The people behind the papers: Thanh Vuong-Brender & Michel Labouesse
Posted by the Node Interviews, on 30 March 2017
This year marks the centenary of D’Arcy Thompson’s On Growth and Form, an attempt to outline the physical and mathematical principles underpinning the generation of biological form. Modern day developmental biologists, bolstered by new technologies, have taken up Thompson’s cause to try to understand the mechanics of development, particularly with regard to morphogenesis. While the generation of forces by the actomyosin cytoskeleton has received a lot of attention, how the material properties of developing tissues influence morphogenesis is less well understood. Today’s paper was recently published in eLife, and investigates the relationship between forces and tissue stiffness in the elongation of the C. elegans embryo. We caught up with lead author Thanh Vuong-Brender and her supervisor Michel Labouesse of the Institut de Biologie Paris-Seine, to hear the story behind the work.

Michel, can you tell us your scientific biography and the questions that your lab is trying to answer?
ML I did my undergrad in Maths/Physics, but chose to do a PhD in Genetics, which appealed to the mathematical neuron I had. I fell in love with C. elegans through a series of seminars by Sydney Brenner – I like the concept of the lineage – and I went to get worm training with Bob Horvitz at MIT. Initially interested by cell fate specification, I rapidly moved to analyse epithelial morphogenesis, and progressively realized that I could thereby feed my second physically-oriented neuron.
Broadly speaking, in the lab we want to understand how mechanical forces impact on cellular processes. Indeed, a cellular phenotype corresponds to its global fate and its 3D organisation; so the challenge is to understand how mechanical forces can modify gene expression programs and/or cell shape determinants, which are defined by junction and cytoskeleton organisation, plus trafficking. Addressing these issues is not as trivial as it may look. At the molecular level, one can think that identifying the structure that senses the force, and the signal transduction that can next modify cell fate or shape should be enough. But it is unlikely to be so. First, the effect or forces is rarely an isolated one-time event, but is often repeated such that the question of timing/periodicity becomes more central than for a chemical signal. Second, understanding how a force can have an effect generally requires thinking, not (only) in biochemical terms, but chiefly in physical terms. Entities to be considered should be energy, entropy, elasticity.
And Thanh, how did you come to work in Michel’s lab?
TV-B I was trained as a physicist. My PhD was about the use of automated imaging and fluorescent markers for diagnosis of cervical cancer. After that, I went to work for a small company that developed automated imaging systems. I realised that I was more interested in academic research to explore and understand natural phenomena. During my PhD, I learnt some biology but not as much as I wanted. So I looked for a postdoc during which I could learn more about biology and came across Michel lab’s papers on mechanical problems of C. elegans embryonic elongation. I found the subject attractive, maybe because it presented to me a mechanical problem to solve. I did not understand all the biology but I thought it was really interesting to learn and to work on it. I sent my postdoc application to Michel and was really lucky to be accepted.
In an interview with Current Biology in 2005, you said you were excited by the challenge of understanding the mechanics of development. I wonder what you think of the progress the field has made in the 12 years since then?
ML The field has evolved tremendously, in part due to progress in imaging and data processing, and in part because a new generation of scientists with strong background in physics has entered the area. In my field, papers that brought key paradigmatic changes, which in retrospect seem quite common sense, include the demonstration that a morphogenetic event requires small increments that progressively modify the cell (Martin et al, Nature; Rauzi et al, Nature; Solon et al, Cell). Another one is that two apposed tissues with distinct mechanical properties will twist (Savin et al, Nature).
You write that the material properties of developing tissues have received less attention than the forces that act on them. Why do you think this is?
ML There are two probable reasons: in vertebrate embryos, the field has more frequently focused often on global movements (although the issue of stiffness has been pointed out more than 30 years ago in Xenopus), whereas in fly embryos, the field has focused on processes dependent on cells having a homogenous behaviour.
TV-B Deformation depends both on forces and material properties, so in theory, one can have as many regulatory pathways of shape formation through the regulation of material properties as through forces. The role of forces has been intensively investigated through the studies of non-muscle myosin and its regulatory pathways. Many studies have suggested the role of anisotropic material properties, like the elongation of Drosophila developing eggs or trachea. The role of material properties has received more attention in plants, but studies and mechanical measurements of material properties in animal morphogenesis are scarce. Our aim was to bring attention to this important parameter.
What makes the C. elegans embryo a good model for developmental mechanics?
ML The worm is very simple and many of its past successes have been linked to cell biology (PAR proteins, netrin, centrosome assembly, apoptosis, EGF/Ras signalling, to name a few). Its only drawback is that the embryo is quite small, very fragile and not easily amenable to approaches available in other species.
TV-B C. elegans embryonic elongation is very simple and different from other animal models, since it does not involve cell division or cell migration, but is mostly driven by cell shape changes. The number of cells is limited so that one can investigate at the cell resolution and the whole embryo. Other advantages are the easy genetics and worm cultivation.
Can you give us the key results of the paper in a paragraph?
