A day in the life of a skate lab
Posted by Kate Criswell, on 7 October 2015
Greetings! My name is Kate (but you can call me skate) Criswell and I am a Ph.D. student in the Department of Organismal Biology and Anatomy at the University of Chicago. I study axial column evolution and development in fishes, and my developmental study organism is the little skate, Leucoraja erinacea. Little skates are little known fish with a cartilaginous skeleton, and they live in the northwestern Atlantic Ocean, from North Carolina up to Nova Scotia.
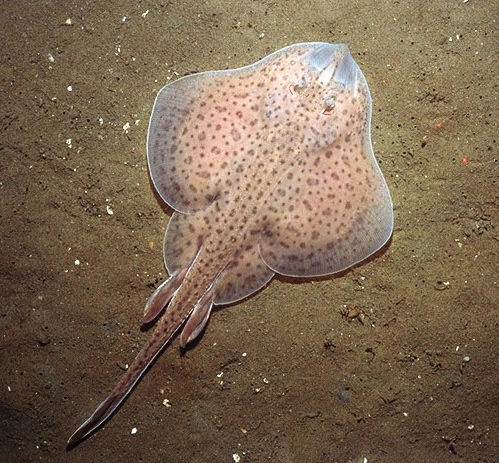

What’s so great about skates?
Skates are dorsoventrally flattened and have broad, flat pectoral fins and long, slender tails. They are closely related to sharks, rays, and chimaeras, and together these groups make up the chondrichthyans, or cartilaginous fishes. Apart from being elegant and charismatic (and for some, toothy and aggressive), chondrichthyans can provide valuable information on the broad range of developmental mechanisms at work across vertebrates. By comparing the development of cartilaginous fishes with more established organisms like zebrafish, chickens, and mice, we can tell which processes or structures are derived (specialized and more recently evolved) and which are ancestral to all vertebrates. As the sister group to all other jawed vertebrates, cartilaginous fishes exhibit many characteristics that may have been lost in their bony counterparts, or they might display features that are unique and are not found in any other vertebrates.
How do we raise skate embryos?
Skates are relatively easy to keep in a lab, and the embryos are very robust, with no need for filtered seawater or antibiotics (as long as you keep them under 18 degrees!). I do most of my live-animal manipulations in the summers at the Marine Biological Laboratory (MBL) in Woods Hole, Massachusetts. The MBL has a fantastic Marine Resources Center (MRC) with large skate brood stock tanks that they use to house adult skates (they also care for many other marine organisms, like snails, bony fish, squid, and sea urchins; a walk through the MRC is better than a trip to the aquarium!). The adult skates mate in the wild and are then collected and housed in the MRC. Females can store sperm for up to 9 months, enabling them to lay eggs throughout the year, which makes it easy to for me to acquire embryos in the winter months. Indeed, there are a number of different labs across the United States that obtain skate embryos from MBL.
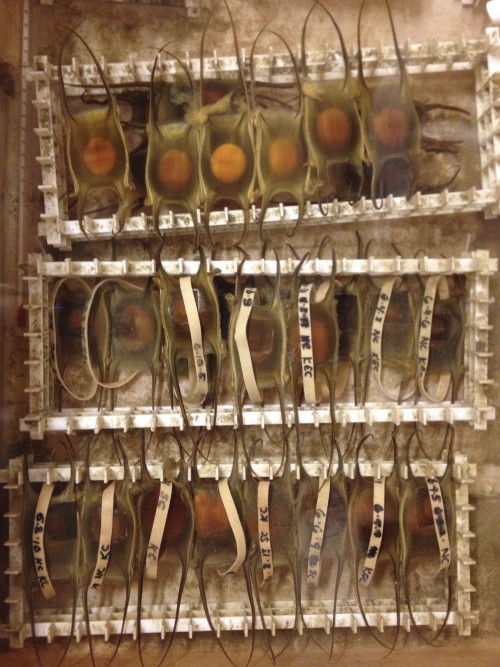
Each egg contains one embryo, which is enclosed in a capsule made of collagen fibers called a mermaid’s purse. These egg cases are rectangular in shape, with two long tendrils at each end. Once laid, staff at the MRC sort the egg cases by week and keep them in 16 degree circulating seawater that is pumped in from the nearby Great Harbor. The egg cases are closed to the environment for the first six or so weeks of development, and then small slits open at the ends of the tendrils to let water flow through. The skates take approximately 5-6 months to hatch into fully-formed, miniature adults, which gives me plenty of time (sometimes too much time) to study their development.
This video was made by my friend and collaborator, Andrew Gillis, who recorded video of the major stages of skate development this past summer at MBL. He’s got a bunch of other great skate videos over on his YouTube channel.
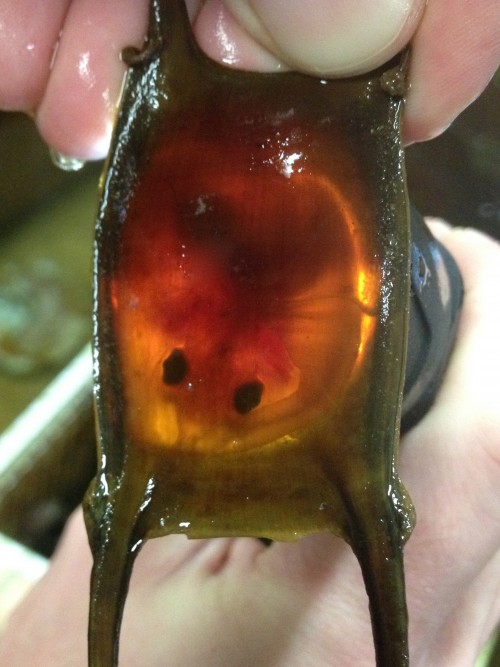
During the summers, while I am doing manipulations of live skate embryos at MBL, I make weekly trips to the MRC to go “shopping” for eggs. I collect eggs of approximately the right age, bring them to the lab space in the Loeb building, and keep them in a sea table until they are at exactly the right stage for experiments. Determining the embryonic stage of skate embryos is actually quite easy, despite their thick egg cases, because the eggs are translucent, and shining a flashlight underneath reveals the embryo curled up inside.

