Mapping the Embryo for Developmental Biologists & Stem Cell Researchers; LifeMap Discovery® – a Roadmap of Mammalian Cell Ontology
Posted by arielrinon, on 30 March 2014
Understanding how cells differentiate during embryonic development is invaluable for the in vitro derivation of functional cells from stem cells. However, mapping the human embryo, including characterization of all the cell types that make up the developing and mature human body, and of all embryonic progenitor cell types that appear in intermediate developmental stages, is an overwhelming challenge.
The LifeMap Discovery database has taken a lead role in this effort, providing the research community with an easy to use data portal describing embryonic development, along with substantial information about stem and progenitor cells, relevant differentiation protocols and cell therapy applications.
The database has been designed to integrate data derived from in vivo and in vitro experimental setups, including gene expression and signaling information in developing cells. This knowledge is essential for identification and classification of laboratory-derived stem cells and can be used to match such with in vivo cells sharing an identical or similar gene expression profile.
The database is divided into the following modules:
LifeMap Discovery provides easy access to a wealth of information on mammalian development, integrated with stem cell biology and regenerative medicine information.
Highlights:
– A detailed description of the developmental ontology of organs/tissues, anatomical compartments and cells
– Manually curated gene expression relating to all developmental stages, as well as data extracted from high throughput experiments and large scale in situ databases
– Detailed information on signals that regulate cells during development
LifeMap Discovery describes cultured stem, progenitor and primary cells, along with related differentiation protocols. Integration of stem cell biology and embryonic development knowledge provided in the database can be harnessed by stem cell researchers to better characterize experimentally obtained cell derivatives and to develop or improve stem cell differentiation protocols.
Highlights:
– Provides valuable information regarding stem cell types, such as gene expression profile and cellular markers
– Contains a large collection of stem cell differentiation protocols
– Enables accurate characterization of cultured stem and progenitor cells during differentiation processes
– Promotes an in-depth understanding of how stem cells differentiate, and of the key signals governing the process
LifeMap Discovery summarizes cell-based therapies that aim to apply stem, progenitor or primary cells towards treatment of degenerative diseases.
Highlights:
– A concentrated source of information on cell therapies spanning the different stages of development
– The provided information has been manually curated from multiple literature sources, such as: scientific publications, press releases, patent applications and clinical trials registries.
– The comprehensive information presented for each cell therapy includes: an overview of the therapy, a list of therapeutic cells utilized in the specific treatment, mode and regimen of cell delivery, mechanism of action, formulation, in vitro data, animal models, preclinical data and related clinical trials.
LifeMap Discovery also functions as a gene expression database, aiming to sketch a systematic map of gene expression profiles within cells and tissues.
Highlights:
– Gene expression profiling of developing and adult mammalian organs, tissues, anatomical compartments and cells, as well as for cultured stem, progenitor and primary cells, or cells derived using differentiation protocols; this genetic information enables characterization of annotated cells by their gene expression patterns.
– GeneAnalyticsTM, a powerful gene expression analysis tool, has recently been added to the LifeMap Discovery tool box. The platform integrates and clusters data extracted from multiple and variable resources and supports simultaneous analysis of multiple genes, applying a novel algorithm to match gene sets to tissues, anatomical compartments and cells within the database. The GeneAnalytics application is the most comprehensive analysis tool currently available for modeling gene expression data in the embryo and the adult body.
LifeMap Discovery is a free tool for academics. We welcome you to watch our short introductory movie to learn more about the database and hope you will find LifeMap Discovery a useful tool for your ongoing and future research!
Please do not hesitate to contact me, by commenting on this post, for more information about LifeMap Discovery.
Dr. Ariel Rinon
LifeMap Discovery Team
References:
- LifeMap Discovery is available at: http://discovery.lifemapsc.com/
- Edgar R, Mazor Y, Rinon A, Blumenthal J, Golan Y, Buzhor E, Livnat I, Ben-Ari S, Lieder I, Shitrit A, Gilboa Y, Ben-Yehudah A, Edri O, Shraga N, Bogoch Y, Leshansky L, Aharoni S, West MD, Warshawsky D, Shtrichman R (2013). LifeMap Discovery™: the embryonic development, stem cells, and regenerative medicine research portal. PLoS One. 2013; 8(7): e66629.


 (4 votes)
(4 votes)
 (No Ratings Yet)
(No Ratings Yet)



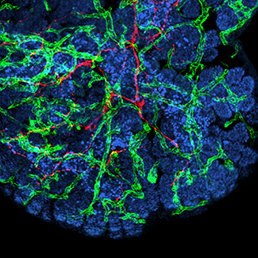 In many organs, the processes of vascularisation and innervation are frequently interdependent. In the pancreas, islet endocrine cells secrete vascular endocrine growth factor (VEGF) to direct vascularisation, but little is known about how islet innervation is regulated. On p.
