The people behind the papers – Kana Ishimatsu, Tom Hiscock & Sean Megason
Posted by the Node Interviews, on 12 June 2018
Somites are segmented structures which give rise to numerous tissues in the vertebrate body. It has long been observed that somites scale in size with the overall size of the embryo, both as development proceeds and between individuals of different sizes, but the molecular underpinnings of this process have remained controversial. A new paper in the current issue of Development uses size-reduced zebrafish embryos to investigate this problem. We caught up with authors Kana Ishimatsu, Tom Hiscock and Sean Megason (Associate Professor of Systems Biology at Harvard Medical School) to find out more about the story.

Sean, can you give us your scientific biography and the questions your lab is trying to answer?
SM I’ve always been interested in how code makes pattern. When I was a kid, my parents bought me a computer (Commodore Vic-20). At some point I got bored with the couple of video games it came with and I taught myself how to program using the built in BASIC interpreter. I ended up making a lot of simulations of patterning based on complex dynamical systems without any training or knowing that it was a field of study. My mom was a high school librarian in the small town I grew up in so she would bring home the old Scientific Americans for me. They used to have a section called “Mathematical Recreations” that inspired a bunch of programming. In college I figured out that how the genome patterns life is the ultimate puzzle and got hooked on developmental biology.
My lab is broadly interested in how groups of cells work together to make things. We are big believers that “mechanism” typically cannot be reduced to a molecule. It emerges as interactions that can span many levels from molecules to cells to tissue mechanics, and that deciphering the mechanisms requires a balanced approach of direct observation, perturbation, and modelling across these scales.
Kana – how did you come to join Sean’s lab and be involved with the project?
KI I came to be interested in somite scaling problem while I was still in Japan, but soon realized I needed more quantitative approaches, especially in imaging. Sean’s lab had an ideal imaging system, a common interest in scaling issue, and good environment to work with people from different backgrounds.
And what about you Tom? You seem to have done a mix of wet and dry science so far in your career?
TH Yes – I started out studying physics and maths… until I joined Sean’s lab for my PhD, and fell in love with embryos and imaging. Now I have returned a little to my physics roots, and am trying to build theories to understand how animals build themselves.
Somites were first shown to scale in 1975, but decades later somite scaling and its relationship to the segmentation clock is still a controversial topic. Why do you think this is?
KI Although there are a number of models explaining somite formation, not only under a normal condition but also experimentally perturbed conditions, we have not been able to distinguish which “class” of model is working in vivo, before going into theoretically and molecularly detailed analyses. This has made more confusion than clarification.
TH Maybe one of the reasons is that there are many models that fit the data, and that it’s been difficult to design experiments to distinguish between the different hypotheses.
SM In our hands distinguishing models required rigorously comparing quantitative data with theoretical predictions, otherwise there’s a tendency to just say “oh it looks close enough”. It also required figuring out new experimental approaches that make different predictions for the different models.
Can you give us the key results of the paper in a paragraph?
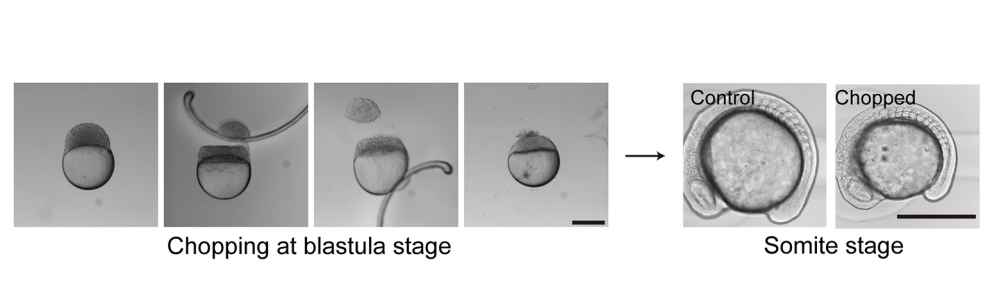
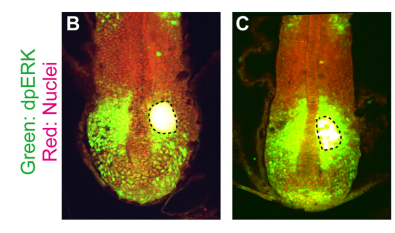
KI In order to tackle the long-standing problem of the somite formation mechanism, we started off from simply asking whether somite size scales with PSM size, which was a disputed topic in the field. We discovered that somite size scales with PSM size in a linear manner, only when the time delay between somite boundary specification and formation is taken into consideration. Based on this observation, we hypothesized the key feature of somite formation should scale with PSM size, and found only the Fgf gradient scales with PSM size, not clock period, axis elongation speed or a spatial wave pattern. Taking the time delay and the gradient scaling in to consideration, we proposed a new model, the “Clock and Scaled Gradient model”, in which scaling of a gradient is responsible for both progression and scaling of the somite formation. This model not only explains all the existing experimental results, but also makes a unique prediction, namely “an echo effect”, which was validated in vivo. Once PSM size is made artificially smaller, the system shows oscillations of somite sizes – becoming smaller and larger over and over again – which cannot be predicted by other classes or version of models.
Do you have any favourite candidates for the molecular players that would control gradient scaling?
KI Though there are several obvious candidates, like Retinoic Acid, it would be interesting if it does not primarily depend on molecules. For example, the scaling might be achieved through a gradient of pH or mechanical force. Anyway, the mechanism has to have fast dynamics because Fgf scaling occurs every somite cycle, implying that mechanisms that require a long time to achieve scaling (such as an expander-repressor mechanism) are not likely to control the gradient scaling in PSM.
SM This has been a hot topic of debate in the lab! There are some obvious candidates for players in the system (e.g. Retinoic Acid) but my best guess is that this is a case where mechanism cannot be reduced to a molecule. There are multiple signalling gradients and antagonists which all regulate each other, and in the context of lots of dynamic cell rearrangements and tissue movements.
How does somite scaling compare to other mechanisms of scaling in development?
KI Based on what we know thus far, using gradient scaling is a fairly general strategy to achieve scaling in development. When I started this project, I was almost sure that we were going to find something other than gradient scaling that is responsible for somite scaling, such as axis elongation speed or spatial patterns of waves in PSM. However, we ended up finding gradient scaling as the underlying mechanism of somite formation. It would be interesting to ask, at least theoretically, if there is any benefit to employ gradient scaling, rather than other mechanisms. Moreover, it is important to keep looking for other scaling mechanisms underlying embryo patterning.
SM The main reason I was interested in looking at somites in the size-reduced embryos was that I thought it would NOT be based on gradient scaling. In my mind, I reasoned that somite size was the product of clock period and axis elongation (wavefront regression) speed. I thought it was very unlikely that clock period would change since it is set by molecular degradation rates, but it seemed likely that the axis would extend more slowly if there were less cells. So I preferred a mechanism of “intrinsic scaling” where if there is a smaller number of precursors, you naturally get smaller products in a system governed by growth. I liked this hypothesis because it was based on geometrical/mechanical considerations rather than molecules but of course it turned out to be wrong! Gradient scaling is certainly an important way to scale pattern given the central importance of gradients in patterning itself, but I’m hopeful that there are other mechanisms to be discovered.

