Using hemimetabolous insects to investigate the origin of the tra-dsx axis
Posted by jrwexler, on 24 September 2019
The story behind our recent paper in eLife.
Rapid turn over of sex determination mechanisms provides biologists with an elegant study system connecting sexual selection to molecular evolution. Striking examples of this turnover are found in African cichlids, where multiple sex determination signals exist not only within the same genera, but sometimes within the same species [1]; in the common house fly, where the primary sex determiner varies along a longitudinal gradient [2], and in the Japanese frog Rana rugosa, a species in which sex chromosomes segregate geographically but whose signals can be overridden by environmental (hormonal) intervention [3].
Downstream of the primary sex determination signal is a developmental pathway that also presents an intriguing puzzle for evolutionary biologists. While the sexual differentiation pathways of mice and humans are fundamentally similar, and while the downstream elements of this pathway are conserved between bees and flies, there are no shared genes in the sex determination pathways of mammals, arthropods, and nematodes with the striking exception of the Doublesex Mab-3 Related Transcription factor (DMRT) family. Furthermore, the fundamental biochemistry of sexual differentiation in these taxa is distinct. In mammals, signaling molecules establish feedback loops that promote one sexual identity and repress the opposite one [4]. In insects, sex-specific RNA splicing determines sexual identity [5]. And in nematodes, a cast of genes including a phosphatase and protease control the sexual fate of a cell [6]. Thus the puzzle: how does a pathway which appears stable on the order of hundreds of millions of years diverge so distinctly between different animal taxa? The Kopp lab was interested in how it is that the sexual differentiation pathway changes on a macro-evolutionary time scale.
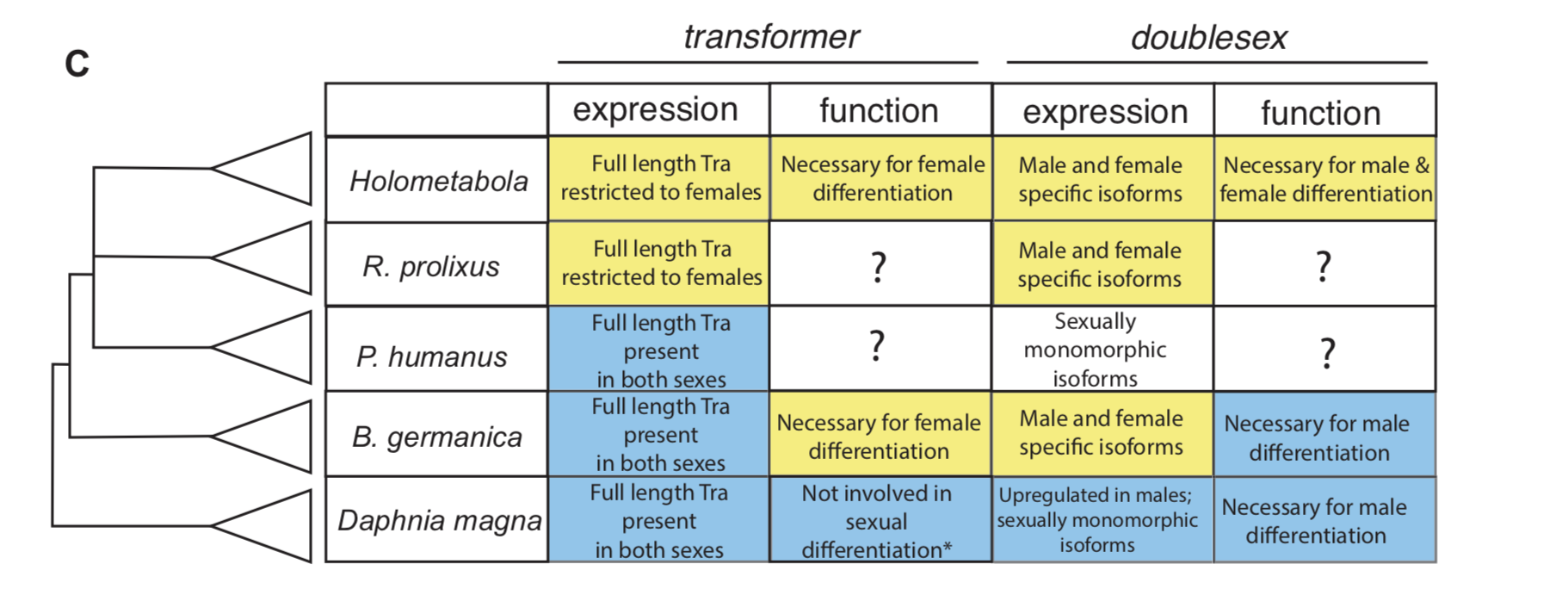
In insect developmental biology Drosophila is the point of reference for all other species. Sex differentiation is no different, where two genes at the bottom of this pathway — transformer (tra) and doublesex (dsx) — were first described in the fruit fly. doublesex (a member of the eponymous Doublesex Mab-3 Related Transcription factor family) is a transcription factor with male- and female-specific isoforms. Both isoforms contain the same DNA binding domain, while