In many organs, the processes of vascularisation and innervation are frequently interdependent. In the pancreas, islet endocrine cells secrete vascular endocrine growth factor (VEGF) to direct vascularisation, but little is known about how islet innervation is regulated. On p. 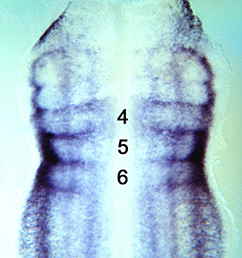
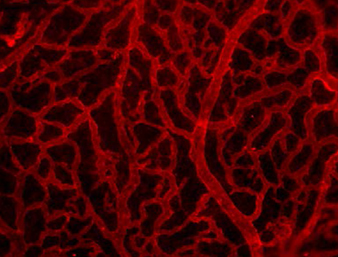 Clathrin-mediated endocytosis regulates the signalling activity and turnover of multiple plasma membrane proteins. Interfering with endocytosis can therefore have complex effects on developmental processes. William Sessa and colleagues (p.
Clathrin-mediated endocytosis regulates the signalling activity and turnover of multiple plasma membrane proteins. Interfering with endocytosis can therefore have complex effects on developmental processes. William Sessa and colleagues (p. 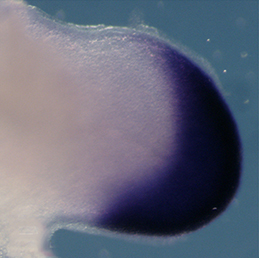 The vertebrate limb is an important model for understanding developmental patterning processes. Along the proximo-distal axis, the limb is segmented into stylopod (upper limb), zeugopod (lower limb) and autopod (hand/foot) regions, marked by the expression of particular homeobox genes – Meis1/2 most proximally and Hoxa13 most distally. Fibroblast growth factor (FGF) is a key inducer of distal fate, while the role of retinoic acid (RA) in promoting proximal fate has been controversial. Miguel Torres and co-workers (p.
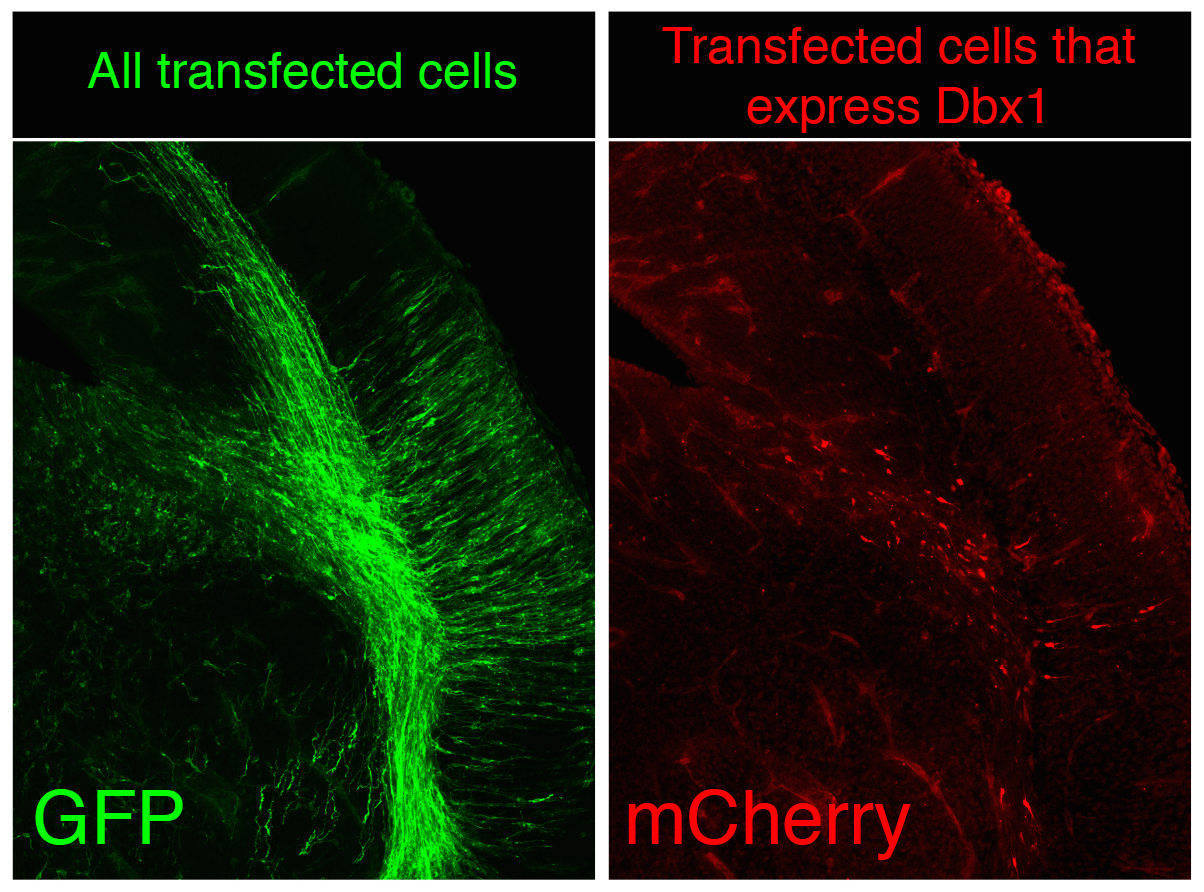
The vertebrate limb is an important model for understanding developmental patterning processes. Along the proximo-distal axis, the limb is segmented into stylopod (upper limb), zeugopod (lower limb) and autopod (hand/foot) regions, marked by the expression of particular homeobox genes – Meis1/2 most proximally and Hoxa13 most distally. Fibroblast growth factor (FGF) is a key inducer of distal fate, while the role of retinoic acid (RA) in promoting proximal fate has been controversial. Miguel Torres and co-workers (p. 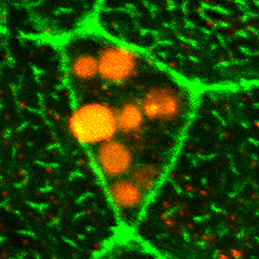 The Xenopus embryonic epidermis is a mucociliary epithelium – analogous to that found in mammalian airways. This tissue is characterised by the presence of multiciliated cells (MCCs) and goblet cells. A third cell type, the ion-secreting cell, was recently discovered in the Xenopus epidermis. Two papers now identify a final cell type in the frog embryonic skin, the small secretory cell (SSC). Eamon Dubaissi and colleagues (p.
