Xenopus represents a prime model for dissecting in vivo the signalling network that controls retinal stem cell behaviour. Its retina indeed retains a reservoir of active neural stem cells in a peripheral region, the ciliary marginal zone (CMZ), that sustains continuous eye growth throughout life. To identify regulatory mechanisms underlying retinal stem cell activity in this model system, we recently focussed our interest on the terminal effector of the Hippo pathway, the co-transcriptional factor YAP (Cabochette et al., 2015). The Hippo pathway acts as a global regulator of organ size during development (Lian et al., 2010; Ramos and Camargo, 2012). We were thus curious to question its function in the context of Xenopus indefinite retinal tissue growth.
Not surprisingly, overexpressing Yap by the mean of blastomere mRNA injections resulted in eye overgrowth, while knocking it down by Morpholino injections led to microphthalmia. In order to know whether the latter phenotype was due to embryonic or post-embryonic growth defects, we adapted in Xenopus the use of photo-cleavable Morpholinos (photo-MO). This technology, previously set up in zebrafish (Tallafuss et al., 2012), allows for inducible or reversible gene knockdowns through UV-induced cleavage of either sense or antisense photo-MOs. Our data support the conclusion that the small eye phenotype rather results from defective post-embryonic CMZ-dependent growth. This raised the hypothesis that YAP may play a specific role in the homeostatic control of post-embryonic retinal stem/precursor cell activity (Figure 1).
 Figure 1. YAP controls post-embryonic eye growth. Dissected eyes from stage 40 Xenopus tadpoles following blastomere microinjection of Yap mRNA or photo-Morpholinos (that allow knocking-down Yap only at post-embryonic stages). Compared to a control situation (on the left) Yap overexpression leads to eye overgrowth (in the middle) while post-embryonic Yap knock-down leads to a microphthalmic phenotype (on the right).
Figure 1. YAP controls post-embryonic eye growth. Dissected eyes from stage 40 Xenopus tadpoles following blastomere microinjection of Yap mRNA or photo-Morpholinos (that allow knocking-down Yap only at post-embryonic stages). Compared to a control situation (on the left) Yap overexpression leads to eye overgrowth (in the middle) while post-embryonic Yap knock-down leads to a microphthalmic phenotype (on the right).
In the post-embryonic retina, Yap expression is restricted to the tip of the CMZ where stem cells reside. We therefore anticipated that Yap knock-down may have exhausted the stem cell pool. But surprisingly stem cells were still present and analysis of their proliferation showed that they were still dividing as well. However, a severe reduction in EdU incorporation was observed, suggesting that something was wrong in their cell cycle progression. We thus employed a variety of approaches dedicated to in vivo analysis of cell cycle phase duration. We very unexpectedly found that although the cell cycle of Yap-morphant retinal stem cells lasts longer, their S-phase length is severely reduced from 17 to 4 hours. How come the S-phase of a stem cell can decrease to such extent?
During S-phase, an eukaryote cell replicates its DNA, starting from multiple replication origins scattered on the genome. This tightly regulated process follows a strict temporal program. The genome is indeed partitioned into early and late replication domains, such that some origins fire during early S-phase while others fire during late S-phase. We found that the precise choreography of the DNA replication program was altered upon Yap knock-down, with a decreased proportion of stem cells exhibiting late S-phase patterns. Late origin may thus have fired prematurely or may have not fired at all. In any cases, this likely explains the shortening in S-phase duration. Among rare factors known to produce such phenotype is c-Myc, whose expression was interestingly found to be increased in Yap morphant CMZ. Although not formally demonstrated, we thus propose that YAP may control S-phase progression through direct or indirect control of c-Myc expression.
Deregulation of DNA replication timing is known to be a source of genomic instability. In line with this, we observed an increased occurrence of DNA damage, enhanced p21 and p53 expression and increased cell death among progenitor cells derived from Yap-depleted stem cells. This ultimately leads to a failure in producing new neurons, which provides an explanation for the reduced post-embryonic growth of Yap morphant retina.
Finally, we also showed that in this temporal control of S-phase progression, YAP physically and functionally interacts with a novel partner, PKNOX1, a mammalian Homothorax ortholog belonging to the Meis/Prep homeodomain factor family, involved in the maintenance of genomic stability (Iotti et al., 2011).
Although relatively young, the research field on Hippo signalling has raised incredibly fast (Lin et al., 2013; Yu and Guan, 2013), with recent interest in the field of stem cell biology (Hiemer and Varelas, 2013; Mo et al., 2014; Piccolo et al., 2014). However, little mechanistic insights are known into how this pathway regulates stem cell properties. Our work revealed a novel role for this factor in the control of the temporal program of DNA replication (Figure 2). We propose a model where this YAP function would protect neural stem cells of the retina from experiencing genomic instability.
