Rare is Everywhere
Posted by Michal Shoshkes-Carmel, on 5 June 2018
The story behind FOXL1+ telocytes
You can find our recently published Nature paper here
Our story began two decades ago when my mentor, Klaus H. Kaestner, identified and cloned the transcription factor FOXL1, as being expressed in the mesenchyme of the mouse fetal gut (Kaestner et al. 1997). The position of FOXL1+ mesenchymal cells in such close proximity to the developing epithelium, as evidenced from In-Situ hybridization studies, suggested that FOXL1-expressing cells might be prime candidates to be key signal givers to the developing and adult gastrointestinal tract.
The early studies focused on the role of FOXL1 itself in regulating the intestinal epithelia by deriving mice homozygous for a FOXL1 null allele (FOXL1 -/-) (Perreault et al. 2001; Katz et al. 2004). FOXL1 null mice exhibit increased epithelial proliferation along with increased activation of the Wnt/β-catenin pathway, linking FOXL1 to Wnt/β-catenin pathway regulation.
The breakthrough came with a change in thinking. Klaus first realized that in order to study the role of FOXL1+ CELLS we should employ FOXL1 as a marker to trace these cells and genetically ablate them. The Kaestner lab derived two mouse models to kill FOXL1+ cells through the use of diphtheria toxin administration; FOXL1hDTR BAC transgenic mice that express the human diphtheria toxin receptor from a 170kb bacterial artificial chromosome that harbors all regulatory elements to direct transgene expression in subepithelial telocytes and Rosa-iDTR mice generated by crossing FOXL1Cre mice to a strain that produces the diphtheria toxin receptor from the ubiquitous Rosa26 locus in a Cre-dependent manner (Buch et al. 2005, Sackett et al. 2007).
Inducible ablation of FOXL1+ cells in adult mice caused a dramatic disruption to the intestinal epithelium, loss of stem and progenitor cell proliferation and the experimental mice died a few days after loss of telocytes had been initiated, demonstrating that FOXL1+ cells play a major role in stem cell function (Aoki et al. 2016).
I joined the Kaestner lab for my post-doctoral training during the time when these cell ablation models where being characterized in details. I am trained as a developmental biologist and therefore studying the potential cross talk between epithelia and mesenchyme was of great interest to me, even though at the time very little was known about the nature and function of FOXL1+ cells. In fact, we had no way to detect FOXL1 protein in tissue sections or biochemically, as multiple attempts at obtaining anti-FOXL1 antibodies by commercial outfits had been unsuccessful.
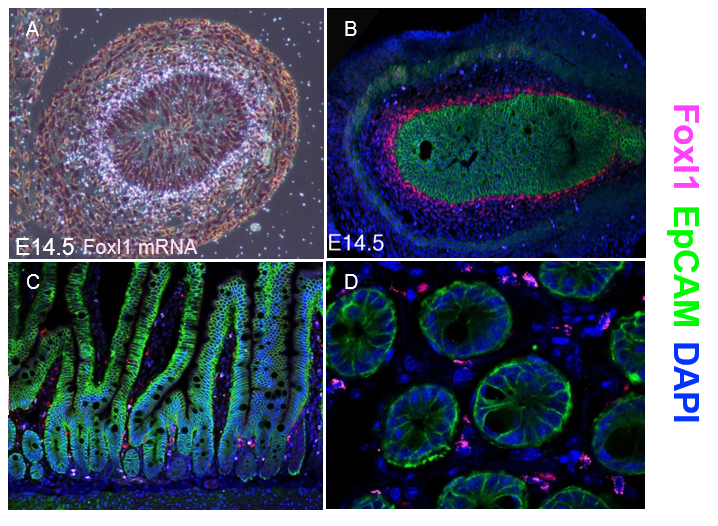
Thankfully, Chris V. E. Wright’s lab at Vanderbilt University came to our aid and generated multiple monospecific anti-FOXL1 antibodies for us. Antibody staining of mouse fetal gut showed a protein expression pattern for FOXL1 that was very similar to the one seen two decades earlier using radioactive In-situ hybridization to detect FOXL1-mRNA (Kaestner et al. 1997) (Figure 1 A-B).
