Research Associate – Bioinformatician
Posted by stemcellsjobs, on 13 March 2015
Closing Date: 15 March 2021
Department/Location: Wellcome Trust – Medical Research Council Cambridge Stem Cell Institute, University of Cambridge
Salary: £28,695-£37,394
Reference: PS04734
Closing date: 20 April 2015
Fixed-term: The funds for this post are available until 30 September 2017 in the first instance.
Applications are invited for a computational biologist to join the research group of Dr Paul Bertone at the Wellcome Trust – Medical Research Council Stem Cell Institute at the University of Cambridge. We are applying state-of-the-art experimental and computational methods toward the understanding of transcriptional regulation in pluripotent stem cells from rodents and primates.
We work closely with the groups of Dr Jennifer Nichols and Professor Austin Smith at the SCI, and seek to fill this position with a postdoctoral scientist who will join a new collaborative project funded by the BBSRC. This arrangement provides an excellent environment for research and career development, as the post holder will benefit from the resources and expertise of both experimental and computational environments to lead this multidisciplinary initiative.
The group applies large-scale gene expression profiling, comparative genomics and regulatory systems modelling to mammalian embryogenesis and stem cell biology. The post holder will have the opportunity to lead related computational projects, but will also work in partnership with experimentalists and contribute to the design and execution of collaborative studies. He or she will have a strong publication record, excellent communication skills, and enjoy working on ambitious projects at the frontiers of genomics and biotechnology.
Candidates will have a solid background in computation and current knowledge of eukaryotic genomics. Expertise in statistical methods and Unix scripting are essential. Additional experience with high-throughput sequencing data, numerical computing or regulatory systems modelling is desirable but not required. Applicants should hold a PhD in a relevant field (e.g. Bioinformatics, Applied Mathematics, Computer Science, Biomedical Engineering, or Biochemistry/Molecular Biology with a computational component). Previous experience in stem cell biology is not necessary.
Once an offer of employment has been accepted, the successful candidate will be required to undergo a health assessment.
To apply online for this vacancy and to view further information about the role, please visit: http://www.jobs.cam.ac.uk/job/5481. This will take you to the role on the University’s Job Opportunities pages. There you will need to click on the ‘Apply online’ button and register an account with the University’s Web Recruitment System (if you have not already) and log in before completing the online application form.
The closing date for all applications is the Monday 20th April 2015.
Informal enquiries about the post are also welcome via email: cscrjobs@cscr.cam.ac.uk.
Please upload your Curriculum Vitae (CV) and a covering letter in the Upload section of the online application to supplement your application. If you upload any additional documents which have not been requested, we will not be able to consider these as part of your application.
Interviews will be held towards the beginning of May 2015.
Please quote reference PS04734 on your application and in any correspondence about this vacancy.
The University values diversity and is committed to equality of opportunity.
The University has a responsibility to ensure that all employees are eligible to live and work in the UK.


 (No Ratings Yet)
(No Ratings Yet) (2 votes)
(2 votes)
 Dmrt1 and its related genes play a key role in sex determination in a broad range of metazoan species. However, Dmrt1 has become dispensable for testis determination in mammals, and this function is instead carried out by Sry, which is a newly evolved gene found on the Y chromosome. Now, Peter Koopman and colleagues show that, even though its function is not normally required, Dmrt1 is able to drive female-to-male sex reversal in mice (p.
Dmrt1 and its related genes play a key role in sex determination in a broad range of metazoan species. However, Dmrt1 has become dispensable for testis determination in mammals, and this function is instead carried out by Sry, which is a newly evolved gene found on the Y chromosome. Now, Peter Koopman and colleagues show that, even though its function is not normally required, Dmrt1 is able to drive female-to-male sex reversal in mice (p.  In plants, stem cell proliferation is negatively regulated by the receptor kinase CLAVATA1 (CLV1) and its peptide ligand CLAVATA3 (CLV3). Previous studies have suggested that CLV1 acts redundantly with other receptor kinases, such as BAM1, 2 and 3, but the molecular mechanisms underpinning this redundancy have been unclear. Now, Elliot Meyerowitz and co-workers interrogate the role of CLV1-CLV3 signalling in the Arabidopsis shoot apical meristem (p.
In plants, stem cell proliferation is negatively regulated by the receptor kinase CLAVATA1 (CLV1) and its peptide ligand CLAVATA3 (CLV3). Previous studies have suggested that CLV1 acts redundantly with other receptor kinases, such as BAM1, 2 and 3, but the molecular mechanisms underpinning this redundancy have been unclear. Now, Elliot Meyerowitz and co-workers interrogate the role of CLV1-CLV3 signalling in the Arabidopsis shoot apical meristem (p.  Hematopoietic stem cells (HSCs) give rise to all cells of the adult blood system, and understanding how these cells first arise during embryogenesis is important for developing regenerative medicine-based strategies for producing HSCs in vitro. Here, David Traver and colleagues demonstrate that Gata2b acts as an early regulator of zebrafish hematopoietic precursors (p.
Hematopoietic stem cells (HSCs) give rise to all cells of the adult blood system, and understanding how these cells first arise during embryogenesis is important for developing regenerative medicine-based strategies for producing HSCs in vitro. Here, David Traver and colleagues demonstrate that Gata2b acts as an early regulator of zebrafish hematopoietic precursors (p. Adherens junctions (AJs), which are specialised E-cadherin-based cell contacts, are continuously remodelled during tissue morphogenesis, as cells change shape and position. The accumulation of Bazooka (Baz), the Drosophila PAR3 homologue, is thought to specify where new E-cadherin complexes are deposited during AJ remodelling, but what regulates Baz localisation? Here, Alexandre Djiane and colleagues show that the scaffold protein Magi regulates Baz localization and hence AJ remodelling inDrosophila eye epithelial cells (p.
