The people behind the papers – You Wu & Mineko Kengaku
Posted by the Node Interviews, on 12 March 2018
Neuronal migration is critical for mammalian brain development. In many migrating neurons, the nucleus translocates from the trailing to the leading edge of the cell in a manner dependent on the actin and microtubule cytoskeletons, but how these cytoskeletons interact and their relative contribution to the forces that move the nucleus has remained unclear. This week we feature a paper published in the latest issue of Development that uses live imaging to understand this process, revealing some fascinating cellular behaviours. We caught up with first author You Wu and his supervisor Mineko Kengaku, Professor at the Institute for Integrated Cell-Material Sciences at Kyoto University, to find out more about the story.

Mineko, can you give us your scientific biography and the questions your lab is trying to answer?
MK We study how neurons are built up into the beautiful architecture of the brain cortex. A major effort of my lab has been devoted to live-imaging of growing neurons for a mechanistic understanding of brain formation, using the clear and simple cerebellar cortex as a model. We especially focus on the mechanisms and dynamics of cytoskeletal organization during migration of granule cells and dendrite arborization of Purkinje neurons, the two major events during the formation of the three-layered cortex of the cerebellum. I did my PhD study on the molecular mechanism of the primary axis formation in the Xenopus CNS, under supervision of Dr Harumasa Okamoto at the University of Tokyo. I wanted to further study the molecular signals regulating morphogenesis, and joined the laboratory of Cliff Tabin at Harvard Medical School as a post-doc. His lab had just discovered Sonic hedgehog as a bona fide morphogen in the chick embryo. I was extremely lucky to be present in the lab at moments when long-standing questions were clarified, such as the mechanisms of left-right asymmetry, bone formation, fore- and hind limb differentiation, etc. As a consequence of this rapid progress, many fundamental questions in morphogenetic signals had been clarified by the time I started my own lab at the RIKEN Brain Science Institute. The major interests in developmental biology then shifted toward stem cells and disease mechanisms. But I was still fascinated with animal morphology, and therefore decided to look into the beautiful shapes of neurons in the well-organized cortex of the brain.
And You, how did you come to be involved with this project?
YW I was attracted by the beautiful morphology of neurons and the structure of the cortex, and joined the Kengaku Lab as a Master’s student. I first worked on hippocampal neurons to seek how they achieve their unique dendritic shape by live-imaging. The dynamics of development really excited me. Around the time I completed my M.S. degree, I saw a movie of nuclear rotation in a migrating neuron, and was simply interested in the phenomenon.
What was known about the role of the cytoskeleton in CGC nuclear migration before your work?
MK & YW Unlike mesenchymal cell migration, neurons typically form a long leading process and move the nucleus into the leading process in a saltatory manner independently of the constant leading process extension. Since the discovery of an evolutionary conserved cytoplasmic dynein complex protein LIS1 as a responsible gene for smooth brain syndrome (type 1 lissencephaly), accumulating evidence has supported the model that the nucleus is pulled forward by dynein motor activity along the microtubules extended from the leading process. Subsequent studies have also implicated the contractile force of actomyosin in nuclear movement, although competing hypotheses for the site of actomyosin force generation have been proposed using different neuron models.
CGC migration has provided an in vitro model for cell biology study of neuronal migration ever since the first time-lapse observations in 1980s. Recent studies from the groups of Xiao-bing Yuan and David Solecki have provided strong evidence that actomyosin localized in the proximal leading process exerts the driving force for nuclear migration in CGCs. However, the precise interplay and relative contribution of actomyosin and microtubules during nuclear migration have remained to be elucidated.
Can you give us the key results of the paper in a paragraph?
