September in preprints
Posted by the Node, on 31 October 2025
Welcome to our monthly trawl for developmental and stem cell biology (and related) preprints.
The preprints this month are hosted on bioRxiv – use these links below to get to the section you want:
- Patterning & signalling
- Morphogenesis & mechanics
- Genes & genomes
- Stem cells, regeneration & disease modelling
- Plant development
- Environment, evolution and development
Research practice and education
Spotted a preprint in this list that you love? If you’re keen to gain some science writing experience and be part of a friendly, diverse and international community, consider joining preLights and writing a preprint highlight article.
Developmental biology
| Patterning & signalling
Tissue-wide, synchronous Erk oscillations time the segmentation of the zebrafish notochord
Priyom Adhyapok, James Norman, Michel Bagnat, Stefano Di Talia
Cellular NAD+ availability and redox state constrain developmental speed in the Drosophila eye
Nisha Veits, Yuting Guo, Jingjing He, Khalil Mazouni, Ivan Nemazanyy, Martin Bres, Cara Picciotto, Claire Mestdagh, Yan Yan, Francois Schweisguth
Cytoneme feedback ensures signaling specificity when multiple ligands converge on a common receptor
Akshay Patel, Molly Maranto, Hind Hitti, Sougata Roy
Kathryn G. Powers, Joshua Hahn, Juliette Wohlschlegel, Olivia Bermingham-McDonogh
Yuan-Chen Tsai, Hajime Ozaki, Xinyi Wang, Axel A. Almet, Isabel Fleming, Kaori Shiraiwa, Matthew Jung Min Noh, Caihao Nie, Sunnyana Trejo, Bret Kiyoshi Sugita, Jiya Dalal, Ruben Alberto Gonzalez, Briana De Jesus, Gregory Li-Min Chen, Michael J Gandal, Qing Nie, Momoko Watanabe
Amber M. Ridgway, Javier Figueras Jimenez, Beñat Yáñez Iturbe-Ormaeche, Maria D. S. Nunes, Alistair P. McGregor
Ectomesenchymal identity emerges via relief of Twist1 transcript destabilization
Lara C. Busby, Jessica R. Patrick, Luke W. Lyons, Megan L. Martik
Spatiotemporal proteomics reveals dynamic antagonistic gradients shaping signalling waves
Wilke H. M. Meijer, Virginia Andrade, Suzan Stelloo, Wouter M. Thomas, Marek J. van Oostrom, Eveline F. Ilcken, Kim T. J. Peters, Michiel Vermeulen, Katharina F. Sonnen
Bruce suppresses autophagy-regulated caspase activity and wing tissue growth in Drosophila
Natsuki Shinoda, Yutaro Hama, Nozomi Hanawa, Masayuki Miura
Nathan Anderson, Sen-Lin Lai, Chris Q. Doe
