In Development this week (Vol. 138, Issue 1)
Posted by Seema Grewal, on 7 December 2010
The first issue of 2011 is out now…here are the highlights:
Geminin control of lineage commitment

The transition between pluripotency and multi-lineage commitment during early embryogenesis must be closely regulated to ensure correct spatial and temporal patterning of the embryo. But what regulates this crucial transition? According to Kristen Kroll and co-workers, part of the answer to this question in Xenopus embryos lies with the nuclear protein Geminin (see p. 33). The researchers show that Geminin overexpression represses many genes associated with cell commitment but increases the expression of genes that promote pluripotent and immature neuroectodermal cell fates. Geminin, they report, represses Activin-, FGF- and BMP-mediated cell commitment. Consistent with this finding, Geminin knockdown enhances commitment responses to growth factor signalling and results in ectopic mesodermal, endodermal and epidermal fate commitment in the embryo. The researchers also report that repression of commitment by Geminin depends on Polycomb repressor function, and show that Geminin promotes Polycomb-mediated repressive histone modifications of mesodermal genes. The researchers propose, therefore, that cooperativity between Geminin and Polycomb plays an essential role in controlling spatial and temporal patterning in early embryos.
Rock-ing between AP and LR axes
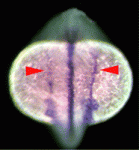
The vertebrate body plan features a left-right (LR) asymmetry, but how the LR axis is orientated correctly with respect to the anteroposterior (AP) and dorsoventral (DV) axes is not known. Here, Jeffrey Amack and co-workers (p. 45) report that the Rho kinase Rock2b links AP patterning to LR patterning in zebrafish embryos. During development, Kupffer’s vesicle (KV) generates a cilia-driven leftward fluid flow that directs LR patterning. The authors demonstrate that depletion of rock2b in whole embryos or in the KV cell lineage alone disrupts asymmetric gene expression during development and perturbs organ asymmetries. They show that, in control embryos, ciliated cells are distributed asymmetrically along the AP axis of the KV and generate asymmetric fluid flow. By contrast, rock2b knockdown embryos show defective KV patterning and cell morphology, and a loss of directional flow. Based on their studies, the authors propose that Rock2b is required for the AP positioning of ciliated cells within the KV and for subsequent LR patterning in zebrafish embryos.
Mesp2 Notches up somite polarity

Somites, the most obviously segmented structures in vertebrate embryos, are subdivided into anterior (rostral) and posterior (caudal) compartments. Repression and activation of Notch signalling are essential for the establishment of the rostral and caudal compartments of the somite, respectively. The mechanism by which Notch is repressed has remained elusive but, on p. 55, Yumiko Saga and colleagues identify the bHLH transcription factor Mesp2 as a novel negative regulator of Notch signalling in mouse somites. In the absence of Mesp2, somites are completely caudalised but, intriguingly, the researchers now show that the introduction of a dominant-negative form of Rbpj (a downstream effector of Notch signalling) into the Mesp2 locus largely rescues the segmental defects of Mesp2-null mice. They also report that Mesp2 represses Notch signalling independently of its function as a transcription factor by inducing the destabilisation of mastermind-like 1, a core regulator of the Notch signalling pathway. These new findings shed light on the molecular mechanisms that control the rostrocaudal patterning of somites.
Cdx1: refining the hindbrain
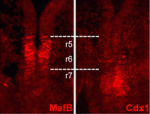
During embryogenesis, the vertebrate hindbrain is segmented along its anteroposterior axis into lineage-restricted compartments, known as rhombomeres (r1-r8), that dictate subsequent neural patterning. The signals that pattern the hindbrain are known, but how each rhombomere-specific gene expression pattern is established is unclear. On p. 65, Sabine Cordes and colleagues reveal that the homeobox protein Cdx1 patterns the mouse hindbrain by spatially restricting the expression of the transcription factor MafB. Mafb is required for r5 and r6 development, and its expression is restricted to these segments. The authors report that the Mafb enhancer contains candidate Cdx-binding sites, and that Cdx1 binds to these sites both in vitro and in vivo. They show that Cdx1 is expressed at the r6/r7 boundary, at the posterior limit of the Mafb-expressing domain. Importantly, in the absence of Cdx1, MafB expression extends beyond its normal r6/r7 boundary. The authors propose that Cdx1 acts as an early and transient repressor of Mafb, and thus plays a role in refining hindbrain identity.