What kinds of experiments can we do with skates?
Their hardy nature and (relatively) short developmental period makes skates ideal chondrichthyans in which to do fate-mapping experiments. To study the embryonic origins of different parts of the axial skeleton using young embryos I cut a small window in the egg case, inject a fluorescent dye into the embryonic tissue of interest, and then glue a piece of donor eggshell over the hole (Krazy glue works best; all other glues are subpar). Skates develop a bit like chickens, with the embryo sitting atop a large yolk. When the embryos are young the yolk is extremely fragile and can’t be removed from the egg case. Slightly older embryos (stage 24 and above) have a tougher yolk, and to work with them I can simply cut around the edges of the egg case to make a large window, deposit the entire embryo + yolk in a petri dish with seawater and the anesthetic tricaine, and inject the embryo while it is in the dish. I then scoop the embryo up (using a very sophisticated tool – a plastic soup spoon from the Woods Hole Market) and place it back in the egg case for continued development. There is no need at this stage to glue the egg case shut.

After dye injection I let the embryos develop for several months in a sea table, checking the temperature every morning and removing any dead embryos. This means constantly having wet and cold hands, and having chilly seawater drip down my back when I’m not expecting it. By the end of the summer (or several weeks post-injection) I can be sure that most of the remaining embryos will survive into old age. When the axial skeleton is well developed, which takes about 3-4 months, I return to the MBL to fix the embryos and bring them back to Chicago. I spend the rest of the year analyzing experiments from the summer, using paraffin histology to cut thin sections of the axial column and then looking for dye-labeled cells in different parts of the vertebrae.
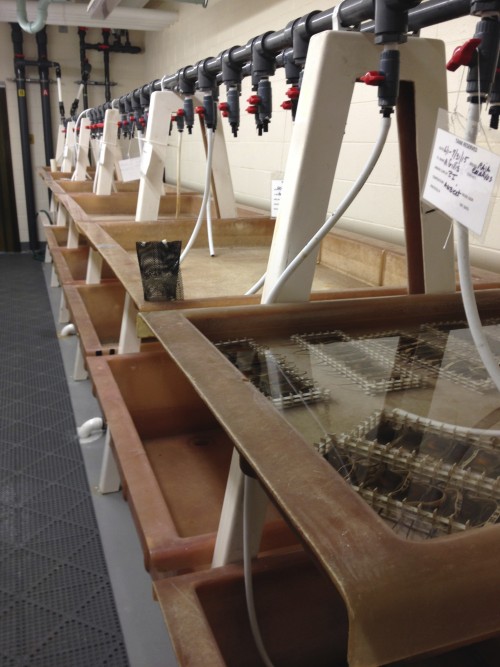