Asfa Kamal, Yash Sheregare, Chaitanya Patkar, Mrunmayi Markam, Rujul Deolikar, Nawaneetan Sriraman, Aditya Seth, Manish Jaiswal, Rohan Jayant Khadilkar
Notch coordinates self-organization of germ layers and axial polarity in cnidarian gastruloids
Sanjay Narayanaswamy, Franziska Haas, Emmanuel Haillot, Elly Tanaka, Ulrich Technau
Daniëlle T.J. Woutersen, Andreas van Impel, Stefan Schulte-Merker, Jeroen den Hertog
Andrew T Plygawko, Jamie Adams, Zack Richards, Kyra Campbell
Nodal/Smad2 signaling sustains developmental pausing by repressing Pparg-mediated lipid metabolism
Giacomo Furlan, S. Bryn Martin, Brandon Cho, Sarah A. McClymont, Elizabeth J. Robertson, Evelyne Collignon, Miguel Ramalho-Santos
Spatio-temporal control of myoblast identity drives muscle diversity in the Drosophila leg
Camille Guillermin, Violaine Tribollet, Mathilde Bouchet, Anne Laurencon, Dan Zhou, Sergio Sarnataro, Laurent Gilquin, Isabelle Stevant, Guillaume Marcy, Emeric Texeraud, Yad Ghavi-Helm, Benjamin Gillet, Sandrine Hughes, Samantha Vonau, Jonathan Enriquez
Rac1 inhibition slows down the segmentation clock and increases the somite size
Maria Pappa, Charisios D. Tsiairis
Sergio Juarez-Carreño, Marco Milán
Mahla Ahmadi, Heike Rudolf, Christine Mau, Jimena Garcia-Guillen, Ezzat El-Sherif
Endothelial Slit2 guides the Robo1-positive sympathetic innervation during heart development
Juanjuan Zhao, Susann Bruche, Konstantinos Lekkos, Carolyn Carr, Joaquim M. Vieira, John G. Parnavelas, William D. Andrews, Mathilda T.M. Mommersteeg
Uncoupling Neocortical Neuron Fate and Migration via a Let-7–RBX2 Axis
Steven Decker, Keiko Hino, Anna La Torre, Sergi Simó
Canonical WNT signalling governs Echinococcus metacestode development
Ruth Herrmann, Luisa Schiegl, Michaela Herz, Kilian Rudolf, Akito Koike, Markus Spiliotis, Monika Bergmann, Nancy Holroyd, Uriel Koziol, Matt Berriman, Klaus Brehm
Vascular Patterning affects Intramembranous Ossification through HIF1α-Vegf Signaling
Soma Dash, Jonathan R. Rettig, Madelaine Gogol, Paul A. Trainor
Alicia Donoghue, Lewis S Mosby, Inês Lago Baldaia, Tamara Hodgetts, Evelina Ursu, Zeynep Erten, Zena Hadjivasiliou, Vilaiwan M Fernandes
Gli3R-mediated inhibition of hedgehog signaling alters the embryonic transcriptome in zebrafish
Anna J. Moyer, Summer B. Thyme
Zhuangzhi Zhang, Zhejun Xu, Tongye Fu,Jialin Li, Feihong Yang, Chuannan Yang, Wenhui Zheng, Zizhuo Sha, Yanjing Gao, Mengge Sun, Zhenmeiyu Li, Jing Ding, Xiaosu Li, Zhengang Yang