Boc: novel roles in Shh regulation
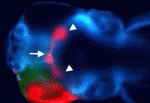
Hedgehog (Hh) signalling gradients control many developmental processes and are influenced by numerous positive and negative regulators. The transmembrane protein Brother of Cdo (Boc) has been implicated in Sonic hedgehog (Shh)-mediated commissural axon guidance in the CNS, but how Boc affects the cellular Hh response in vivo is unclear. Here, Rolf Karlstrom and colleagues reveal that Boc is cell-autonomously required for Hh-mediated ventral CNS patterning in zebrafish (see p. 75). The umleitung (uml) zebrafish mutant is characterised by defects in retinotectal projections. The researchers show first that uml encodes Boc. Then, by analysing the phenotypes of uml mutants, they show that Boc is a positive regulator of Hh signalling in the spinal cord, hypothalamus, pituitary, somites and upper jaw, but that Boc might be a negative regulator of Hh signalling in the lower jaw. Overall, these results reveal a role for Boc in ventral CNS cells that receive high levels of Hh, and uncover novel roles for Boc in vertebrate development.
Expanding the zebrafish toolkit

The zebrafish genetics toolkit has been missing a particularly handy piece of kit: a promoter to drive ubiquitous transgene expression throughout development, equivalent to the Rosa26 locus used in mouse genetics. But no longer, for in one of Development‘s inaugural Technical papers (p. 169), Leonard Zon and co-workers report that the zebrafish ubiquitin (ubi) promoter can drive constitutive