In Development this week (Vol. 143, Issue 6)
Posted by Seema Grewal, on 15 March 2016
Here are the highlights from the current issue of Development:
Knocking the SOX off obesity

Growth restriction in utero is associated with increased risk of obesity in later life. Recently, epigenetic inheritance was identified as an important component of this phenomenon, but the precise molecular mechanisms that underpin the association between growth restriction and obesity remain unknown. Now, on p. 950, Walter Stünkel and colleagues report a role for SOX6 in adipocyte differentiation, and suggest that SOX6 may be a key player in the association between growth restriction and obesity. By comparing adipocytes differentiated from mesenchymal stem cells from normal and growth-restricted newborn umbilical cords, the authors show that SOX6 is upregulated in growth-restricted adipocytes and that it activates key adipogenic players including PPARγ, C/EBPα and MEST. The authors also show that SOX6 interacts with β-catenin, possibly inhibiting WNT/β-catenin signalling to promote adipogenesis. Importantly, the role of SOX6 in regulating adipogenesis was also demonstrated in vivo, using Sox6 antisense oligonucleotides to target white adipose tissue in mice. Taken together, these data demonstrate a clear role for SOX6 in regulating adipocyte differentiation and adipogenesis in vivo, and provide a possible mechanistic link between growth restriction in utero and obesity in later life.
New player in neonatal heart repair

The mammalian heart has a transient regenerative ability during the neonatal stage. This ability depends on the replicative potential of endogenous cardiomyocytes; however, the underlying transcriptional network that controls cardiomyocyte replication during neonatal heart regeneration remains poorly understood. In this issue (see p. 936), Bin Zhou and colleagues investigate the role of GATA4 – a transcription factor that is crucial for cardiac specification and development – in cardiomyocyte turnover and neonatal heart repair. The authors utilised cryoinjury and apex resection models in a neonatal transgenic mouse in which they could control expression of GATA4 specifically in the cardiomyocytes. Following injury, the authors observed severely compromised ventricular function in Gata4-ablated mice, which was accompanied by reduced cardiomyocyte replication and hypertrophy. Importantly, the authors identified FGF16 as a downstream effector of the Gata4-ablated phenotype, and showed that cardiac-specific overexpression of FGF16 promoted cardiomyocyte replication and improved heart function after injury. These data identify GATA4 and FGF16 as important mediators of neonatal heart repair and bring hope for the possibility of paracrine-mediated repair in the adult heart.
A better MAP(K) for tubule elongation

Branching morphogenesis is fundamental to the development of multiple organs, including the lungs, kidneys, vasculature and mammary glands. Tubule elongation is a crucial part of branching morphogenesis and relies on the balance between cellular proliferation and migration to achieve appropriate growth. In the developing mammary gland, receptor tyrosine kinases (RTKs) regulate tubule elongation, but the relative contribution of proliferation and migration is largely unknown. Now, on p. 983, Andrew Ewald and colleagues use fluorescent reporters and real-time imaging to investigate the role of RTK signalling in tubule elongation in a 3D mouse mammary tissue culture system. The authors show that ERK signalling is required for cell migration and elongation of budding branches, but that cell proliferation is not acutely required for branch elongation. Importantly, the authors show that mosaic expression of MEK, which activates MAPK, is sufficient to induce initiation and elongation of mammary branches. This study provides a fundamental advance in our understanding of the cellular mechanism and molecular control of tube elongation in the developing mammary gland.
PLUS…
Closing the circle: from organoids back to development
 This Editorial, from our Guest Editor Melissa Little, looks at the emerging field of in vitro organogenesis and discusses how organoid technology can be applied to better understand developmental processes. Read the Spotlight on p. 905
This Editorial, from our Guest Editor Melissa Little, looks at the emerging field of in vitro organogenesis and discusses how organoid technology can be applied to better understand developmental processes. Read the Spotlight on p. 905
An interview with Melissa Little
 Melissa Little chats about her research and career, the potential of the organoid and stem cell fields, and what she hopes to achieve during her guest editorship with Development. Read the Spotlight on p. 907
Melissa Little chats about her research and career, the potential of the organoid and stem cell fields, and what she hopes to achieve during her guest editorship with Development. Read the Spotlight on p. 907
*** find out more about our upcoming Special Issue on Organoids ***
From single genes to entire genomes: the search for a function of nuclear organization
 Here, , Read the Review on p. 910
Here, , Read the Review on p. 910
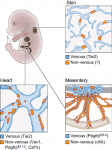
Development of the lymphatic system: new questions and paradigms
,


 Jonathan Semo
Jonathan Semo (No Ratings Yet)
(No Ratings Yet) (1 votes)
(1 votes)

 Each adult mammalian skeletal muscle has a unique complement of fast and slow myofibers, a consequence of cell-autonomous and non-cell-autonomous patterning decisions during development. Intriguingly, following either acute muscle regeneration or deinnervation/reinnervation, the proportionality and patterning of muscle fibers is largely preserved, suggesting a mechanism for ongoing maintenance of fiber type specification during homeostasis and regeneration/repair. We have recently reported that interactions between the repulsive guidance ligand ephrin-A3 [which is expressed on all and only slow (Type I) myofibers] and EphA8 [an ephrin receptor expressed by terminal Schwann cells at all and only neuromuscular junctions of fast (Type II) myofibers] promote and preserve slow myofiber identity by preventing stable innervation of slow muscle fibers by fast motor neurons [
Each adult mammalian skeletal muscle has a unique complement of fast and slow myofibers, a consequence of cell-autonomous and non-cell-autonomous patterning decisions during development. Intriguingly, following either acute muscle regeneration or deinnervation/reinnervation, the proportionality and patterning of muscle fibers is largely preserved, suggesting a mechanism for ongoing maintenance of fiber type specification during homeostasis and regeneration/repair. We have recently reported that interactions between the repulsive guidance ligand ephrin-A3 [which is expressed on all and only slow (Type I) myofibers] and EphA8 [an ephrin receptor expressed by terminal Schwann cells at all and only neuromuscular junctions of fast (Type II) myofibers] promote and preserve slow myofiber identity by preventing stable innervation of slow muscle fibers by fast motor neurons [