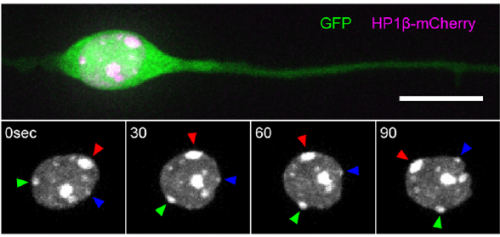
MK & YW For better understanding of the complicated interplay between microtubule- and actin-dependent mechanisms driving nuclear migration in CGCs, we performed a high spatiotemporal resolution imaging using spinning-disk confocal microscopy. We found that the nucleus displayed unexpectedly dynamic motion during migration, including fast rotation in the soma. By quantitative 3D image analysis combined with molecular and pharmacological manipulations, we demonstrated that the nuclear rotation is driven by the same strain energy inducing nuclear translocation. Furthermore, among the two major cytoskeletal forces driving nuclear migration, rotation is solely dependent on microtubule systems but not actomyosin, while nuclear translocation requires both actomyosin and microtubules. We thus used nuclear rotation as a readout of microtubule-dependent tensile stress during nuclear migration, which led to the second major finding that both dynein and kinesin-1 are involved in the force transmission. Our results suggest that microtubules and associated bidirectional motors exert force to small points on the nuclear envelope via interaction with nesprins, and steer rotation or translocation depending on the position of the force points.
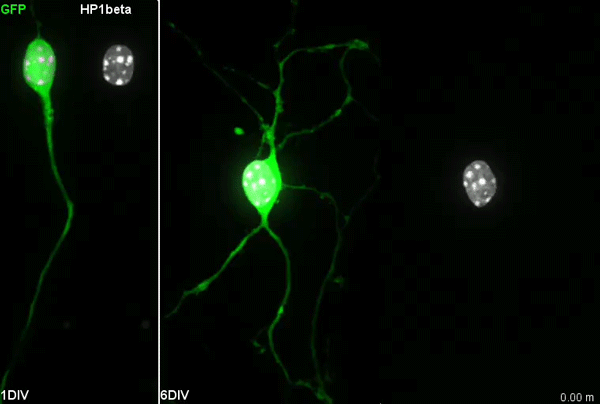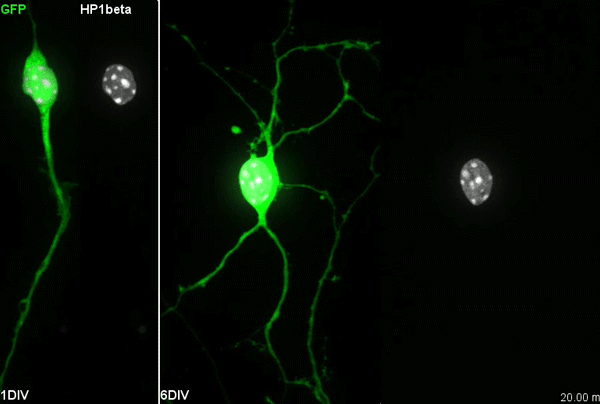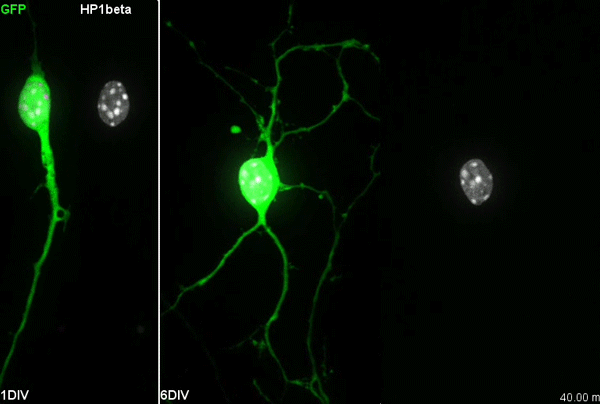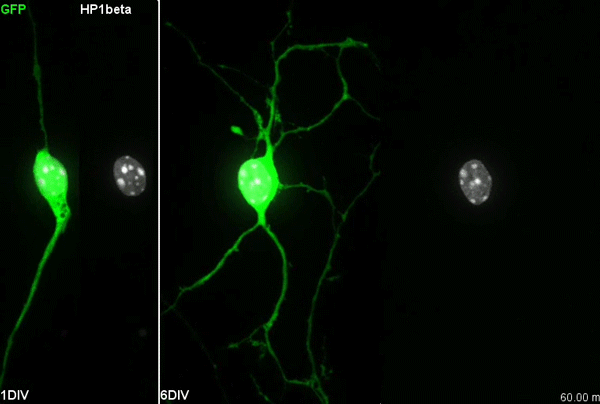
I have to say that the videos in your paper are remarkably beautiful! Can you tell me what it was like to first see nuclear rotation in migrating CGCs?