alternative, sex-specific splicing at the 3’ end of dsx transcripts changes the effect the male and female dsx isoforms have on their targets [7]. In Drosophila, the male isoform of dsx is produced by default. Active intervention by Tra changes the pattern of dsx splicing from a male to a female isoform. tra itself is also sex specifically spliced, such that a premature stop codon appears in the male, but not the female, transcript [8]. The outgroup to insects with the best studied developmental model is the class Branchiopoda, containing the model crustacean Daphnia magna. In Daphnia, dsx is upregulated in males and lacks male and female isoforms. It is required for male, but not female differentiation [9]. tra is not sex-specifically spliced in this species [10].
To investigate how this insect-specific pathway of sexual differentiation based on sex-specific RNA splicing evolved from an ancestral state without male and female isoforms, we had to look beyond the Kopp lab’s traditional Drosophila domain. Way beyond it. Drosophila-like splicing and function have been documented in Coleopterans and in Hymenopterans, suggesting that the tra–dsx axis of sexual differentiation was present in the common ancestor of Holometabola [11,12,13,14]. The phylogenetic interval between crustaceans and Holometabola is filled in large part by the paraphyletic group of hemimetabolous insects — or those that go through a partial metamorphosis. And because insect pests are more likely to have sequenced genomes, we settled on the following delightful trio of hemimetabolous insects: the kissing bug Rhodnius prolixus (order: Hemiptera), the louse Pediculus humanus (order: Pthiraptera), and the German cockroach Blattella germanica (order: Blattodea). Because R. prolixus and P. humanus are both obligate blood feeders, they cannot be cultured in a lab without strict regulations. Thus, we were reliant on the kindness of Ian Orchard’s lab in Toronto for R. prolixus tissue shipped in RNALater. For P. humanus, our tissue collection was somewhat more exciting — we struck up a relationship with a local louse collector, who is hired to comb nits and lice from the heads of infested humans. In contrast, my experience getting to know German roaches was much more civilized. I was lucky to receive an NSF EDEN grant to study in Barcelona for two months with Xavier Belles, where I learned how to knock down genes in B. germanica. The Belles lab has done extensive work developing B. germanica into an evo-devo model.
