In Development this week (Vol. 142, Issue 1)
Posted by Seema Grewal, on 16 December 2014
Here are the highlights from the current issue of Development:
Planar cell polarity squeezes in on the action
 The planar cell polarity (PCP) pathway regulates the polarization of epithelial tissues in various contexts, but recent studies suggest that the PCP pathway also influences other aspects of morphogenesis. Here, Sergei Sokol and colleagues uncover a role for PCP signalling during apical constriction in Xenopus embryos (p. 99). They show that the core PCP protein Vangl2 accumulates at the apical surface of blastopore bottle cells, which undergo apical constriction during gastrulation. The depletion of Vangl2 perturbs apical constriction and hence blastopore formation. The authors further demonstrate that Rab11, a marker of the recycling endosome, localizes to the apical surface of constricting cells in a Vangl2-dependent manner; apical staining of Rab11 is absent in Vangl2-depleted embryos, suggesting that PCP signalling modulates endocytic trafficking. Finally, the authors show that Rab11 in turn modulates Vangl2 distribution and that it cooperates with Myosin V to regulate apical constriction. Together, these studies highlight a novel role for the PCP pathway during apical constriction and support a positive-feedback model in which both PCP signalling and endocytic trafficking function to regulate apical constriction.
The planar cell polarity (PCP) pathway regulates the polarization of epithelial tissues in various contexts, but recent studies suggest that the PCP pathway also influences other aspects of morphogenesis. Here, Sergei Sokol and colleagues uncover a role for PCP signalling during apical constriction in Xenopus embryos (p. 99). They show that the core PCP protein Vangl2 accumulates at the apical surface of blastopore bottle cells, which undergo apical constriction during gastrulation. The depletion of Vangl2 perturbs apical constriction and hence blastopore formation. The authors further demonstrate that Rab11, a marker of the recycling endosome, localizes to the apical surface of constricting cells in a Vangl2-dependent manner; apical staining of Rab11 is absent in Vangl2-depleted embryos, suggesting that PCP signalling modulates endocytic trafficking. Finally, the authors show that Rab11 in turn modulates Vangl2 distribution and that it cooperates with Myosin V to regulate apical constriction. Together, these studies highlight a novel role for the PCP pathway during apical constriction and support a positive-feedback model in which both PCP signalling and endocytic trafficking function to regulate apical constriction.
Epiblast development: getting up to speed
 The epiblast of mammalian embryos undergoes a period of rapid growth shortly after implantation, thereby establishing a population of cells that will give rise to the embryo proper. Here, Miguel Ramalho-Santos and co-workers show that chromodomain helicase DNA-binding protein 1 (Chd1) is required for the transcriptional output that drives this rapid growth (p. 118). They first show that Chd1–/– mouse embryos display post-implantation defects; analyses of lineage and patterning markers indicate that Chd1–/–embryos arrest in the transition between E5.5 and E6.5, prior to anterior-posterior patterning and the onset of gastrulation. The researchers further show that transcriptional output per cell is reduced inChd1–/– mouse embryonic stem cells (ESCs) compared with control ESCs. In line with this, the amount of RNA polymerase II present at gene bodies and transcriptional start sites is decreased in mutant ESCs. Finally, the authors document that Chd1 also directly regulates the output of ribosomal RNA in both ESCs and the epiblast. In summary, the authors propose that Chd1 promotes a global increase in transcriptional output by both RNA polymerase I and II that, in turn, sustains the rapid growth of the epiblast.
The epiblast of mammalian embryos undergoes a period of rapid growth shortly after implantation, thereby establishing a population of cells that will give rise to the embryo proper. Here, Miguel Ramalho-Santos and co-workers show that chromodomain helicase DNA-binding protein 1 (Chd1) is required for the transcriptional output that drives this rapid growth (p. 118). They first show that Chd1–/– mouse embryos display post-implantation defects; analyses of lineage and patterning markers indicate that Chd1–/–embryos arrest in the transition between E5.5 and E6.5, prior to anterior-posterior patterning and the onset of gastrulation. The researchers further show that transcriptional output per cell is reduced inChd1–/– mouse embryonic stem cells (ESCs) compared with control ESCs. In line with this, the amount of RNA polymerase II present at gene bodies and transcriptional start sites is decreased in mutant ESCs. Finally, the authors document that Chd1 also directly regulates the output of ribosomal RNA in both ESCs and the epiblast. In summary, the authors propose that Chd1 promotes a global increase in transcriptional output by both RNA polymerase I and II that, in turn, sustains the rapid growth of the epiblast.