Adherens junctions (AJs), which are specialised E-cadherin-based cell contacts, are continuously remodelled during tissue morphogenesis, as cells change shape and position. The accumulation of Bazooka (Baz), the Drosophila PAR3 homologue, is thought to specify where new E-cadherin complexes are deposited during AJ remodelling, but what regulates Baz localisation? Here, Alexandre Djiane and colleagues show that the scaffold protein Magi regulates Baz localization and hence AJ remodelling inDrosophila eye epithelial cells (p.  Skeletal stem cells (SSCs) reside in the postnatal bone marrow and give rise to cartilage, bone, hematopoiesis-supportive stroma and marrow adipocytes. Here, Paolo Bianco and Pamela Robey discuss the biology of SSCs in the context of the development and postnatal physiology of skeletal lineages, to which their use in medicine must remain anchored. See the Development at a Glance poster article on p.
Skeletal stem cells (SSCs) reside in the postnatal bone marrow and give rise to cartilage, bone, hematopoiesis-supportive stroma and marrow adipocytes. Here, Paolo Bianco and Pamela Robey discuss the biology of SSCs in the context of the development and postnatal physiology of skeletal lineages, to which their use in medicine must remain anchored. See the Development at a Glance poster article on p. 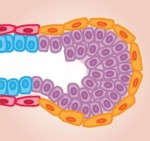 The mammary gland provides an excellent model for studying ‘stem/progenitor’ cells, which – in this context – allow for the repeated expansion and renewal of the gland during adult life. Here, Mina Bissell and colleagues discuss the various cell types that constitute the mammary gland, highlighting how they arise and differentiate, and how the microenvironment influences their development. See the Review on p.
The mammary gland provides an excellent model for studying ‘stem/progenitor’ cells, which – in this context – allow for the repeated expansion and renewal of the gland during adult life. Here, Mina Bissell and colleagues discuss the various cell types that constitute the mammary gland, highlighting how they arise and differentiate, and how the microenvironment influences their development. See the Review on p.  Today, a few button clicks gives access to vast troves of knowledge, and a few dollars buys technologies that even well-funded labs could not get a few decades ago. So, it should be much easier for amateurs and hobbyists to do scientific research now than it was in Darwin’s era. The
Today, a few button clicks gives access to vast troves of knowledge, and a few dollars buys technologies that even well-funded labs could not get a few decades ago. So, it should be much easier for amateurs and hobbyists to do scientific research now than it was in Darwin’s era. The  Facilitating amateur-professional interactions would also improve public understanding of science. This is especially important in areas that intersect with developmental biology; voters are routinely called upon to make decisions related to stem cells, genetics, or evolution. The premise of every graduate school is that the best way to learn how science works is to do it, yet there are few opportunities for adult non-scientists to experience the creative and intellectual side of research. The success of the citizen science movement shows that many people are interested in participating in science. However, most citizen science projects are designed to get a large number of volunteers to do a defined task, rather than to help non-scientists plan research and interpret results. This leaves a big gulf between non-scientists and professionals.
Facilitating amateur-professional interactions would also improve public understanding of science. This is especially important in areas that intersect with developmental biology; voters are routinely called upon to make decisions related to stem cells, genetics, or evolution. The premise of every graduate school is that the best way to learn how science works is to do it, yet there are few opportunities for adult non-scientists to experience the creative and intellectual side of research. The success of the citizen science movement shows that many people are interested in participating in science. However, most citizen science projects are designed to get a large number of volunteers to do a defined task, rather than to help non-scientists plan research and interpret results. This leaves a big gulf between non-scientists and professionals. Developmental biology is ripe for this. Although a lot of developmental biology depends on expensive reagents and high-tech equipment, plenty of high-value, low-tech research remains to be done. Two of my all-time favorite papers (1, 2) used nothing more than glass needles and intelligence to identify, and partially solve, a paradox of ctenophore development: when an embryo is split in two, each half develops into half an embryo; yet the adults can regenerate an entire half of their body. The authors documented ontogenetic transitions in these phenomena, and then deciphered the roles of specific cell lineages in patterning and regeneration. In my own work, I’ve found that the most useful biomechanical techniques for working with embryos are things like micropipette aspiration, which would be easily accessible to amateur microscopists (it was developed in the 1950’s (3)). There are myriad questions in developmental biology that could be investigated with low-budget techniques.
Developmental biology is ripe for this. Although a lot of developmental biology depends on expensive reagents and high-tech equipment, plenty of high-value, low-tech research remains to be done. Two of my all-time favorite papers (1, 2) used nothing more than glass needles and intelligence to identify, and partially solve, a paradox of ctenophore development: when an embryo is split in two, each half develops into half an embryo; yet the adults can regenerate an entire half of their body. The authors documented ontogenetic transitions in these phenomena, and then deciphered the roles of specific cell lineages in patterning and regeneration. In my own work, I’ve found that the most useful biomechanical techniques for working with embryos are things like micropipette aspiration, which would be easily accessible to amateur microscopists (it was developed in the 1950’s (3)). There are myriad questions in developmental biology that could be investigated with low-budget techniques.