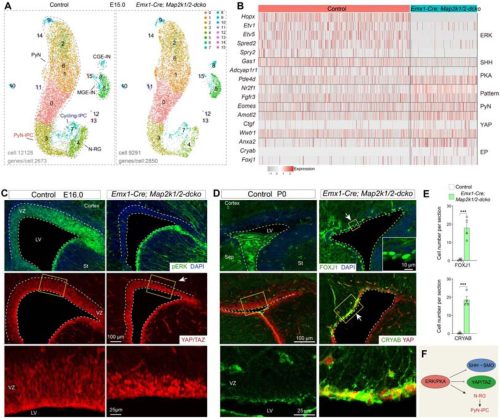
The Wnt co-receptor Arrow-LRP5/6 is required for Planar Cell Polarity establishment in Drosophila
Ursula Weber, Reza Farhadifar, Marek Mlodzik
Rachel A. Minerath, Rajesh K. Kasam, Casey O. Swoboda, Vikram Prasad, Kelly M. Grimes, N. Scott Blair, Hadi Khalil, Christina M Alfieri, Logan Eads, Anthony J. Saviola, Mohamad Azhar, Lianjie Miao, Mingfu Wu, Michelle Tallquist, Kirk C. Hansen, Matthew T Weirauch, Katherine E. Yutzey, Douglas P. Millay, Jeffery D. Molkentin
Chaperone AIP Couples mTORC1 Activation and Catabolic Metabolism During Neonatal Development
Márta Korbonits, Xian Wang, Sayka Barry, Chung Thong Lim, Oniz Suleyman, Stefano De Tito, Nazia Uddin, Maria Lillina Vignola, Charlotte Hall, Laura Perna, J. Paul Chapple, Gabor Czibik, Sian M Henson, Valle Morales, Katiuscia Bianchi, Viðar Örn Eðvarðsson, Kristján Ari Ragnarsson, Viktoría Eir Kristinsdóttir, Anne Debeer, Yoeri Sleyp, Rena Zinchenko, Glenn Anderson, Michael Duchen, Kritarth Singh, Chih Yao Chung, Yu Yuan, Sandip Patel, Artem O. Borovikov, Hans Tómas Björnsson, Hilde Van Esch, Sharon Tooze, Ezra Aksoy, Caroline Brennan, Oliver Haworth
Ayami Nakagawa, Hidekazu Iwakawa, Hiro Takahashi, Simon Vial-Pradel, Mari Takahashi, Kazuomi Ohga, Takuma Ito, Motoki Tamai, Eri Takada, Sumie Keta, Nanako Ishibashi, Sayuri Ando, Iwai Ohbayashi, Masaki Ito, Mami Yamazaki, Byung-Yoon Cha, Je-Tae Woo, Michiko Sasabe, Munetaka Sugiyama, Shoko Kojima, Yasunori Machida, Chiyoko Machida
Zukai Liu, Chengxiang Qiu, Connor A. Kubo, Stella Xu, Riza M. Daza, Eva Nichols, Wei Yang, Anh Vo, Mary B. O’Neill, Choli Lee, Jay Shendure, Nobuhiko Hamazaki
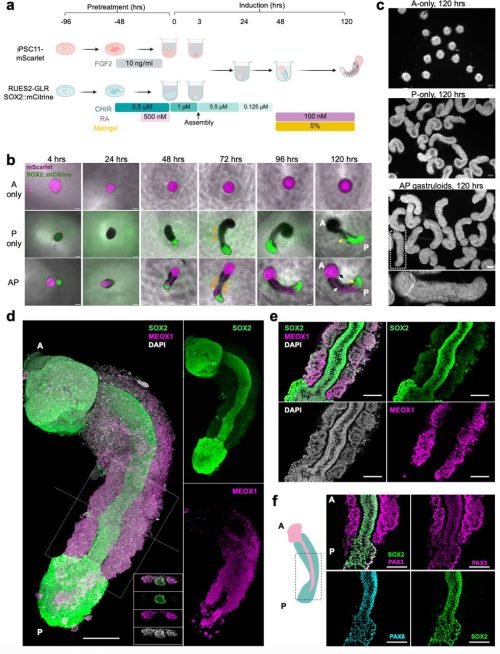
Martina Bohuslavová (née Stiborová), Andrea Hauserová, Rebecca Collier, Joaquin Lilao-Garzón, Silvia Muñoz-Descalo, Alexander W. Bruce
Cell autonomous polarization by the planar cell polarity signaling pathway
Alexis T Weiner, Silas Boye Nissen, Kaye Suyama, Bomsoo Cho, Gandhy Pierre-Louis, Jeffrey D Axelrod
E(spl)m4 Directly Antagonizes Traf4 to Inhibit JNK Signaling in Drosophila Katrin Strobel, Jennifer Falconi, Cédric Leyrat, Rémi Logeay, Sarah J. Bray, Alexandre Djiane
| Morphogenesis & mechanics
Lineage domains and cytoskeletal cables organize a cellular square grid in a crustacean
Beatrice L. Steinert, Leo Blondel, Chandrashekar Kuyyamudi, Evangelia Stamataki, Anastasios Pavlopoulos, Cassandra G. Extavour