Ezh2: balancing cell differentiation in the lung
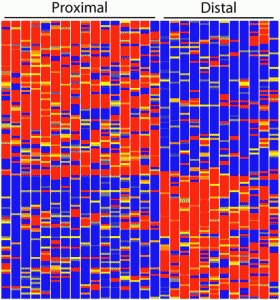
 During development, the lung endoderm is patterned along its anterior-posterior axis, giving rise to distinct epithelial lineages, such as the alveolar cells that mediate gas exchange, and the basal and secretory cells that line the airways. In this issue (p. 108), Edward Morrisey and colleagues show that the polycomb repressive complex 2 component Ezh2 restricts the basal cell lineage during lung development, thereby allowing correct patterning of the lung. The researchers report that Ezh2 is broadly expressed in the lung during early development but then gradually becomes downregulated as development progresses. Importantly, they demonstrate that the endoderm-specific deletion of Ezh2 impairs secretory cell differentiation while inducing the ectopic and premature development of basal cells that express the transcription factor Trp63 and other basal cell markers. Furthermore, they report that Ezh2 deletion gives rise to a cell population that might represent an intermediate state between basal and secretory states. These and other findings indicate that Ezh2 controls the phenotypic switch between basal cells and secretory cells, and regulates both the temporal and spatial patterning of the lung.
During development, the lung endoderm is patterned along its anterior-posterior axis, giving rise to distinct epithelial lineages, such as the alveolar cells that mediate gas exchange, and the basal and secretory cells that line the airways. In this issue (p. 108), Edward Morrisey and colleagues show that the polycomb repressive complex 2 component Ezh2 restricts the basal cell lineage during lung development, thereby allowing correct patterning of the lung. The researchers report that Ezh2 is broadly expressed in the lung during early development but then gradually becomes downregulated as development progresses. Importantly, they demonstrate that the endoderm-specific deletion of Ezh2 impairs secretory cell differentiation while inducing the ectopic and premature development of basal cells that express the transcription factor Trp63 and other basal cell markers. Furthermore, they report that Ezh2 deletion gives rise to a cell population that might represent an intermediate state between basal and secretory states. These and other findings indicate that Ezh2 controls the phenotypic switch between basal cells and secretory cells, and regulates both the temporal and spatial patterning of the lung.
MRTFs at the heart of epicardial motility
 The epicardium – the single-cell layer of mesothelium that surrounds the heart – harbours a population of progenitor cells that modulates heart development and contributes to various cardiac lineages. During heart development, these epicardium-derived progenitor cells (EPDCs) undergo epithelial-to-mesenchymal transition and migrate into the sub-epicardial space, but the mechanisms regulating their mobilization remain unclear. On p. 21, Eric Small and colleagues show that myocardin-related transcription factors (MRTFs) regulate the motility of mouse EPDCs as well as the maturation of coronary vessels. They demonstrate that MRTF-A and MRTF-B are enriched within the epicardium, where they localize to the perinuclear space. The researchers further demonstrate that, in epicardial-mesothelial cells cultured in vitro, TGFβ signalling leads to the nuclear accumulation of MRTFs and the activation of a cell motility gene expression program. Importantly, the epicardial-specific ablation of Mrtfa and Mrtfb causes sub-epicardial haemorrhage; mutant hearts display a disorganised epicardial layer. In addition, lineage-tracing studies reveal a novel epicardial-derived coronary pericyte population that contributes to coronary vessel integrity and that is depleted in mutant embryos. Together, these findings, which link EPDC motility to cell differentiation in the heart, highlight novel approaches that could be used to manipulate EPDCs for cardiac repair.
The epicardium – the single-cell layer of mesothelium that surrounds the heart – harbours a population of progenitor cells that modulates heart development and contributes to various cardiac lineages. During heart development, these epicardium-derived progenitor cells (EPDCs) undergo epithelial-to-mesenchymal transition and migrate into the sub-epicardial space, but the mechanisms regulating their mobilization remain unclear. On p. 21, Eric Small and colleagues show that myocardin-related transcription factors (MRTFs) regulate the motility of mouse EPDCs as well as the maturation of coronary vessels. They demonstrate that MRTF-A and MRTF-B are enriched within the epicardium, where they localize to the perinuclear space. The researchers further demonstrate that, in epicardial-mesothelial cells cultured in vitro, TGFβ signalling leads to the nuclear accumulation of MRTFs and the activation of a cell motility gene expression program. Importantly, the epicardial-specific ablation of Mrtfa and Mrtfb causes sub-epicardial haemorrhage; mutant hearts display a disorganised epicardial layer. In addition, lineage-tracing studies reveal a novel epicardial-derived coronary pericyte population that contributes to coronary vessel integrity and that is depleted in mutant embryos. Together, these findings, which link EPDC motility to cell differentiation in the heart, highlight novel approaches that could be used to manipulate EPDCs for cardiac repair.
