In Development this week (Vol 137, Issue 19)
Posted by Seema Grewal, on 7 September 2010
Here are the research highlights from the current issue of Development:
Nr5a receptors reset EpiSC pluripotency
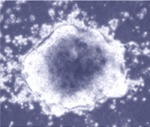 Rodent embryonic stem (ES) cells that are derived from blastocysts self-renew without mitogenic growth factors and robustly colonize chimaeras, whereas egg cylinder-derived stem cells (EpiSCs) require fibroblast growth factor and contribute poorly to chimaeras. Nevertheless, expression of a single reprogramming gene, such as Klf4 or Nanog, can return EpiSCs to a molecular and developmental pluripotent ‘ground state’. Now, on p. 3185, Ge Guo and Austin Smith use a genome-wide genetic screen to identify other molecules that can reprogramme EpiSCs. By using piggyBac transposition to randomly activate endogenous gene expression in mouse EpiSCs and by selecting for undifferentiated colonies in the absence of growth factors, the researchers unexpectedly identify the Nr5a nuclear receptors as potent inducers of ground state pluripotency. Intriguingly, they also show that, unlike previously identified reprogramming factors, Nr5a receptors do not play a role in ES cell renewal. Together, these results highlight the usefulness of EpiSC conversion (in defined culture) as an experimental system for studying molecular reprogramming.
Rodent embryonic stem (ES) cells that are derived from blastocysts self-renew without mitogenic growth factors and robustly colonize chimaeras, whereas egg cylinder-derived stem cells (EpiSCs) require fibroblast growth factor and contribute poorly to chimaeras. Nevertheless, expression of a single reprogramming gene, such as Klf4 or Nanog, can return EpiSCs to a molecular and developmental pluripotent ‘ground state’. Now, on p. 3185, Ge Guo and Austin Smith use a genome-wide genetic screen to identify other molecules that can reprogramme EpiSCs. By using piggyBac transposition to randomly activate endogenous gene expression in mouse EpiSCs and by selecting for undifferentiated colonies in the absence of growth factors, the researchers unexpectedly identify the Nr5a nuclear receptors as potent inducers of ground state pluripotency. Intriguingly, they also show that, unlike previously identified reprogramming factors, Nr5a receptors do not play a role in ES cell renewal. Together, these results highlight the usefulness of EpiSC conversion (in defined culture) as an experimental system for studying molecular reprogramming.
EGFR-Notch signalling makes (proneural) waves
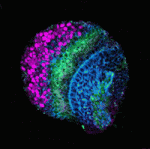 During neurogenesis in the Drosophila optic lobe, a wave of differentiation that converts neuroepithelial cells into neuroblasts sweeps across the neuroepithelial sheet in a medial to lateral direction. This differentiation wave is preceded by the ‘proneural wave’: the transient expression of the proneural gene lethal of scute [l(1)sc]. Now, Tetsuya Tabata and colleagues report that EGFR and Notch signalling play pivotal and coordinated roles in proneural wave progression in the Drosophila optic lobe (see p. 3193). They show that EGFR signalling is activated in neuroepithelial cells and induces l(1)sc expression. Transient, spatially restricted expression of Rhomboid regulates EGFR, they report, and Rhomboid expression is regulated by the EGFR signal, a feedback loop that moves the proneural wave laterally. The researchers also report that Notch signalling, which prolongs the proneural state, is regulated both by itself and by EGFR signalling. Based on these results, the researchers propose that coordinated sequential EGFR and Notch signalling regulates proneural wave progression, which, in turn, induces neuroblast formation in a precisely ordered manner.
During neurogenesis in the Drosophila optic lobe, a wave of differentiation that converts neuroepithelial cells into neuroblasts sweeps across the neuroepithelial sheet in a medial to lateral direction. This differentiation wave is preceded by the ‘proneural wave’: the transient expression of the proneural gene lethal of scute [l(1)sc]. Now, Tetsuya Tabata and colleagues report that EGFR and Notch signalling play pivotal and coordinated roles in proneural wave progression in the Drosophila optic lobe (see p. 3193). They show that EGFR signalling is activated in neuroepithelial cells and induces l(1)sc expression. Transient, spatially restricted expression of Rhomboid regulates EGFR, they report, and Rhomboid expression is regulated by the EGFR signal, a feedback loop that moves the proneural wave laterally. The researchers also report that Notch signalling, which prolongs the proneural state, is regulated both by itself and by EGFR signalling. Based on these results, the researchers propose that coordinated sequential EGFR and Notch signalling regulates proneural wave progression, which, in turn, induces neuroblast formation in a precisely ordered manner.