transgene expression throughout development. The authors initially identified ubi in BLAST searches using human ubiquitin. They then tested a 3.5 kb 5′ region upstream of its translational start site for transcriptional regulatory sequences and found that it drives strong and ubiquitous EGFP expression within 4 hours of injection into a single-cell embryo. Moreover, in stable ubi-EGFP transgenic lines, EGFP is strongly expressed in all external and internal organs they analysed, in all blood cell types, and from embryo to adulthood. The authors also created inducible ubi-driven CreERt2 transgenes and loxP lineage-tracer transgenes that give strong reporter activity upon Cre exposure, which further enhances and expands the zebrafish transgenesis toolkit.
To find out more, and to read the first author’s “behind the scenes” account of this work, see the related post on the Node
Plus…
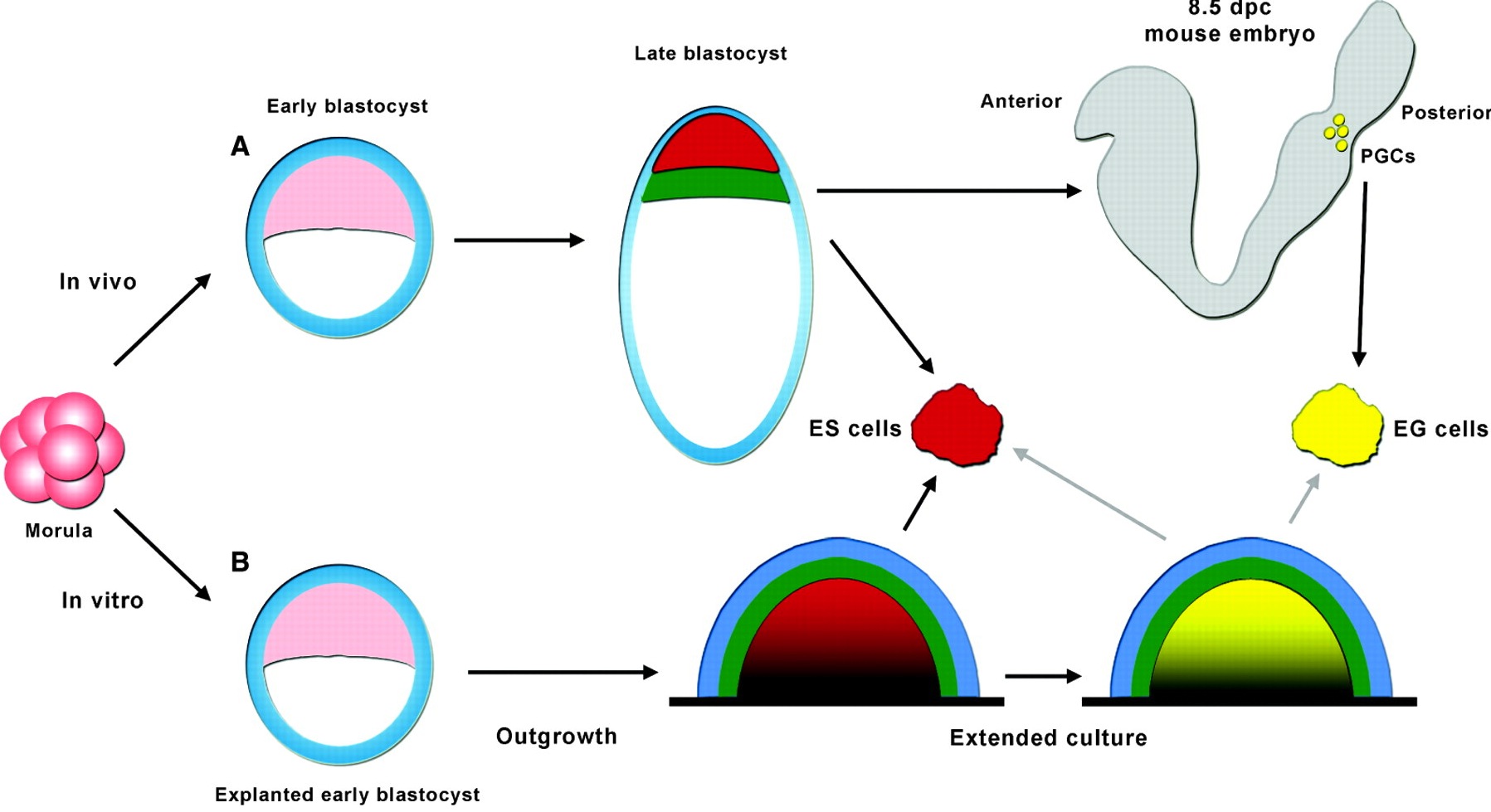
The origin of ES cells has been debated in recent years. Jenny Nichols and Austin Smith now propose that there are, in fact, two possible routes by which ES cells can arise that are dictated by culture conditions.
See the Hypothesis article on p. 3
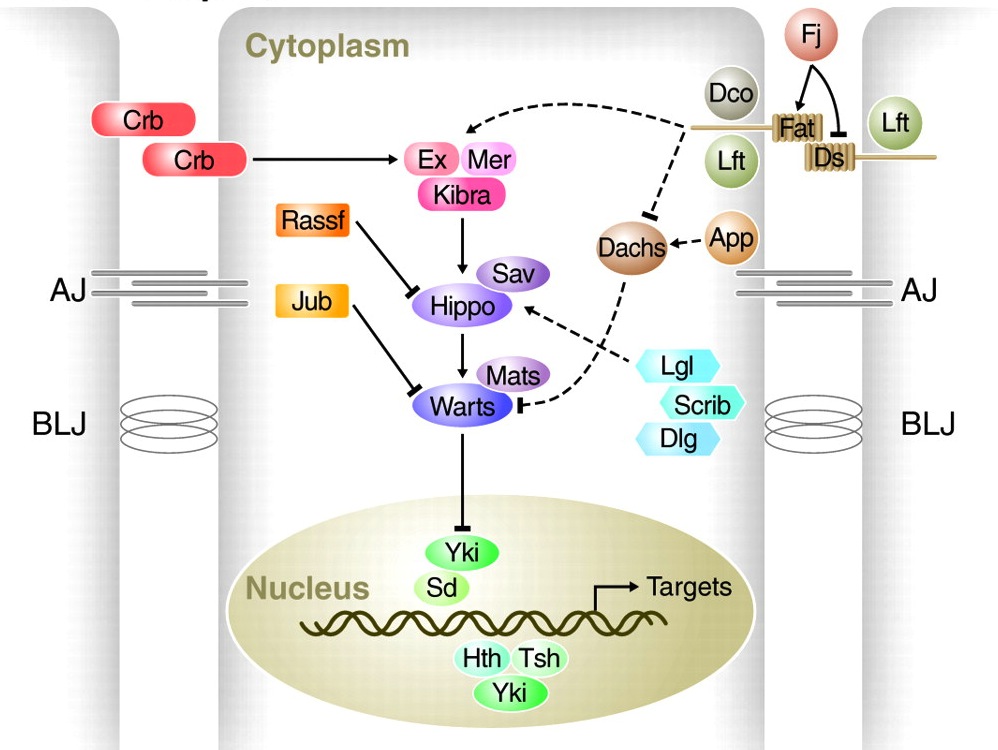
The Hippo pathway regulates growth in Drosophila and vertebrates, and, as Georg Halder and Randy Johnson now discuss, recent studies have shed light on how it governs organ size control and regeneration, and on how it is dynamically regulated during development.
See the Review article on p. 9.


 (No Ratings Yet)
(No Ratings Yet)