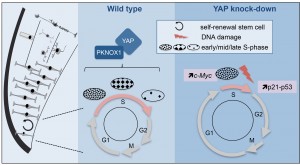 Figure 2: Model illustrating YAP function in retinal stem cells. We found that YAP is expressed in CMZ retinal stem cells (left panel). The middle panel shows the cell cycle of wild type retinal stem cells and the putative role of the YAP/PKNOX1 complex in the control of S-phase temporal progression (represented by the distinct patterns of DNA replication foci). YAP knock-down (right panel) leads to a dramatic reduction of S-phase length likely due to c-Myc-dependent premature firing of late replication origins. This results in genomic instability (increased occurrence of DNA damage, enhanced p21 and p53 expression and eventually cell death).
Figure 2: Model illustrating YAP function in retinal stem cells. We found that YAP is expressed in CMZ retinal stem cells (left panel). The middle panel shows the cell cycle of wild type retinal stem cells and the putative role of the YAP/PKNOX1 complex in the control of S-phase temporal progression (represented by the distinct patterns of DNA replication foci). YAP knock-down (right panel) leads to a dramatic reduction of S-phase length likely due to c-Myc-dependent premature firing of late replication origins. This results in genomic instability (increased occurrence of DNA damage, enhanced p21 and p53 expression and eventually cell death).
S-phase duration recently emerged as a major target of cell cycle regulation in different neural progenitor types during cortical development. Neural stem cells exhibit a substantially longer S-phase than progenitors committed to neuron production (Arai et al., 2011; Turrero García et al., 2015), a feature proposed to be dedicated to high quality control of replicated DNA, as errors would be inherited by all the progeny (Arai et al., 2011). Unique mechanisms may therefore operate in neural stem cells to control S-phase duration and ensure genomic integrity. We propose that YAP is part of the genetic network involved in this stem cell-specific regulation of the replication temporal program.
Main paper:
Cabochette, P., Vega-Lopez, G., Bitard, J., Parain, K., Chemouny, R., Masson, C., Borday, C., Hedderich, M., Henningfeld, K. A., Locker, M., et al. (2015). YAP controls retinal stem cell DNA replication timing and genomic stability. eLife 4, e08488.
References
Arai, Y., Pulvers, J. N., Haffner, C., Schilling, B., Nüsslein, I., Calegari, F. and Huttner, W. B. (2011). Neural stem and progenitor cells shorten S-phase on commitment to neuron production. Nat. Commun. 2, 154.
Hiemer, S. E. and Varelas, X. (2013). Stem cell regulation by the Hippo pathway. Biochim. Biophys. Acta 1830, 2323–34.
Iotti, G., Longobardi, E., Masella, S., Dardaei, L., De Santis, F., Micali, N. and Blasi, F. (2011). Homeodomain transcription factor and tumor suppressor Prep1 is required to maintain genomic stability. Proc. Natl. Acad. Sci. 108, E314.
Lian, I., Kim, J., Okazawa, H., Zhao, J., Zhao, B., Yu, J., Chinnaiyan, A., Israel, M. a, Goldstein, L. S. B., Abujarour, R., et al. (2010). The role of YAP transcription coactivator in regulating stem cell self-renewal and differentiation. Genes Dev. 24, 1106–18.
Lin, J. I., Poon, C. L. C. and Harvey, K. F. (2013). The hippo size control pathway–ever expanding. Sci. Signal. 6, pe4.
Mo, J.-S., Park, H. W. and Guan, K.-L. (2014). The Hippo signaling pathway in stem cell biology and cancer. EMBO Rep. 15, 642–56.
Piccolo, S., Dupont, S. and Cordenonsi, M. (2014). The Biology of YAP/TAZ: Hippo Signaling and Beyond. Physiol. Rev. 94, 1287–1312.
Ramos, A. and Camargo, F. D. (2012). The Hippo signaling pathway and stem cell biology. Trends Cell Biol. 1–8.
Tallafuss, a., Gibson, D., Morcos, P., Li, Y., Seredick, S., Eisen, J. and Washbourne, P. (2012). Turning gene function ON and OFF using sense and antisense photo-morpholinos in zebrafish. Development 139, 1691–1699.
Turrero García, M., Chang, Y., Arai, Y. and Huttner, W. B. (2015). S-phase duration is the main target of cell cycle regulation in neural progenitors of developing ferret neocortex. J. Comp. Neurol. [Epub ahead of print].
Yu, F.-X. and Guan, K.-L. (2013). The Hippo pathway: regulators and regulations. Genes Dev. 27, 355–371.
 (No Ratings Yet)
(No Ratings Yet)
 Loading...
Loading...


 (No Ratings Yet)
(No Ratings Yet)