The Xenopus embryonic epidermis is a mucociliary epithelium – analogous to that found in mammalian airways. This tissue is characterised by the presence of multiciliated cells (MCCs) and goblet cells. A third cell type, the ion-secreting cell, was recently discovered in the Xenopus epidermis. Two papers now identify a final cell type in the frog embryonic skin, the small secretory cell (SSC). Eamon Dubaissi and colleagues (p. 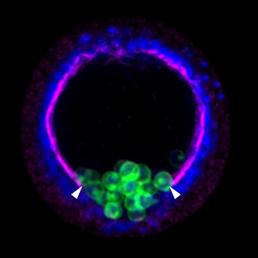 Epithelial-mesenchymal transition (EMT) is a cell state change used repeatedly during development across evolution, while aberrant EMT is associated with cancer metastasis. The transcriptional control of EMT has been extensively studied, and several transcription factors (TFs) shown to play crucial roles; notably, TFs such as Twist and Snail have been proposed to be ‘master regulators’ of EMT. On p.
Epithelial-mesenchymal transition (EMT) is a cell state change used repeatedly during development across evolution, while aberrant EMT is associated with cancer metastasis. The transcriptional control of EMT has been extensively studied, and several transcription factors (TFs) shown to play crucial roles; notably, TFs such as Twist and Snail have been proposed to be ‘master regulators’ of EMT. On p.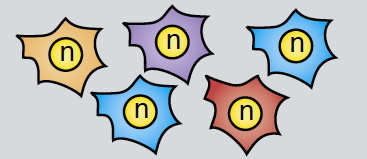  Haploid genetics holds great promise for understanding genome evolution and function. Much of the work on haploid genetics has previously been limited to microbes, but possibilities now extend to mammals. Here, Anton Wutz examines the potential use of haploid cells and puts them into a historical and biological context. See the Development at a Glance poster article on p.
Haploid genetics holds great promise for understanding genome evolution and function. Much of the work on haploid genetics has previously been limited to microbes, but possibilities now extend to mammals. Here, Anton Wutz examines the potential use of haploid cells and puts them into a historical and biological context. See the Development at a Glance poster article on p.  Cilia play many essential roles in fluid transport and cellular locomotion, and as sensory hubs for a variety of signal transduction pathways. Here, Sudipto Roy and colleagues review our understanding of the transcriptional control of ciliary biogenesis, highlighting the activities of FOXJ1 and the RFX family of transcriptional regulators. See the Review article on p.
Cilia play many essential roles in fluid transport and cellular locomotion, and as sensory hubs for a variety of signal transduction pathways. Here, Sudipto Roy and colleagues review our understanding of the transcriptional control of ciliary biogenesis, highlighting the activities of FOXJ1 and the RFX family of transcriptional regulators. See the Review article on p.