AI tackles variability of metastatic conversion triggered by bioelectric disregulation
Posted by Mike Levin, on 7 November 2015
One of the most important problems in experimental biology has to do with variability / heterogeneity (Rubin, 1990): why do different organisms react differently to the same perturbation or reagent? This is observed even among clonal populations (e.g., cohorts of planarian flatworms descended from the fission of 1 animal and living in the same environment), and has huge implications for both efficacy and side-effects of biomedical interventions. Understanding and being able to predict such outcomes requires quantitative, fully-specified models and an analysis of the attractors in the state space describing their dynamics (Davies et al., 2011; Huang et al., 2009). Unfortunately, uncovering such models is very difficult – far more challenging than just describing the epistasis between proteins necessary for a process to occur. We recently addressed this problem by creating a machine learning platform to help infer mechanistic models of complex processes with stochastic outcomes (Lobikin et al., 2015a).
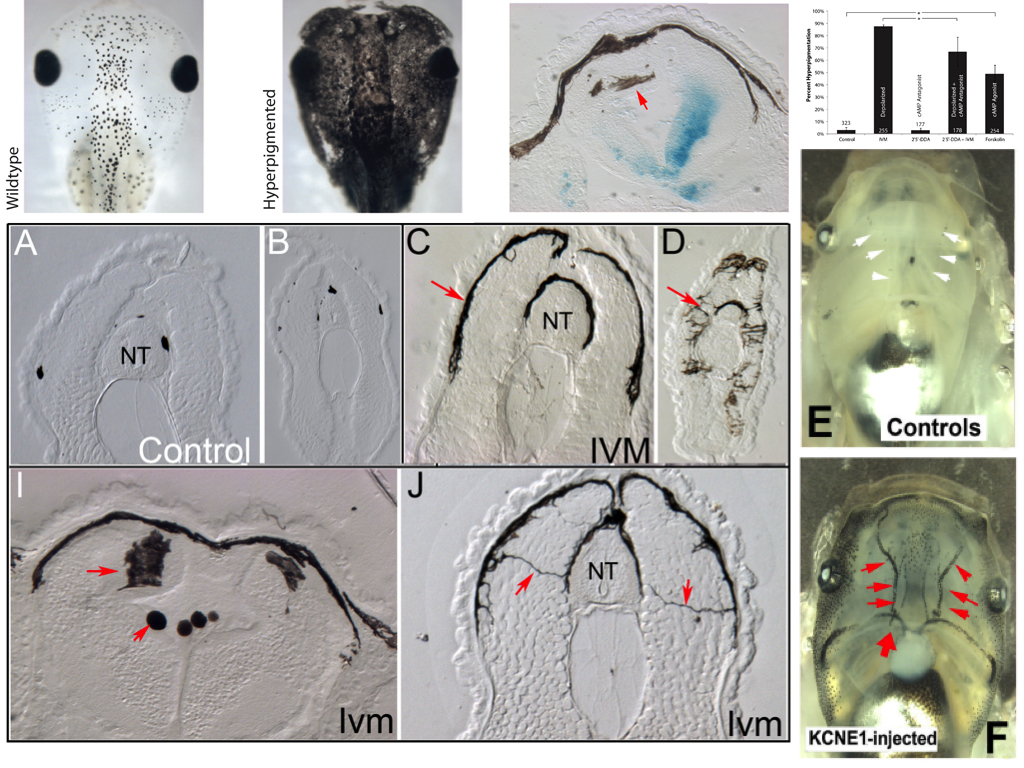
Our lab studies developmental bioelectricity – the mechanisms by which cells (not just neurons) coordinate their activity using voltage gradients (Levin, 2014; Tseng and Levin, 2013). About 10 years ago, we began using the frog embryo to ask what would happen if a small number of widely-distributed somatic cells during embryogenesis was selectively depolarized (resting potential brought closer to 0) (Morokuma et al., 2008). We took advantage of the glycine gated chloride channel, which turned out to be expressed in a ubiquitous but sparse cell population (Blackiston et al., 2011). We used ivermectin, a drug that specifically locks this channel in an open state, to allow negative chloride ions to leave cells down their concentration gradient and thus depolarize that cell population – akin to clonal analysis. To our surprise, the first effect we noticed was not in those cells themselves, but in melanocytes (pigment cells derived from the neural crest). In depolarized embryos, the melanocytes converted to a metastatic-like phenotype: they over-proliferated, changed shape to a drastically-arborized morphology, and invaded inappropriate areas of the body (brain, blood vessels, soft internal organs) in an MMP-dependent manner.
