The people behind the papers – Guillaume Blin, Manuel Thery & Sally Lowell
Posted by the Node Interviews, on 21 September 2018
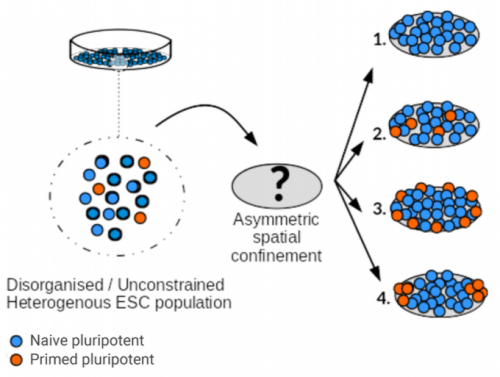
During early mouse development, a series of signalling interactions breaks the symmetry of the egg cylinder, spatially organising the embryo into territories that define the future axes of the body. Symmetry breaking can also be observed in embryonic stem cell (ESC) colonies cultured on micropatterned substrates, which thus provide a powerful system to test the role of signalling as well as other features of the cell population in early patterning. A recent paper in Development uses micropatterned colonies to test the role of geometry in symmetry breaking. We caught up with three of the paper’s authors – Guillaume Blin, Manuel Thery and Sally Lowell – to hear all about the history.

Guillaume, can you give us your scientific biography and the questions that drive your research?
GB I did my undergrad in the south of France where I was trained as a biochemist and biophysicist. I spent one of my first lab experience with Prof. Catherine Picart (Material biophysicist) and Dr. Christian Roy (Cell Biochemist) in the DIMNP lab in Montpellier. I worked on a biomimetic system to model and quantify the interaction between the protein ezrin (known for its role in cell polarity) and PIP2 (a constituent of the plasma membrane). This work was truly interdisciplinary and as I built skills in good old biochemistry, I also had the chance to learn about techniques more alien to the biologist world I came from – zeta potential, DLS, FCS, QCM, this was fascinating.
Catherine, Christian, as well as Prof. Andrea Parmeggiani (theortical physicist and teacher who changed my understanding of bio-molecular interactions), succeeded in creating a synergistic environment spiced with intrinsic ‘cultural differences’ between actors which meant that the coffee table was often the scene of some of the most hilarious banter between physicists and biologists. I am very grateful to them as this experience gave me the will and confidence to keep on building my own interdisciplinary skill set later on.
A question that came out of this work is how macroscopic phenomenon emerge as a result of microscopic events and for this reason I next changed system scale and went on to study cell biology in the lab of Prof. R. Foisner as an Erasmus student in Vienna. Again I am grateful to the people who have trained me Dr. Andy Brachner and Dr Josef Gotzmann. I don’t think I would have adapted well to what would come next if I had not spent such a great time with them in the Max Perutz lab. My Viennese experience coincided with the publication of seminal papers from the D. Discher and C. Chen lab showing that the fate of stem cells could be largely influenced by the physical properties of their environment.
And so, I then started a PhD with Catherine who had elaborated a collaboration with stem cell scientist Dr. Michel Puceat, in order to test the influence of stiffness on the cell fate of embryonic stem cells. During my PhD, I was also involved in another project aiming at modeling cardiac development with ESC to isolate a cardiac progenitor population to use as a source for cell therapy. This jump from purely basic science to a more translational approach has also brought its load of teaching and I am grateful to Michel for introducing me to this important dimension of research.
While working with ESC I noticed that even a clonal population of cells would create a wide diversity of behaviors in the dish and at that time the literature was flourishing with examples of heterogeneous expression of genes associated with cell fate biases. As a result, the next question I became interested in is how can we reconcile the propensity of stem cells to diversify in a seemingly erratic fashion in the dish with the remarkable reproducibility and robustness of developmental patterning in vivo.