MK Our co-author Hiroki Umeshima is highly skilled in live imaging, and he set up most of the imaging systems used in the present study. His first movie of nuclear rotation caught my eye, but I did not begin the present study for more than a year, as I was unsure if the rotation is a biologically significant event or an idling motion caused by excessive force to the nucleus in culture. But, the more movies I watched, the more confident I became that the rotation was driven by active forces on small points of the nucleus during nuclear migration. Around that time, You had developed his skill in image analysis, so I asked him if he was interested in analyzing this mysterious phenomenon, and he said ‘Yes’.
YW It was really interesting. I never imagined the nucleus would rotate so fast and frequently during migration. The movies also showed the dynamic deformation of the nucleus. I reckoned these motions could be indicating force transmission, and started to find a way to analyze the dynamics.
How do you think the opposing microtubule motors dynein and kinesin work together to drive unidirectional nuclear motion?
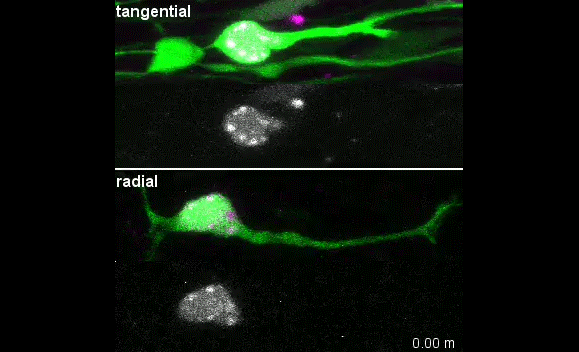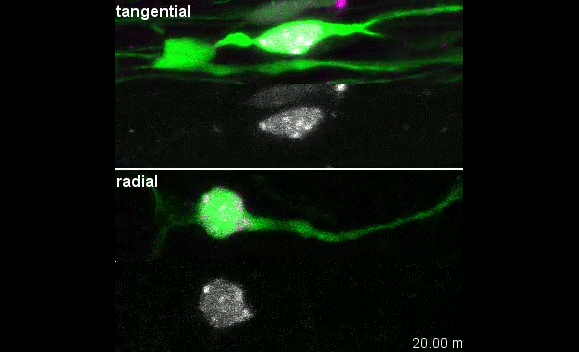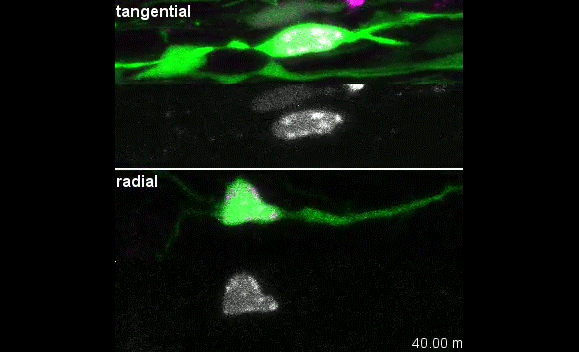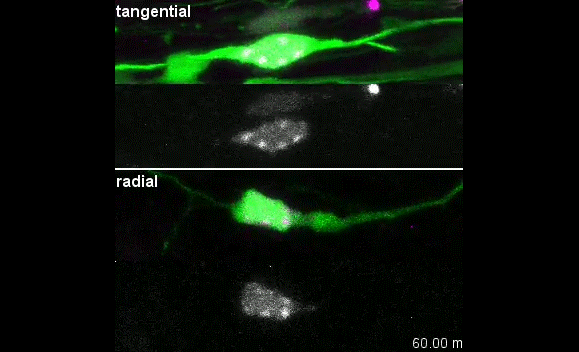
MK & YW We observed frequent switches of rotation direction both in vivo and in vitro (Movie 4 and 5), indicating that the nuclear surface readily moves in both anterograde and retrograde directions by dynamic interplay of the bidirectional motors on bipolar microtubules. Given that the polarity of perinuclear microtubules is mixed but strongly biased, dynein probably acts as the predominant motor and moves the nucleus along the retrograde microtubules (with minus-end toward the leading process) as previously indicated, while kinesin drives back-step movement or rotation. Involvement of bidirectional motors in nuclear migration along uniformly oriented microtubules has been found in long hypodermal precursor cells in nematode embryos (Bone et al., Development 2016; Fridolfsson and Starr, Dev. Biol. 2010). Both hypodermal cells and CGCs are elongated and move nuclei through narrow interstitial and tissue spaces, where dynamic bi-directional movement by opposing motors might adjust the precise positioning of the nucleus and help it squeeze through constrictions.
When doing the research, did you have any particular result or eureka moment that has stuck with you?