We called the GlyR-expressing cells “instructor” cells, since they were able to change the behavior of a remote cell population, and used a variety of targeting and rescue strategies to show that the effect did occur at long range (was not cell-autonomous). The phenotype recapitulated the metastatic phase of melanoma, but without DNA damage, mutations in oncogenes, or carcinogen exposure (Chernet and Levin, 2013; Lobikin et al., 2012). The bioelectric disregulation of instructor cell state was sufficient to kickstart this process; the phenotype was very clean – the tadpoles developed normally, although later we also found subtle effects on muscle development and vasculature (Lobikin et al., 2015b). The affected tadpoles were not hard to identify: they turned pitch black because of the excess and spread-out shape of the melanocytes which took over their bodies.
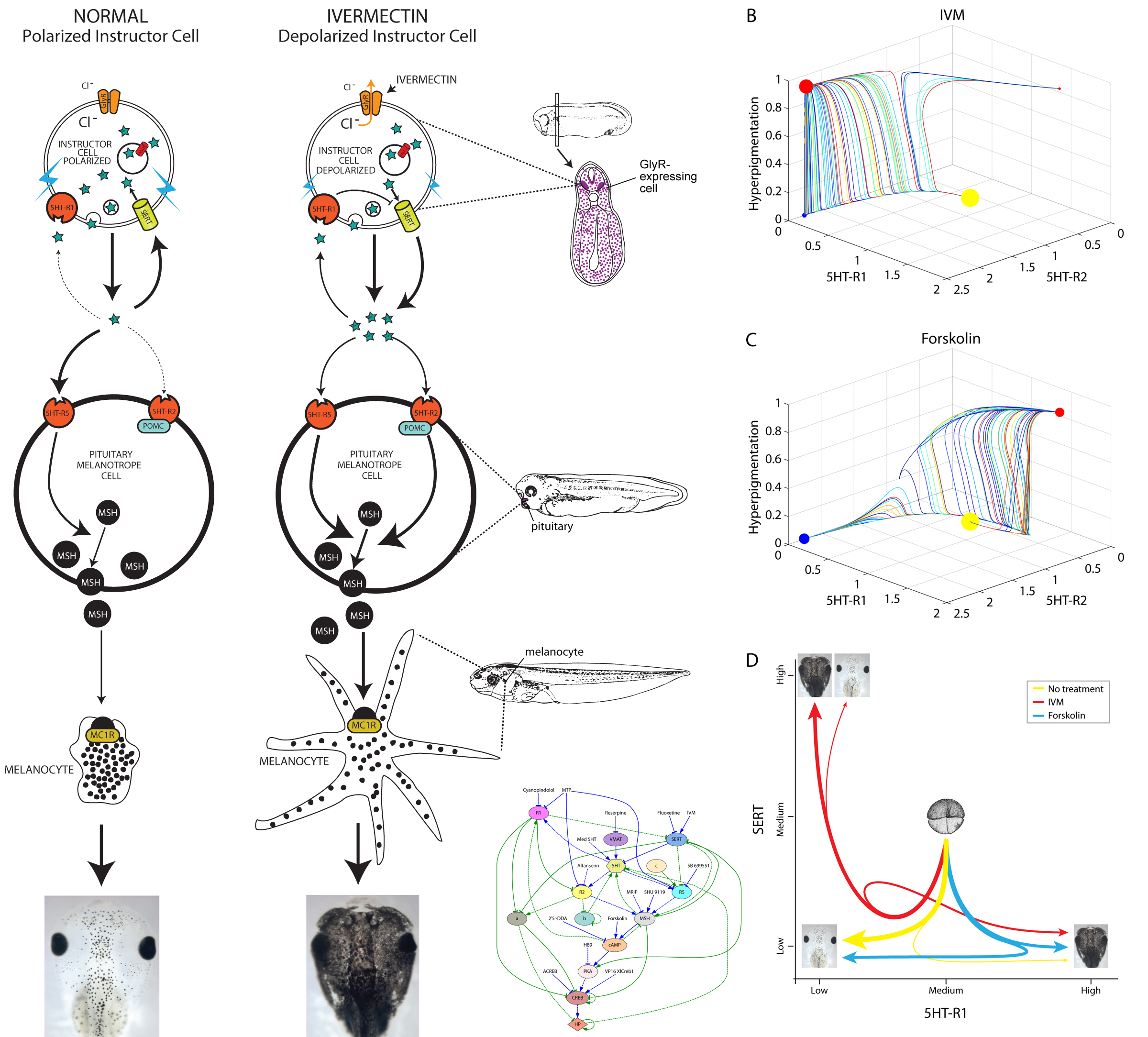
Investigating the mechanism of this effect was relatively straightforward. We dissected the signaling pathway and showed that it relied on a number of components related to serotonergic signaling, via the serotonin transporter (which was under voltage control) and cAMP. We implicated a number of signaling proteins in this cascade, tested their functional relationships with each other, and made the usual “arrow model” of the process (Blackiston et al., 2011). But one aspect remained unsatisfying. The phenotype was all-or-none: we never observed a partially-converted animal. Depending on the penetrance of any given functional treatment, some percentage of the tadpoles would convert (entirely), and some would remain unaffected. It’s as if they were flipping a (biased) coin to decide, but all of the cells in the animal were flipping the same coin. And our arrow diagram model could not predict or explain the precise percentage of melanoma-converted animals that would result from any given experiment. Indeed, the more experiments we did, the harder the problem got, because the dataset that had to be matched by any candidate model was getting more and more complex. This is a pervasive problem in many areas of biology, because the ever-growing mountain of data that is being published makes it ever more difficult for scientists to come up with models that fit the data.