When doing the research, did you have any particular result or eureka moment that has stuck with you?
KI One was that we discovered that the relationship between PSM size and somite size looks completely different when we take the delay into consideration. Until then, I was a big fan of the idea that travelling waves are playing a central role in determining somite size, but this discovery changed my view 180 degrees. The other was when we came up with echo experiment that can be uniquely predicted by our model. We spent months trying to come up with one experiment that not only existing models but also any modified versions of them cannot predict. It was such an exciting moment when we finally came up with the idea and were able to see the predicted result in vivo!
TM For me, it was when we realized that scaling was a central feature of how somites are made. We’d started by thinking about why smaller embryos have smaller somites – but it got really exciting once we realized that scaling was happening throughout somitogenesis.
And what about the flipside: any moments of frustration or despair?
KI&TM There was a long time where we were really confused by the data – particularly since there is so much known about somites, it’s hard to make sense of it all. But these struggles – particularly afternoons of intense headaches and confusion together – are our fondest memories of this project!
What next for you two after this paper?
KI I would like to expand the strategy we took in this study to a non-repetitive, higher dimensional (2D and 3D) system. The advantages of studying somite formation is that it is simple enough to describe its shape as 1D and that it is a repetitive structure allowing us to easily extract its characteristic feature (length). These advantages allowed us to find its input-output relation (PSM size and somite size) and find the transfer function (gradient scaling). It is exciting if we can use the same strategy more generally to study developmental systems. Of course, I am also excited to study (1) the underlying mechanism of gradient scaling in PSM, and (2) the mechanism that integrates the positional information given by the gradient and temporal information given by the oscillator.
TM I’ve just started a postdoc position with Ben Simons and John Marioni in Cambridge, where I think about how we can use lineage tracing data and single cell sequencing to understanding development.
And where will this work take the Megason lab?
SM The mechanism of PSM gradient scaling is clearly an interesting and open question as is the regulation of the speed and duration of axis extension in general given that somite number and body length vary widely between species but are very constant within a species. The size-reduction technique also opens up the rest of the embryo to scaling questions. I am a big fan of the work of Naama Barkai and colleagues on scaling of dorsal-ventral pattern in the early embryo by Expansion-Repression, but the scaling problem must separately be solved for every subsequent part of the embryo. There’s also the question of why these scaling systems are even there. It’s not so we can chop embryos in a dish and watch them recover. Where we actually take the work just depends on who joins the lab and their interests.
Finally, let’s move outside the lab – what do you like to do in your spare time in Boston?
KI I like to eat a lot of good food, especially Chinese!
TH One of the things I miss a lot about Boston is the lovely cycle ride to Walden Pond, followed by a refreshing swim in the lake, and a big ice-cream in Concord!
SM I chase my three little monsters around and occasionally do some recreational coding and gardening.
Size-reduced embryos reveal a gradient scaling-based mechanism for zebrafish somite formation
Kana Ishimatsu, Tom W. Hiscock, Zach M. Collins, Dini Wahyu Kartika Sari, Kenny Lischer, David L. Richmond, Yasumasa Bessho, Takaaki Matsui, Sean G. Megason
Development 2018 145: dev161257 doi: 10.1242/dev.161257
This is #44 in our interview series. Browse the archive here.


 (2 votes)
(2 votes)