In addition to fate-mapping, skates respond well to other experimental manipulations, like drug treatments and bead implantations. For drug treatments, the drug of choice can either be injected into the egg case before it opens to let seawater flow through or, for later stages, the embryos can be removed from their egg cases and placed in baths for several days. Once returned to circulating seawater the embryos can continue developing perfectly well sitting in glass bowls in the sea table, outside of their egg cases.
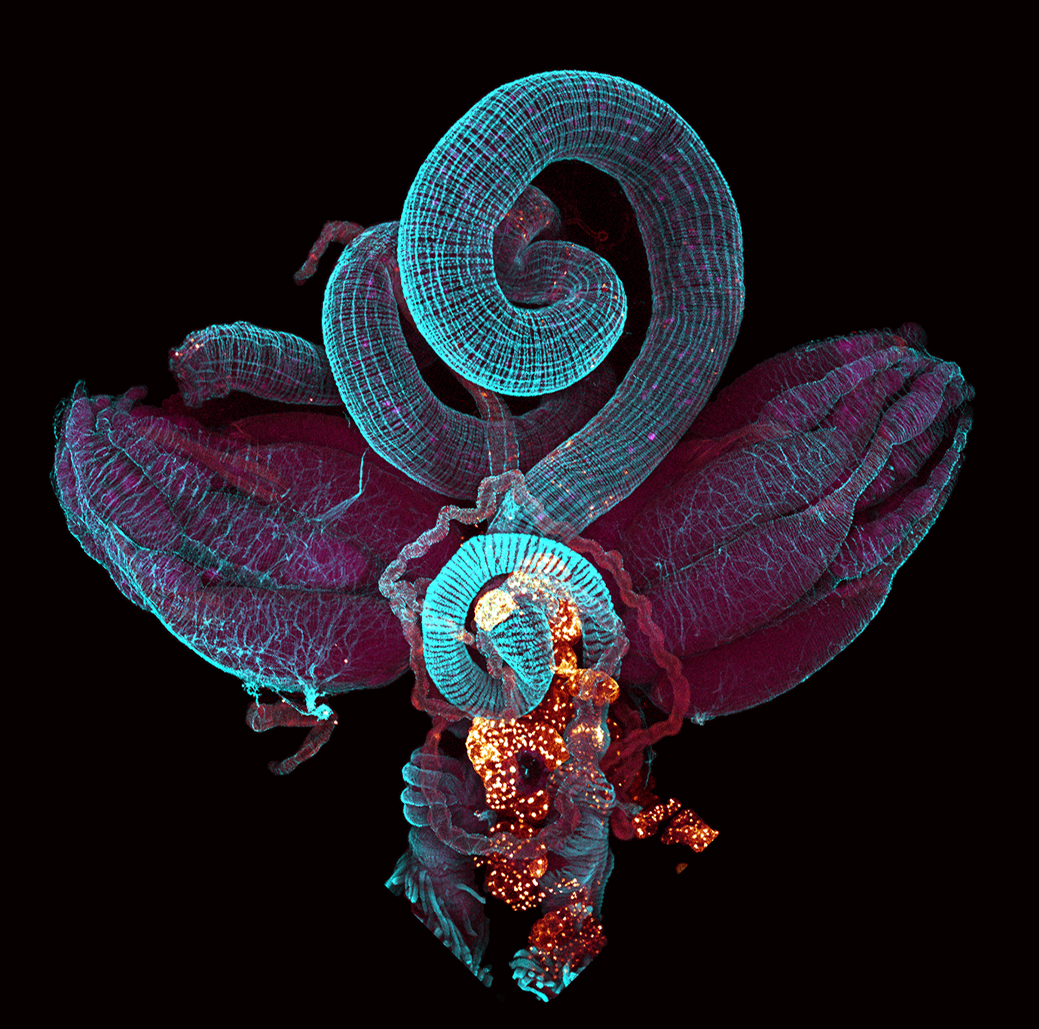
Even though studying skate development can be difficult (especially when your experiments take four months!) it is extremely rewarding. I am excited to continue working in this system and to expand upon the tools currently available to learn as much as I can about the evolution of vertebrate development.
 This post is part of a series on a day in the life of developmental biology labs working on different model organisms. You can read the introduction to the series here and read other posts in this series here.
This post is part of a series on a day in the life of developmental biology labs working on different model organisms. You can read the introduction to the series here and read other posts in this series here.


 (16 votes)
(16 votes)
 (7 votes)
(7 votes)


 Apoptosis was originally regarded as a ‘silent’ mechanism of cell elimination designed to degrade the contents of doomed cells. However, during the past decade it has become clear that apoptotic cells can produce diverse signals that have a profound impact on neighboring cells and tissues. Here,
Apoptosis was originally regarded as a ‘silent’ mechanism of cell elimination designed to degrade the contents of doomed cells. However, during the past decade it has become clear that apoptotic cells can produce diverse signals that have a profound impact on neighboring cells and tissues. Here, 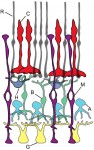 Photoreceptors – the light-sensitive cells in the vertebrate retina – have been extremely well-characterized with regards to their biochemistry, cell biology and physiology. They therefore provide an excellent model for exploring the factors and mechanisms that drive neural progenitors into a differentiated cell fate in the nervous system. Here,
Photoreceptors – the light-sensitive cells in the vertebrate retina – have been extremely well-characterized with regards to their biochemistry, cell biology and physiology. They therefore provide an excellent model for exploring the factors and mechanisms that drive neural progenitors into a differentiated cell fate in the nervous system. Here, 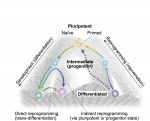 During development, cells transition from a pluripotent to a differentiated state, generating all the different types of cells in the body. Although development is generally considered an irreversible process, it is now possible to reprogram mature cells to pluripotency. Here,
During development, cells transition from a pluripotent to a differentiated state, generating all the different types of cells in the body. Although development is generally considered an irreversible process, it is now possible to reprogram mature cells to pluripotency. Here, (No Ratings Yet)
(No Ratings Yet)