To address this, we turned to a platform we recently designed, using artificial intelligence techniques to help human scientists infer predictive models from published functional data on planarian regeneration (Lobo and Levin, 2015). The system (Lobikin et al., 2015a) used evolutionary computation to search the space of all possible networks comprised of the elements we knew were involved in melanocyte regulation by instructor cells’ voltage. Each network was evaluated against a set of our data, to see if it correctly predicted what percentage of animals would become converted if a specific reagent was used to perturb the pathway. What makes this system powerful is that it does not exhaustively test all possible networks, but uses mutation and a survival of the fittest strategy to home in on the correct answer. The system literally evolved a network specifying the functional connections among the pieces and the strength of each connection, which could correctly recapitulate the complex probabilistic dataset.
Remarkably, not only did it identify a network that correctly explained the data against which we searched, but that same network correctly predicted new experiments it had never seen (which were not present in the training phase). One of the surprises revealed by the discovered model was that there are actually 2 different molecular states that lead to hyperpigmentation. Their resultant phenotype is the same, and would not have been recognized as different based on cell- or tissue-level characterization, but the network model showed that there are two distinct attractors corresponding to the converted state, and thus two molecularly-different ways to reach the same outcome.
This model can now be interrogated for testable mechanistic predictions, and can be used to generate suggested interventions for getting the desired outcome in specific situations. We think this is a proof of principle for using this strategy to derive predictive models matching a complex dataset (with multiple stochastic outcomes for the same input), which could be used by many labs to infer mechanisms from functional data (in basic research), or to identify models matching individual physiological and genetic circuits (for personalized biomedicine approaches). We believe that this is an early step in the creation of the next generation of bioinformatics tools (Lobo et al., 2014; Lobo et al., 2013) – a part of the nascent “robot scientist” field (King et al., 2009; Sparkes et al., 2010), which must augment the efforts of human researchers if we are to glean insights and actionable intelligence from the ever-growing deluge of data.
We welcome collaborations with researchers interested in applying these techniques to their own functional data.