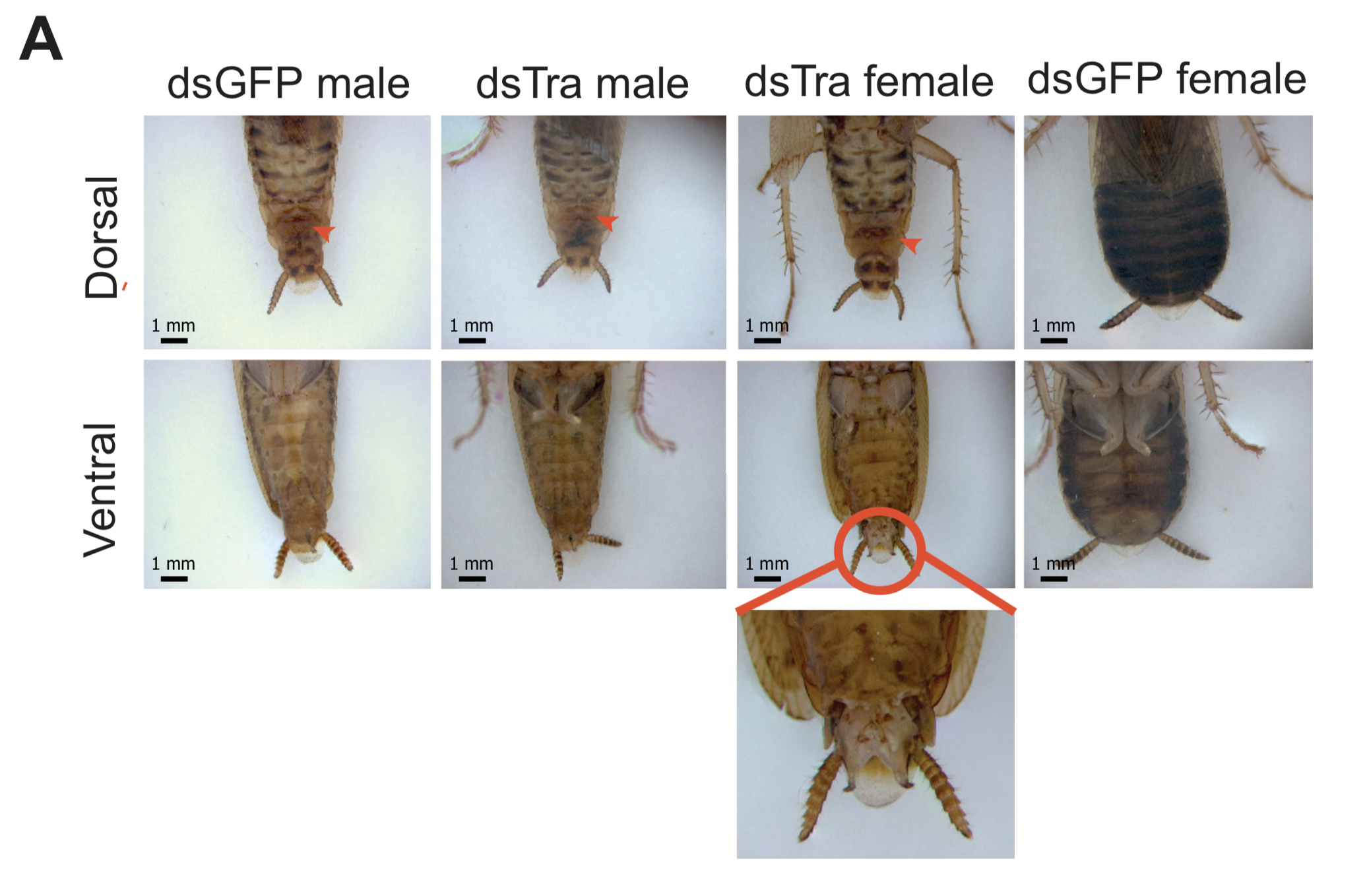

For all three species, we investigated the isoforms of tra and dsx present in males and females. In the German cockroach, we knocked down both genes to probe their functions. The phylogenetic breadth of our samples allowed us to observe the evolutionary trajectory of dsx and tra across a large evolutionary interval. We discovered that the holometabolous pattern of tra splicing, with a male-specific premature stop codon, evolved after tra evolved a functional role as a regulator of female development. Our results from R. prolixus, where we isolated tra with a male specific stop codon, showed that the “canonical” pattern of sex-specific splicing of this gene did evolve before the common ancestor of Holometabola. In contrast, it appears that the familiar holometabolous pattern of male- and female-specific dsx isoforms differing at their 3’ end evolved before dsx became a key regulator of female differentiation. So for each gene, we observed a different story: with dsx, the “holometabolous” splicing pattern evolved before the “holometabolous” functional role; for tra, this pattern was reversed.
The knockdown experiments lead to some very fun collaborations as we attempted to probe the resulting phenotypes. After I returned from Spain and set up my own B. germanica colony at UC Davis, I was reliant on the generosity of Coby Schal, who shared a few roaches from his colony, allowing me to set up my own. The Schal lab specializes, among other things, in the study of insect behavior. When I attended the conference of the International Congress of Entomology with Coby a year later, I excitedly sat down to show him images of the masculinized females we observed upon knocking down tra. Coby then offered to quantify the degree to which these roaches’ behavior was masculinized. The experiments he did with his lab technician Ayako Wada-Katsumata revealed that like in Drosophila, sex-specific behavior of the German roach is under the control of tra. Separately, I began chatting with my teammate from the UC Davis cycling team about the work. This teammate and friend, Matt Amicucci, also happened to be a graduate student in the chemistry department, who then offered to help investigate the oligosaccharides produced by masculinized females in their tergal glands. Thanks to Matt’s expertise, we learned that tra also controls the chemical physiology of German roaches.