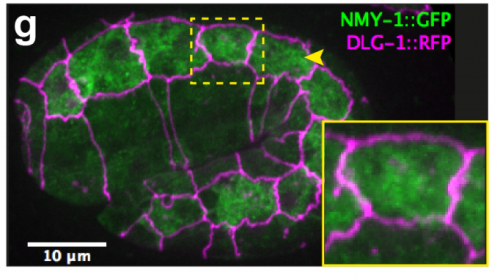
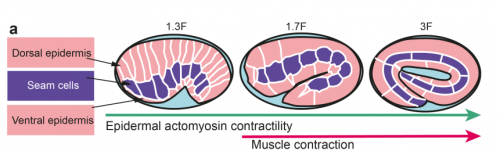
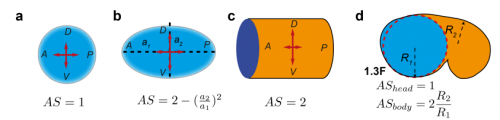
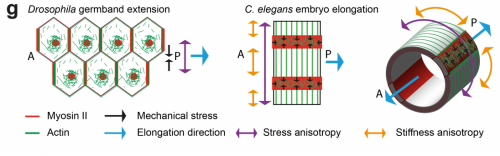
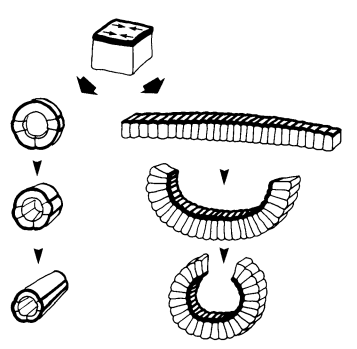
TV-B In the last step of embryonic development, the embryos of C. elegans transform from a ball of cells into the characteristic cylindrical shape of a worm. This process is powered by the association of the molecular motor myosin II, and the actin cytoskeleton in the embryonic epidermis. The epidermis is made up of six strips of cells running along the head-to-tail axis. Myosin II is mostly active in two strips of cells on the two sides of the embryo (lateral cells), but has low activity in the upper and lower strips of cells (dorsally and ventrally to the lateral cells). It is unclear how this distribution of myosin causes embryos to elongate only along the head-to-tail axis. Using laser nano-ablation, we have probed the forces exerted and the material properties in the embryonic epidermal cells. The results show that myosin’s activity in the lateral cells induced constriction around the embryo, sort of similar to the effect of a boa constrictor tightening around its prey. At the same time, the actin filaments in the dorsal and ventral strips form rigid bundles oriented along the circumference. They form a “belt” preventing the constriction from causing the cells at the dorsal and ventral strips embryo to expand. Finally, the only direction the embryo can elongate is along the head-to-tail axis.
How important was the modelling to complement and reinforce your experimental data?
ML It was critical in two ways. First to guide the interpretation of experimental data; second as suggested in the answer to the first question, because understanding a process involving a mechanical force should include a physical dimension whenever possible.
TV-B The mathematical modelling helps us to understand mechanistically what we observed. It can also predict the behaviours of the system in another situation or the role of another component in the process. So it can be your initial hypothesis or feedback, which combines with the experiment data to make the work evolve.
Are the anisotropies of stress and stiffness established by particular signals in the cell, and could this be this linked to cell fate choice in the embryo?
ML These are key mostly unsolved questions for the future.
TV-B I think that the anisotropies of stress and stiffness are linked to the cell specific myosin/actin regulator distribution, which have been shown to be different between lateral and dorso-ventral epidermal cells. It is likely to be linked to the lateral/dorso-ventral cell fate, but remains to be proved.
When doing the research, was there a particularly exciting result or eureka moment that has stayed with you?
TV-B I did not really have a “eureka”, but for me, everyday is like an adventure. There were problems (technical or theoretical) to solve and challenges to overcome because we always tried new things. For more routine stuff, I tried to make it better or quicker. There are small victories like “yes ! it (laser ablation) works”, “I got my CRISPR knock-in strain”, “ the experiment data matches (the theoretical one))”… which led me through the days and frustrations sometime. Well, like every scientist, I hope to have a “bigger eureka” in the future.
And what about the flipside: any moments of frustration or despair?
TV-B Yes, I was trying desperately to make optogenetically-controlled gene expression work in worms. I have tried 3 different systems, none of these worked. Finally I tried a different optogenetic method controlling protein aggregation, but did not have the time to finish it. One consolation is that my preliminary results were used by someone else. I told myself that the failures are part of the quest and became now more resilient to them.
Where next for you following this work?
TV-B I am trying to shift (again) to another domain of research. During my postdoc, my interest for microbiology, symbiosis between microbes and the origin of eukaryotes has grown. So I decided to go to study the microbial diversity. It will be totally different but surely I will learn a lot of things.
And what is the next step for the Labouesse lab?
ML I am quite proud of the work we did with Thanh and our two other co-authors, as I think it accounts in physical terms for a big part of the early phase of elongation. Incidentally, I want to pay tribute to a visionary landmark paper of our field written in 1986 by Jim Priess, a PhD student at the time, who had foreseen almost everything in early worm elongation. Now, I want to account for the second phase of C. elegans elongation that requires the mechanical input of muscle contractions (which Jim had not touched upon), and want to reach a similar level of understanding. We are nearing this phase.
Thanh Thi Kim Vuong-Brender, Martine Ben Amar, Julien Pontabry, Michel Labouesse. 2017. The interplay of stiffness and force anisotropies drive embryo elongation. eLife 23866
Browse the People behind the Papers archives here


 (No Ratings Yet)
(No Ratings Yet)