However, the immunostaining for FOXL1 protein in the adult mouse intestine was disappointing at first. FOXL1 protein was present in the nuclei of selected mesenchymal cells; However, the abundance was very low (Figure 1C-D). On average, there were two to three FOXL1+ cells per crypt and their location was at mid-crypt region and along the villi core, in-addition to the crypt base where the stem cells reside.
It is well known that the driving force for intestinal stem cell function is the Wnt/β-catenin pathway, which acts as a short range signaling-system. Potential cell types that might provide Wnt ligands should be in close contact with the stem cells. Do FOXL1+ cells touch stem cells? Do FOXL1+ cells express Wnts? Do they provide the essential Wnts that maintain stem cell identity? These were the key questions that I set out to answer.
With these questions in mind, I met Chris Wright for the first time in person, at the Gastrointestinal tract FASEB meeting in August 2015. During our little discussion, Chris mentioned “cytonemes”, cellular projections that are specialized for exchange of signaling molecules, and suggested that I investigate FOXL1+ cell structure. If FOXL1+ cells have long extensions and/or posses a unique cell structure, this might allow them to contact all epithelial cells.
Back in the lab, I had a clearer understanding as to which steps I should take –
- I planned to inhibit all Wnt secretion from FOXL1+ cells and ask: Are stem cells affected?
- I needed to sort GFP-labeled FOXL1+ cells (using a FOXL1-Cre ;Rosa-YFP mice) and prepare RNA seq libraries. Determine their gene expression profile: Do FOXL1+ express Wnts, and if so which ones?
- I had to label FOXL1+ cells with a membrane reporter so that I could determine the extent of FOXL1+ cell structure
Targeting secretion of all 19 mammalial Wnt proteins can be done by deleting either Wntless or Porcupine, two essential Wnt processing enzymes, for which floxed mutant mice were already available. In order to inhibit Wnt secretion from adult and not fetal FOXL1+ cells, I need an inducible-FOXL1 driven Cre. With the help of fellow postdoc Kirk Wangensteen, I built a FOXL1-CreERT2 BAC and generated a new transgenic mouse line.
The next challenge arose when I tried to sort GFP-labeled FOXL1+ cells. To be able to sort FOXL1+ cells I had to selectively digest the mesenchyme from the intestinal epithelium in a single cell suspension. It was difficult to determine the optimal conditions to digest the mesenchyme as harsh digestion killed FOXL1+ cells, while mild digestion did not liberate any GFP-positive cells. Together with fellow postdoc Yue Wang, we devised a strategy to enable the selection to be successful. We isolated FOXL1+ cells, made RNA seq libraries and submitted them for sequencing.
The last piece of the puzzle was determinning FOXL1+ cell structure. To achieve this goal, I crossed our FOXL1 Cre mice to the Rosa-mTmG reporter in which FOXL1 promoter driven Cre activity lead to the expression of a membrane-bound version of GFP, which labeled the plasma membrane and allowed me to see the full extent of FOXL1+ cell.
The results were impressive; FOXL1+ GFP labeled cells were very large in extent and thus in contact with the entire epithelium from crypt base to the tip of the villi, with each and every single epithelial cell being touched by FOXL1+ mesenchymal cells!
During the same week I determined FOXL1+ cell structure, the RNAseq data was returned from sequencing. FOXL1+ cells indeed express a specific subset of Wnts and also the Wnt pathway inducers, R-Spondins. However, FOXL1+ cells also made high levels of Wnt inhibitors as well as BMPs, which are known to oppose Wnt signaling. How could this be, as we had shown through cell ablation that critical Wnt signals emanate from FOXL1+ cells?
I reasoned that since I had sorted FOXL1+ cells from anywhere along the crypt-villus axis for my RNAseq study, FOXL1+ cells might compartmentalize expression of signaling molecules based on their specific position along the crypt-villus axis.
To test this hypothesis, I contacted Shalev Itzkovitz from the Weizmann Institute, who had optimized a single molecule mRNA-Fluorescence In-Situ Hybridization (smFISH) technique for the mouse intestine. My goal was to focus on mRNA localization of different signaling molecules in FOXL1+ cell projections along the crypt-villus.