References
Blackiston, D., Adams, D. S., Lemire, J. M., Lobikin, M. and Levin, M. (2011). Transmembrane potential of GlyCl-expressing instructor cells induces a neoplastic-like conversion of melanocytes via a serotonergic pathway. Dis Model Mech 4, 67-85, http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Citation&list_uids=20959630
Chernet, B. and Levin, M. (2013). Endogenous Voltage Potentials and the Microenvironment: Bioelectric Signals that Reveal, Induce and Normalize Cancer. J Clin Exp Oncol Suppl 1, http://www.ncbi.nlm.nih.gov/pubmed/25525610
http://www.ncbi.nlm.nih.gov/pmc/articles/PMC4267524/pdf/nihms621827.pdf
Davies, P. C., Demetrius, L. and Tuszynski, J. A. (2011). Cancer as a dynamical phase transition. Theor Biol Med Model 8, 30, http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Citation&list_uids=21867509
Huang, S., Ernberg, I. and Kauffman, S. (2009). Cancer attractors: a systems view of tumors from a gene network dynamics and developmental perspective. Seminars in cell & developmental biology 20, 869-876, http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Citation&list_uids=19595782
King, R. D., Rowland, J., Oliver, S. G., Young, M., Aubrey, W., Byrne, E., Liakata, M., Markham, M., Pir, P., Soldatova, L. N., et al. (2009). The automation of science. Science 324, 85-89, http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Citation&list_uids=19342587
Levin, M. (2014). Molecular bioelectricity: how endogenous voltage potentials control cell behavior and instruct pattern regulation in vivo. Mol. Biol. Cell 25, 3835-3850, http://www.ncbi.nlm.nih.gov/pubmed/25425556
Lobikin, M., Chernet, B., Lobo, D. and Levin, M. (2012). Resting potential, oncogene-induced tumorigenesis, and metastasis: the bioelectric basis of cancer in vivo. Phys Biol 9, 065002, http://www.ncbi.nlm.nih.gov/pubmed/23196890
Lobikin, M., Lobo, D., Blackiston, D. J., Martyniuk, C. J., Tkachenko, E. and Levin, M. (2015a). Serotonergic regulation of melanocyte conversion: A bioelectrically regulated network for stochastic all-or-none hyperpigmentation. Sci Signal 8, ra99, http://www.ncbi.nlm.nih.gov/pubmed/26443706
Lobikin, M., Pare, J. F., Kaplan, D. L. and Levin, M. (2015b). Selective depolarization of transmembrane potential alters muscle patterning and muscle cell localization in Xenopus laevis embryos. Int J Dev Biol, http://www.ncbi.nlm.nih.gov/pubmed/26198143
Lobo, D., Feldman, E. B., Shah, M., Malone, T. J. and Levin, M. (2014). A bioinformatics expert system linking functional data to anatomical outcomes in limb regeneration. Regeneration, n/a-n/a, http://dx.doi.org/10.1002/reg2.13
Lobo, D. and Levin, M. (2015). Inferring Regulatory Networks from Experimental Morphological Phenotypes: A Computational Method Reverse-Engineers Planarian Regeneration. PLoS Comput Biol in press,
Lobo, D., Malone, T. J. and Levin, M. (2013). Towards a bioinformatics of patterning: a computational approach to understanding regulative morphogenesis. Biol Open 2, 156-169, http://www.ncbi.nlm.nih.gov/pubmed/23429669
Morokuma, J., Blackiston, D., Adams, D. S., Seebohm, G., Trimmer, B. and Levin, M. (2008). Modulation of potassium channel function confers a hyperproliferative invasive phenotype on embryonic stem cells. Proc Natl Acad Sci U S A 105, 16608-16613, http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Citation&list_uids=18931301
Rubin, H. (1990). The significance of biological heterogeneity. Cancer Metastasis Rev 9, 1-20, http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Citation&list_uids=2208565
Sparkes, A., Aubrey, W., Byrne, E., Clare, A., Khan, M. N., Liakata, M., Markham, M., Rowland, J., Soldatova, L. N., Whelan, K. E., et al. (2010). Towards Robot Scientists for autonomous scientific discovery. Autom Exp 2, 1, http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=PubMed&dopt=Citation&list_uids=20119518
Tseng, A. and Levin, M. (2013). Cracking the bioelectric code: Probing endogenous ionic controls of pattern formation. Communicative & Integrative Biology 6, 1-8, http://www.landesbioscience.com/journals/cib/article/22595/


 (5 votes)
(5 votes)