Mga fuels pluripotent cells
 The dual specificity T-box/bHLH-zipper transcription factor Mga is expressed in pluripotent cells of the mouse embryo and in embryonic stem cells (ESCs), but its function in these cells is unclear. Here, Virginia Papaioannou and colleagues examine the role of Mga in early development and show that it is essential for the survival of pluripotent cells (p. 31). They first show that Mga depletion in early mouse embryos and ESCs causes growth defects; increased cell death is observed in the inner cell mass (ICM) of mutant embryos in vivoand in vitro, and in Mga mutant ESCs cells in vitro. Lineage specification, in contrast, is unaffected by Mgadepletion. The researchers further identify the enzyme ornithine decarboxylase (ODC), which converts ornithine to putrescine in the polyamine synthesis pathway, as a candidate downstream target of Mga. Accordingly, they demonstrate that exogenous putrescine can rescue the ICM in Mga mutant embryos and the survival of Mga mutant ESCs. These findings highlight a role for polyamines in pluripotent cells and suggest that Mga controls cell survival in early embryos and ESCs by regulating polyamine pools.
The dual specificity T-box/bHLH-zipper transcription factor Mga is expressed in pluripotent cells of the mouse embryo and in embryonic stem cells (ESCs), but its function in these cells is unclear. Here, Virginia Papaioannou and colleagues examine the role of Mga in early development and show that it is essential for the survival of pluripotent cells (p. 31). They first show that Mga depletion in early mouse embryos and ESCs causes growth defects; increased cell death is observed in the inner cell mass (ICM) of mutant embryos in vivoand in vitro, and in Mga mutant ESCs cells in vitro. Lineage specification, in contrast, is unaffected by Mgadepletion. The researchers further identify the enzyme ornithine decarboxylase (ODC), which converts ornithine to putrescine in the polyamine synthesis pathway, as a candidate downstream target of Mga. Accordingly, they demonstrate that exogenous putrescine can rescue the ICM in Mga mutant embryos and the survival of Mga mutant ESCs. These findings highlight a role for polyamines in pluripotent cells and suggest that Mga controls cell survival in early embryos and ESCs by regulating polyamine pools.
Plus…
In recognition of recent breakthroughs and in line with Development’s recent expansion into the stem cell field, we recently organized a workshop – ‘From Stem Cells to Human Development’ – that was held in September 2014. In this issue, you will find a report from this meeting as well as an Editorial and several Spotlight articles that address key issues in this field.
Looking inwards: opening a window onto human development
Development Editors announce a new focus on human developmental biology and discuss how they hope to support this expanding field. See the Editorial on p. 1
Ethical considerations in chimera research
The use of human tissue, particularly in the generation of chimeric animals, throws up important ethical considerations that scientists and policy-makers must consider. See the Spotlight article by Göran Hermerén on p. 3
From naïve pluripotency to chimeras: a new ethical challenge?
The ability to create chimeric animal models using naïve human pluripotent stem cells is now on the horizon. Should we be concerned about using such chimeric animals? Insoo Hyun discusses these concerns on p. 6
Mouse and human blastocyst-derived stem cells: vive les differences
Early human and mouse embryos exhibit significant differences in their development and, as discussed by Janet Rossant, these are reflected in the properties of stem cell lines derived from these embryos. See the Spotlight on p. 9
Modeling human lung development and disease using pluripotent stem cells
Successful disease therapy often requires an in-depth knowledge of basic developmental biology, not only through study of model organisms but crucially also of human tissue. Hans-Willem Snoeck discusses the importance of using human stem cell models for understanding human lung development and disease. See the Spotlight on p. 13
On human development: lessons from stem cell systems
In September 2014, over 100 scientists from around the globe gathered at Wotton House near London for the Company of Biologists’ workshop ‘From Stem Cells to Human Development’. The workshop covered diverse aspects of human development, from the earliest stages of embryogenesis to differentiation of mature cell types of all three germ layers from pluripotent cells. , Here, Alexander Medvinsky and Frederick Livesey summarise some of the exciting data presented at the workshop and draw together the main themes that emerged. See the Meeting Review on p. 17


 (No Ratings Yet)
(No Ratings Yet) (3 votes)
(3 votes)
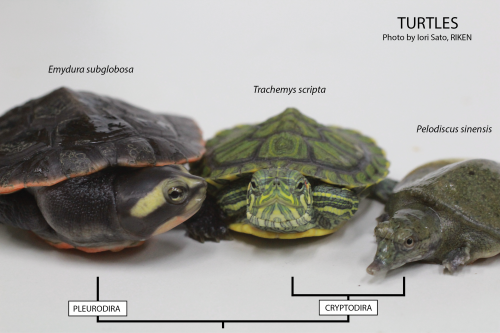


 This post is part of a series on a day in the life of developmental biology labs working on different model organisms. You can read the introduction to the series
This post is part of a series on a day in the life of developmental biology labs working on different model organisms. You can read the introduction to the series  (15 votes)
(15 votes)