Inspiring work came out from the lab of P.W Zandstra who was using micropatterns to dissect endogenous signaling in ESC and the role of the micro-environment. This gave me the idea that we may be able to guide patterning with geometrical confinement. I was also very impressed with Manuel’s work and so I contacted Manuel to learn how to make micropatterns. Manuel was even more enthusiastic and supportive than what I could have hoped for. I learned a lot with him and I must say, his good mood and energy are very contagious, beware! The first preliminary results were obtained as early as 2010. However, this work also coincided with the end of my PhD and the work was held back by the writing of my thesis and also by some technical challenges I was facing (notably in imaging).
Motivated by the will to better understand the origin of the heterogeneous response of stem cells to differentiation signals, I then joined the lab of Dr. Sally Lowell in the MRC-Center for Regenerative Medicine in Edinburgh. I was awarded a 4-year Wellcome Trust post-doctoral fellowship in order to determine how changes in tissue geometry may feedback onto early cell fate decisions. As part of my post-doc project, I developed new image analysis tools to quantify patterning in complex 3D cell populations at single cell resolution. With this new computational tool in hand, we decided to resume the story initiated about 6 years earlier. Sally has been incredible at providing support to ‘resurrect’ this project. Like the dream PI for a post-doc, she provided the pitch-perfect environment to reveal the hidden potential of this work, a bit like geometry in our study!
So there I stand now, looking to establish my own group in order to continue identifying general principles about how the cells self-organise using synthetic and quantitative methods.
Sally, what are the main aims of the Lowell Lab, and what led you to investigate the role of geometry in patterning?
SL I’m fascinated by how cells talk to each other to generate patterns. We think that when these conversations are disrupted, for example through changes in adhesion and tissue morphology, this can help to reinforce differentiation decisions to make development more robust. Paradoxically these same mechanisms may well be confounding our ability to direct differentiation in cells in culture. My wonderful and creative lab members are working hard to understand how all this is happening. When Guillaume joined the lab he set to work revolutionising the way we were able to approach these questions. For example, thanks to a set of image analysis tools that Guillaume has developed we can now measure cell-cell interactions and changes in morphology at single cell resolution even within enormous (to us) post-gastrulation mouse embryos. Watch this space for a couple more of Guillaume’s papers which will report these tools to the community. The idea to investigate the role of geometry came entirely from Guillaume, who is the brains and the driving force behind this project. Indeed he very presciently began this project eight years ago during his PhD, taking inspiration from the work of Manuel Thery. Manuel will be able to tell you more about how it all began…
And how did you become involved with the project Manuel?
MT I was working on cell architecture and developing micropatterning methods to play with cell shape and adhesion. Meanwhile I was reading the papers by Christopher Chen who has always been a great source of inspiration for me. He was constraining cell groups in crazy shapes where mesenchymal stem cells were differentiating toward adipocytes or osteoblast depending on their position in the group. Osteoblasts appeared at the periphery while adipocytes sat at the center of the colony. In a very thoughtful experiment he plated cells on sinusoidal avenues. All cells were close to the periphery, but osteoblast appeared along convex edges while adipocytes appeared in concave edges. The proof was shown that stem cells sense geometrical cues and not only the local cell density. I was super excited by this results and used it a lot as an example of the importance of geometry to cell physiology. But I could not work on it. In 2010 I was starting my group with two students and we were focusing on intra-cellular architecture. It was very frustrating. Then Guillaume run into my office and proposed to study the role of geometry in ES cell colony! I jumped on my chair like the wolf facing Little Red Ridding Hood in the nightclub in Tex Avery’s cartoons. We immediately started to draw shapes that could contradict the density dogma. The issue was that Guillaume was not equipped to perform microfabrication experiments in his lab. We had to set up a new protocol. Guillaume worked a lot and got very enthusiastic with what I would consider pretty unsatisfying results. So he insisted more and more, tried new chemicals and got it to work. From there I started reading a lot on ES cells but it was too complex to me. Guillaume was visiting me and teaching me mouse embryonic development. I learned so much! It was really great. I became fascinated by the gastrulation and started working on epithelial-to-mesenchymal transition.
What was known about the role of geometry in early mammalian patterning prior to your work?
GBGeometry is known to have a strong influence during the 1st cell fate decision, when the blastomeres become segregated either to the ICM or to the TE. Geometry dictates cell polarity which in turns define the activity of key transcription factors. This is a phenomenon that is now well documented.