For this, I had to devise a way to label the full extent of FOXL1 cells, not just their nuclei. Unfortunately, I could not employ my Foxl1Cre Rosa-mTmG mice, as the reporter has a global tomato fluorescent protein expression, which bleeds through all analysis channels, thereby interfering with the smFISH signal. FOXL1 antibody staining would also not work, as it would label only the nuclei. What I needed was to identify a surface marker that could be specifically used to label the cells.
The RNAseq data revealed high expression of platelet derived growth factor receptor α (PDGFRα), member of the “villus cluster genes” characterized previously during gut development (Walton et al., 2012, Shyer et al., 2013, Shyer et al., 2015, Walton et al., 2016). Does PDGFRα label FOXL1+ cells? And if so, would it be possible to use it as a marker to label FOXL1+ cell extensions? YES! We demonstrated that all FOXL1+ cells are PDGFRα+.
In Shalev’s lab, Beáta Tóth combined PDGFRα immunostaining with smFISH and we were able to show regional differentiation in mRNA localization of different signaling molecules along FOXL1+ projections, with FOXL1+ cells near the crypt bottom producing abundant Wnt2b, a canonical Wnt pathway activator, while those further up the crypt-villus axis expressed high levels of Wnt pathway inhibitors.
I was puzzled by the unusual structure of FOXL1+ cells, which I also confirmed by electron microscopy. These cells are extremely thin but very large, with diameters in excess of 200 micrometer (for comparison, an intestinal epithelial cell is only about 10 micrometer in size). I was wondering if such unique stromal cells had been described in the literature, based on histological technologies.
My literature search came up with several reports by Popescu (the late eminent Romanian pathologist) and Faussone-Pellegrini (Popescu et al., 2005, Popescu and Pellegrini 2010, Cantarero et al., 2011, Cretoiu et al. 2012, Vannucchi et al., 2013) describing primarily through the use of electron microscopy, the existence of a new stromal cell type that is present in many organs.
Popescu named these cells “Telocytes” from the Greek words “telos” meaning end, “cytes” meaning cells. Telocytes are cells characterized by extremely long and thin projections called telopodes that may reach millimeters long and express PDGFRα in both human and mouse gut.
Apparently, neurons are not unique; telocytes also have long extensions that make direct contact with each other. Would it be possible to use the neuroscientists’ technology to study the 3D network of these cells? X-CLARITY is a method designed by neuroscientists for clearing tissues in order to visualize neurons in their 3D structure within the brain without the need for sectioning (Chung and Deisseroth 2013). Could we apply this technique to clear the intestine and visualize telocytes in their 3D structure?
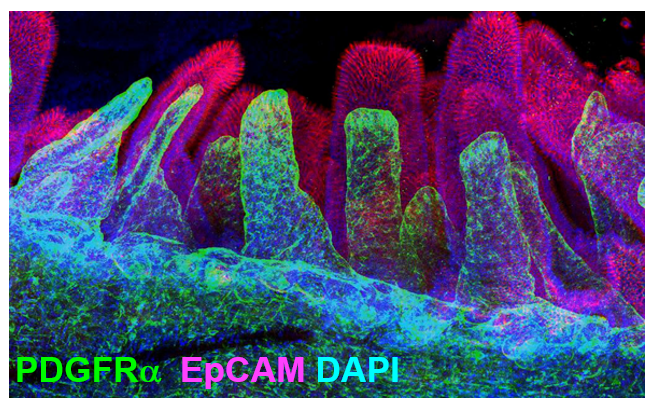
In fact, clearing whole intestine and immunostaining for PDGFRα in green and EpCAM to label epithelial cells in red, allowed me to visualize the comprehensive stromal network of telocytes that form a plexus that supports the entire epithelium (Figure 2).
My journey has just begun, and many exciting questions remain: Does FOXL1 label all telocytes? What is the origin of this remarkable cell? How and when do cells acquire telocyte characteristics? How do these cells compartmentalize signaling? What are the mechanisms by which telocytes signal to the epithelium.
The next decade of research will be a lot of fun!


 (3 votes)
(3 votes)