The breadth of the questions motivating this project and the number of questions that came out of it; the diversity of skills among the paper’s authors, and the generous support of everyone involved really made this work a wonderful and memorable PhD.
Above: Wild type males respond to the stimulus of a wild type female antenna by raising their wings and presenting their tergal gland.
Above: Females treated with dstra respond as wild type males do to the stimulus of a wild type female antenna. These treated females raise their wings in a stereotypical male courtship display.
[1] Ser JR, Roberts RB, Kocher TD. 2010. Multiple interacting loci control sex determination in lake Malawi cichlid fish. Evolution 64:486–501. DOI: https://doi.org/10.1111/j.1558-5646.2009.00871.x
[2] Hamm R., Shono T., Scott J. G. 2005 A cline in frequency of autosomal males is not associated with insecticide resistance in house fly (Diptera: Muscidae). J. Econ. Entomol. 98: 171–176.
[3] Nakamura M. 2009. Sex determination in amphibians. Seminars in Cell & Developmental Biology 20:271–282. DOI: https://doi.org/10.1016/j.semcdb.2008.10.003
[4] Matson CK, Murphy MW, Sarver AL, Griswold MD, Bardwell VJ, Zarkower D. 2011. DMRT1 prevents female reprogramming in the postnatal mammalian testis. Nature 476:101–104. DOI: https://doi.org/10.1038/ nature10239
[5] Geuverink E., and Beukeboom L.W. 2014. Phylogenetic Distribution and Evolutionary Dynamics of the Sex Determination Genes doublesex and transformer in Insects. Sex Dev 2014;8:38-49 https://doi-org.10.1159/000357056
[6] Zarkower, D. 2006. Somatic sex determination. WormBook, doi/10.1895/wormbook.1.84.1
[7] Coschigano KT, Wensink PC. 1993. Sex-specific transcriptional regulation by the male and female doublesex proteins of Drosophila. Genes & Development 7:42–54. DOI: https://doi.org/10.1101/gad.7.1.42, PMID: 8422 987
[8] Inoue K, Hoshijima K, Higuchi I, Sakamoto H, Shimura Y. 1992. Binding of the Drosophila transformer and transformer-2 proteins to the regulatory elements of doublesex primary transcript for sex-specific RNA processing. PNAS 89:8092–8096. DOI: https://doi.org/10.1073/pnas.89.17.8092, PMID: 1518835
[9] Kato Y, Kobayashi K, Watanabe H, Iguchi T. 2011. Environmental sex determination in the branchiopod crustacean Daphnia magna: deep conservation of a doublesex gene in the sex-determining pathway. PLOS Genetics 7:e1001345. DOI: https://doi.org/10.1371/journal.pgen.1001345, PMID: 21455482
[10] Kato Y, Kobayashi K, Oda S, Tatarazako N, Watanabe H, Iguchi T. 2010. Sequence divergence and expression of a transformer gene in the branchiopod crustacean, Daphnia magna. Genomics 95:160–165. DOI: https://doi. org/10.1016/j.ygeno.2009.12.005, PMID: 20060040
[11] Shukla JN, Palli SR. 2012a. Sex determination in beetles: production of all male progeny by parental RNAi knockdown of transformer. Scientific Reports 2:602. DOI: https://doi.org/10.1038/srep00602, PMID: 22924109
[12] Shukla JN, Palli SR. 2012b. Doublesex target genes in the red flour beetle, Tribolium castaneum. Scientific Reports 2:948. DOI: https://doi.org/10.1038/srep00948, PMID: 23230513
[13] Hasselmann M, Gempe T, Schiøtt M, Nunes-Silva CG, Otte M, Beye M. 2008. Evidence for the evolutionary nascence of a novel sex determination pathway in honeybees. Nature 454:519–522. DOI: https://doi.org/10. 1038/nature07052, PMID: 18594516
[14] Cho S, Huang ZY, Zhang J. 2007. Sex-specific splicing of the honeybee doublesex gene reveals 300 million years of evolution at the bottom of the insect sex-determination pathway. Genetics 177:1733–1741. DOI: https://doi. org/10.1534/genetics.107.078980, PMID: 17947419


 (3 votes)
(3 votes)