Now for the developmental stage that we are looking at in our article (onset of gastrulation), two studies come straight into my mind: Mesnard et al. Current Biology, 2004 and Hiramatsu et al 2013, Dev Cell.
In the first article Mesnard and colleagues have shown that the axis defined by the anterior and posterior domains aligns with the longest axis of the embryo. They also show that this is a dynamic process as the AP axis initially aligns with the shorter axis early on.
In the second article, the authors grow pre-gastrulating mouse embryos ex-vivo within agarose microwells to mimic constraints from the endometrium. They show that the soft physical constraints provided by microwells facilitate the elongation of the embryo resulting in the most distal region of the embryo ‘moving’ far from the source of BMP signalling (the extra-embryonic ectoderm). According to the authors, the resulting decrease in BMP signalling in the distal region participates in the specification of the DVE.
Overall, I believe that everyone has this intuition that geometry participate in defining the shape of signalling gradients, but there is still work to be done to understand how the cells sense the physics of their environment locally and how changes in geometry may feedback onto morphogenetic events.
Can you give us the key results of the paper in a paragraph?
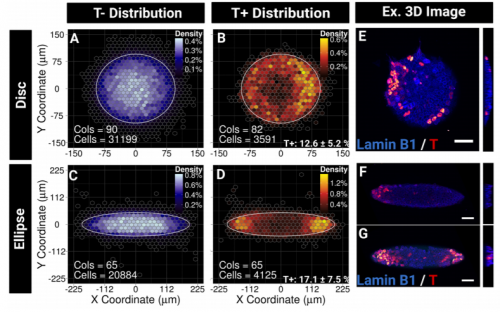
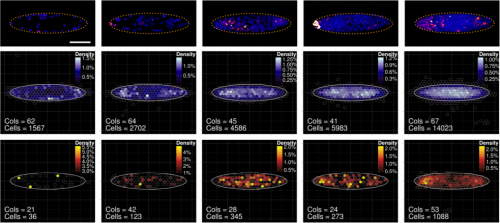
GB, MT, SL We show that imposing physical boundaries to ESC colonies in culture drives the patterning of a T+ primitive streak-like population of cells which is initially present but spatially disorganised. Importantly, if we introduce asymmetry in the shape of colonies, the resulting positioning of the cells follows the asymmetry which raises the possibility that geometry plays an important role in guiding precise positioning of the cells at the streak.
We also show that patterning is robust to changes in the relative number of T+ cells present initially. This is an important result because it indicates that the mechanisms which regulate the number of cells which will ingress into the streak may be distinct from the mechanism which regulates the precise positioning of the cells. To me, this is an indication that it is important to reflect on the notion of control during development and how processes are articulated with one another. Having a global signal triggering local events leading to patterning rather than tightly controlled signalling gradients micromanaging both patterning and specification is an elegant solution. The need for control is minimised and a certain level of robustness is achieved by the system. I can only recommend one of my favourite reviews on the topic from Arthur Lander ‘Pattern Growth and Control’
These ideas can also have great implications for our ability to engineer synthetic systems and actually one of our plans is to see if we can induce patterning in cells which do not normally sort.
Have you got any ideas for how your T+ cells end up at the periphery of the colonies?
GB/SL/MT There are quite a few hypothesis that we can formulate. A possibility is that the cells change fate if they end up in the ‘wrong place’. There may be some selective apoptosis going on as well. If the cells are surrounded by cells with a different level of T, maybe the cells are eliminated. If the cells reorganise, then again this can happen in various ways. As you suggest, cell sorting via differential adhesion may be involved or alternatively, T+ cells may become more motile and migrate faster and with more directional persistence than T- cells. This differential motility may be sufficient to drive sorting even in the absence of chemotaxic cues. It may be a combination of all of the above, we know that the embryo can use a diversity of cell behaviours in order to achieve patterning in the late blastocyst (Plusa et al, Development 2008). Our plan is to use mathematical modelling in combination with live imaging in order to further address this question.