I was excited to observe the dynamics of the nucleus in the organotypic slice culture. Until then, I was focused on the in vitro culture system, where the rotation and deformation occurs sporadically. I found the nucleus in 3D tissue much more dynamic, and there was almost no moment when the nucleus stayed still. It was interesting to see that forces seemed to be always applied to the nucleus, even if it was not translocating
And what about the flipside: any moments of frustration or despair?
While it was an exciting moment to see the nuclear motion in tissue, it was very challenging to analyse it quantitatively. It would be interesting and informative to characterize its properties in tissue precisely, but it was difficult to acquire enough resolution in live-imaging.
What next for you, You?
It is always fun for me to observe dynamic things. I am still interested in neural development and also in brain function, which is achieved by such well-organized structure, and am thinking about doing postdoc in related fields.
Where will this work take the Kengaku lab?
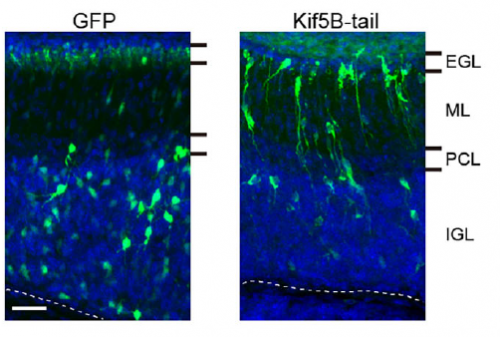
We want to further elucidate the actual crosstalk of microtubule motors and nuclear envelope during nuclear motion. New imaging techniques with higher spatiotemporal resolution might be required to pursue the next problem. Also, the dynamic deformation and rotation shown in the present study suggest that the nucleus is very soft and flexible in newborn neurons. We are interested in how the mechanical properties of the nucleus is determined, and if they contribute to neuronal migration.
Finally, let’s move outside the lab – what do you like to do in your spare time?
MK I must confess that I do not have so much spare time for special activities after research and family. I am a food fan and enjoy cooking every day. I sometimes eat-out with my family and friends on weekends. Luckily Kyoto is a great place to explore nice restaurants of various styles, from traditional Japanese Kaiseki to casual French bistro.
YW I like to sing songs, sometimes in a bit loud voice if nobody is by.
Nesprins and opposing microtubule motors generate a point force that drives directional nuclear motion in migrating neurons. You Kure Wu, Hiroki Umeshima, Junko Kurisu, Mineko Kengaku. Development 2018 145: dev158782
This is #37 in our interview series. Browse the archive here.


 (2 votes)
(2 votes)