Hand2 on heart: promoting cardiac fusion
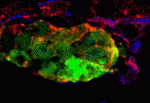 The embryonic heart tube forms from bilateral groups of cardiomyocytes that move towards the embryonic midline where they merge. The transcription factor Hand2 is essential for this ‘cardiac fusion’ but its downstream effectors are unknown. By studying zebrafish heart development, Deborah Yelon and colleagues now identify Fibronectin as a component of the Hand2 pathway that mediates cardiac morphogenesis (see p. 3215). By performing transplantation experiments between wild-type and hand2 mutant embryos, the researchers show that hand2 regulates cardiac fusion by altering the environment through which the cardiomyocytes migrate. Next, they show that fibronectin 1 (fn1) expression is increased in hand2 mutant embryos. Finally, they report that reduction of fn1 function rescues cardiac fusion in hand2 mutant embryos but not the apicobasal polarity defect that is also seen in these embryos. Thus, the Hand2 pathway regulates cardiac morphogenesis by establishing an appropriate environment for cardiac fusion by limiting Fibronectin function but it establishes the apicobasal polarity that is needed for heart tube extension through another, unidentified, effector.
The embryonic heart tube forms from bilateral groups of cardiomyocytes that move towards the embryonic midline where they merge. The transcription factor Hand2 is essential for this ‘cardiac fusion’ but its downstream effectors are unknown. By studying zebrafish heart development, Deborah Yelon and colleagues now identify Fibronectin as a component of the Hand2 pathway that mediates cardiac morphogenesis (see p. 3215). By performing transplantation experiments between wild-type and hand2 mutant embryos, the researchers show that hand2 regulates cardiac fusion by altering the environment through which the cardiomyocytes migrate. Next, they show that fibronectin 1 (fn1) expression is increased in hand2 mutant embryos. Finally, they report that reduction of fn1 function rescues cardiac fusion in hand2 mutant embryos but not the apicobasal polarity defect that is also seen in these embryos. Thus, the Hand2 pathway regulates cardiac morphogenesis by establishing an appropriate environment for cardiac fusion by limiting Fibronectin function but it establishes the apicobasal polarity that is needed for heart tube extension through another, unidentified, effector.
Wise up to Wnt’s role in tooth development
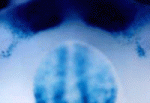 The number, size and shape of mammalian teeth vary widely – just compare a person’s smile with a dog’s ‘smile’. But what controls the patterning of dentition? Mutations in Wise (Sostdc1), which encodes an inhibitor of Lrp5- and Lrp6-dependent Wnt signalling, cause patterning defects in tooth development in mice. Now, by investigating the pathways modulated by Wise, Robb Krumlauf and co-workers show that crosstalk between Wnt and other signalling pathways controls mouse tooth development (see p. 3221). The researchers use genetic experiments to reveal that Wise suppresses the survival of vestigial tooth buds in the normally toothless region between the incisors and molars by inhibiting Lrp5- and Lrp6-dependent Wnt signalling. They also identify the Fgf and Shh signalling pathways as major downstream targets of Wise-regulated Wnt signalling, and show that Shh acts as a negative-feedback regulator of Wnt signalling. Thus, the researchers suggest, variations in the expression of signalling modulators such as Wise could underlie the evolutionary diversity in mammalian dentition.
The number, size and shape of mammalian teeth vary widely – just compare a person’s smile with a dog’s ‘smile’. But what controls the patterning of dentition? Mutations in Wise (Sostdc1), which encodes an inhibitor of Lrp5- and Lrp6-dependent Wnt signalling, cause patterning defects in tooth development in mice. Now, by investigating the pathways modulated by Wise, Robb Krumlauf and co-workers show that crosstalk between Wnt and other signalling pathways controls mouse tooth development (see p. 3221). The researchers use genetic experiments to reveal that Wise suppresses the survival of vestigial tooth buds in the normally toothless region between the incisors and molars by inhibiting Lrp5- and Lrp6-dependent Wnt signalling. They also identify the Fgf and Shh signalling pathways as major downstream targets of Wise-regulated Wnt signalling, and show that Shh acts as a negative-feedback regulator of Wnt signalling. Thus, the researchers suggest, variations in the expression of signalling modulators such as Wise could underlie the evolutionary diversity in mammalian dentition.