Can you translate your findings to the early embryo – where might the guiding hand of geometry be acting?
GB, MT, SL This is a great and difficult question because it is still unclear how the shape of the streak is maintained as the embryo grows. In the chick, a community effect mediated by Nodal signalling and cell-cell interactions as well as Wnt-PCP, have been identified as important elements regulating this process (Voiculescu et al., eLife 2014). The apical polarity regulator crumbs2 has also been involved (Ramkumar et al, NCB 2016). These are examples of molecular determinants which may be part of the machinery which enable the cells to sense the folding of the streak. This geometry sensing may be key to maintain the stable geometry of the streak as the embryo grows. However much work remains to be done to fully understand this process and to assess inter-species differences as well.
When doing the research, did you have any particular result or eureka moment that has stuck with you?
GB The first one is when I saw T+ cells localising at the tips of the ellipses. Despite variability between colonies, the patterning was very clear under the microscope! I remember thinking, I need to quantify this, maybe I am just imagining it, so when we found a way to create the density maps and saw the figure showing a clear result, I was both a little a bit euphoric and very intrigued.
And what about the flipside: any moments of frustration or despair?
GB I had moments when I felt the results were not coming as fast as I wanted, this was mainly due to the fact that imaging was a bottleneck in our initial setup. As a result I was spending very long hours at the confocal in the (cold and dark) room of the imaging facility, skipping activities with family or friends. This was particularly difficult in the autumn when the colours outside were so vibrant. Not being able to see the season pass by was frustrating. But in the end we managed to streamline the whole process and this solved the issue. This was not so bad but I am lucky to have a very understanding and supportive family.
What next for you after this paper?
GB I am currently working on publishing other outputs from my post-doc which include methodological papers describing computational tools for quantitative image analysis, I hope you will be able to hear soon about it and that my tools will be useful to the community. I am also trying to establish my own group in order to continue working on synthetic approaches to identify general principles about how the cells self-organise.
Any plans for future Franco-Scottish collaboration?
MT, SL Yes definitely! We will continue to work together, the combination of microfabrication and stem cells offers a fertile ground with almost limitless possibilities. We will look into remodellable materials and 3D solutions to further guide the development of the cells in vitro. Maybe this is a good place to advertise our recent review on the topic also (Laurent et al. NBE 2018).
Finally, let’s move outside the lab – what do you like to do in your spare time in Edinburgh and Paris?
GB I like to spend time with my family, whatever we do does not matter as long as my 2 daughters keep being cheeky with their sly faces hinting at their next mischiefs. When I find time I also enjoy cycling in the beautiful moors and hills around Edinburgh.
MT I like to get high, and I know many ways for it. Apart from the classical ones, I enjoyed art exhibitions very much. Artists observe the world, analyse it interpret it, and then try to fing original means to convey their thoughts into something that will question people imagination and help them to consider reality differently. Isn’t it exactly what we do every day in our labs? Except that we have much more tools to play with … ;-) Artists also want to understand how things work, how light makes things visible, how images print in our mind, how interactions change intrinsic properties, how context change identity. So I am pretty sure most of them would enjoy Guillaume’s paper very much. Too bad they don’t dare reading Development, while we all go to the Tate.
SL There is no such thing as spare time, but during my not-doing-science time my kids have been teaching me how to mix music on a computer. I’ve only had two lessons so far but probably I will soon become an internationally famous and unfeasibly wealthy DJ. My other hobby is choosing theme tunes for papers that I like and then posting them on twitter, as my service to the developmental biology community*.
*The theme song for Guillaume’s Development paper, “Down on the Corner” by Creedence Clearwater Revival, was suggested by new Development Editor in Chief and incurable hippy James Briscoe, for which we thank him.
Geometrical confinement controls the asymmetric patterning of brachyury in cultures of pluripotent cells
Guillaume Blin, Darren Wisniewski, Catherine Picart, Manuel Thery, Michel Puceat, Sally Lowell. Development 2018 145: dev166025 doi: 10.1242/dev.166025
This is #50 in our interview series. Browse the archive here.


 (2 votes)
(2 votes)