 29: 571-592.
Mazmanian, S. K., C. H. Liu, A. O. Tzianabos and D. L. Kasper (2005). “An immunomodulatory molecule of symbiotic bacteria directs maturation of the host immune system.”Cell
122(1): 107-118.
Mazmanian, S. K., J. L. Round and D. L. Kasper (2008). “A microbial symbiosis factor prevents intestinal inflammatory disease.” Nature 453(7195): 620-625.
Rakoff-Nahoum, S., J. Paglino, F. Eslami-Varzaneh, S. Edberg and R. Medzhitov (2004). “Recognition of commensal microflora by toll-like receptors is required for intestinal homeostasis.” Cell 118(2): 229-241.
Shin, N. R., T. W. Whon and J. W. Bae (2015). “Proteobacteria: microbial signature of dysbiosis in gut microbiota.” Trends Biotechnol 33(9): 496-503.
Wagner, D. E., I. E. Wang and P. W. Reddien (2011). “Clonogenic neoblasts are pluripotent adult stem cells that underlie planarian regeneration.” Science 332(6031): 811-816.


 (16 votes)
(16 votes)