Del1-ving into forebrain development
 During early embryogenesis, morphogen gradients specify the neural plate along the anterior-posterior axis. Canonical Wnt signalling causes the posteriorization of neural tissues. Consequently, Wnt signal attenuation in the embryo’s anterior region is required for the determination of the head region; but how is this achieved? On p. 3293, Hidehiko Inomata, Yoshiki Sasai and co-workers reveal that modulation of canonical Wnt signalling by the extracellular matrix protein Del1 (Developmental endothelial locus-1) is essential for forebrain development in Xenopus embryos. Del1 overexpression expands the forebrain domain, the researchers report, whereas Del1 functional inhibition represses forebrain development. They show that Del1 function in neural plate patterning is mediated mainly by inhibition of canonical Wnt signalling downstream of β-catenin. Notably, however, Del1 inhibition of canonical Wnt signalling involves the Ror2 (receptor tyrosine kinase-like orphan receptor 2) pathway, which is implicated in non-canonical Wnt signalling. These data suggest that Del1 promotes forebrain development by creating a local environment that attenuates the cellular response to Wnt signals via a unique pathway.
During early embryogenesis, morphogen gradients specify the neural plate along the anterior-posterior axis. Canonical Wnt signalling causes the posteriorization of neural tissues. Consequently, Wnt signal attenuation in the embryo’s anterior region is required for the determination of the head region; but how is this achieved? On p. 3293, Hidehiko Inomata, Yoshiki Sasai and co-workers reveal that modulation of canonical Wnt signalling by the extracellular matrix protein Del1 (Developmental endothelial locus-1) is essential for forebrain development in Xenopus embryos. Del1 overexpression expands the forebrain domain, the researchers report, whereas Del1 functional inhibition represses forebrain development. They show that Del1 function in neural plate patterning is mediated mainly by inhibition of canonical Wnt signalling downstream of β-catenin. Notably, however, Del1 inhibition of canonical Wnt signalling involves the Ror2 (receptor tyrosine kinase-like orphan receptor 2) pathway, which is implicated in non-canonical Wnt signalling. These data suggest that Del1 promotes forebrain development by creating a local environment that attenuates the cellular response to Wnt signals via a unique pathway.
Extracellular signal PARtners asymmetric division
 Asymmetric cell divisions generate cell diversity during development, and the orientation of the axis of these divisions determines the future position of differentiated cells. But is the asymmetrical localization of the polarity (PAR) proteins that control asymmetric cell division regulated by extracellular or intracellular signals? On p. 3337, Yukinobu Arata and colleagues answer this controversial question. In C. elegans embryos, the P0 zygote and the P1, P2 and P3 germline cells undergo a series of asymmetric divisions. By examining the development of these germline cells in vitro, the researchers show that, although PAR-2 is distributed asymmetrically in P2 and P3 cells in the absence of extracellular signals, the orientation of PAR-2 localization in these cells depends on their contact with endodermal precursor cells. Other experiments indicate that the endodermal precursor cells control the orientation of PAR-2 localization by extracellular signalling via the MES1/SRC1 pathway. The researchers propose, therefore, that Src is an evolutionarily conserved molecular link that coordinates extrinsic cues with PAR protein localization during asymmetric cell divisions.
Asymmetric cell divisions generate cell diversity during development, and the orientation of the axis of these divisions determines the future position of differentiated cells. But is the asymmetrical localization of the polarity (PAR) proteins that control asymmetric cell division regulated by extracellular or intracellular signals? On p. 3337, Yukinobu Arata and colleagues answer this controversial question. In C. elegans embryos, the P0 zygote and the P1, P2 and P3 germline cells undergo a series of asymmetric divisions. By examining the development of these germline cells in vitro, the researchers show that, although PAR-2 is distributed asymmetrically in P2 and P3 cells in the absence of extracellular signals, the orientation of PAR-2 localization in these cells depends on their contact with endodermal precursor cells. Other experiments indicate that the endodermal precursor cells control the orientation of PAR-2 localization by extracellular signalling via the MES1/SRC1 pathway. The researchers propose, therefore, that Src is an evolutionarily conserved molecular link that coordinates extrinsic cues with PAR protein localization during asymmetric cell divisions.
Plus…
KNOX genes: versatile regulators of plant development and diversity
 Plant KNOX homeodomain transcription factors maintain pluripotent stem cells in the shoot apical meristem, and recent studies have uncovered novel roles for the KNOX proteins in sculpting plant form and its diversity, which Angela Hay and Miltos Tsiantis review. See the Review on p 3153
Plant KNOX homeodomain transcription factors maintain pluripotent stem cells in the shoot apical meristem, and recent studies have uncovered novel roles for the KNOX proteins in sculpting plant form and its diversity, which Angela Hay and Miltos Tsiantis review. See the Review on p 3153


 (No Ratings Yet)
(No Ratings Yet)