Rio Tsutsumi, Antoine N. Diez, Steffen Plunder, Ryuichi Kimura, Shinya Oki, Kaori Takizawa, Haruhiko Akiyama, Yusuke Mii, Ritsuko Takada, Shinji Takada, Mototsugu Eiraku
Julie Warin, Estelle Balissat, Pauline Colombier, Lily Paillat, Maeva Dutilleul, Anne Camus
Biophysical mechanisms of morphogenesis in lizard lungs
Kaleb Hill, Aaron H. Griffing, Michael A. Palmer, Bezia Lemma, Aria Lupo, Tony Gamble, Natalia A. Shylo, Andrej Košmrlj, Paul A. Trainor, Celeste M. Nelson
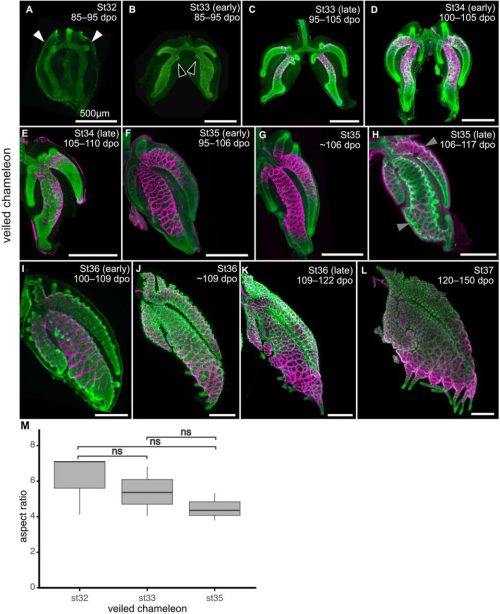
VANGL2 shapes the mouse heart tube from adjacent epithelia and without planar polarity
Paul Palmquist-Gomes, Gaëlle Letort, Ayushi U. Hegde, José María Pérez-Pomares, Sigolène M. Meilhac
Olga D. Jarosińska, Amalia Riga, Hala Zahreddine Fahs, Joren M. Woeltjes, Ruben Schmidt, Fathima S. Refai, Suma Gopinadhan, Kristin C. Gunsalus, Mike Boxem
Bhavik Rathod, Jasmine Samvelyan, Nicole Gustafsson, Aneta Liszka, Narelle McGregor, Jianyao Wu, Claes Ohlsson, Anna Fahlgren, Natalie Sims, Jonas Fuxe, Göran Andersson, Jessica J Alm, Sara H Windahl
Kelli Johnson, Xiangning Dong, Zhiwei Xiao, Hamza Islam, Meghan Anderman, Ian Glass, Jason R. Spence, Katherine D. Walton
Favour Ononiwu, Melissa Mikolaj, Christopher Dell, Abdalla Wael Shamil, Kedar Narayan, Heidi Hehnly
Critical phenomenon underlies de novo luminogenesis during mammalian follicle development
Kim Whye Leong, Yuting Lou, Arikta Biswas, Jue Yu Kelly Tan, Boon Heng Ng, Xixun Lu, Xin Ping Joan Teo, Thong Beng Lu, Carlo Bevilacqua, Isabelle Bonne, Robert Prevedel, Tetsuya Hiraiwa, Chii Jou Chan
Distinct dermal fibroblasts direct mechano-chemical signaling to the epidermis during pregnancy
Yoshihiko Kobayashi, Kazunori Sunadome, Koichiro Maki, Hiroki Fukunaga, Hitomi Matsubara, Sahomi Ohkubo, Ritsuko Maki, Maki Yoshikawa, Aleksandra Tata, Purushothama Rao Tata, Ken-ichi Matsumoto, Taiji Adachi, Mitsuhiro Iwaki, Takuya Yamamoto, Fumiko Toyoshima
Patterned integrin–laminin adhesion coordinates epithelial collective cell migration
Anna Mertens, Nicola Moratscheck, Petra A Klemmt, Chaitanya Dingare, Marion Basoglu, Virginie Lecaudey
Single-cell morphodynamics predict cell fate decisions during mucociliary epithelial differentiation
Mari Tolonen, Ziwei Xu, Ozgur Beker, Varun Kapoor, Bianca Dumitrascu, Jakub Sedzinski
Sarah N. Steiner, Eric Horst, Mitre Athaiya, Craig N. Johnson, Joseph Y. Shen, Michelle L. Kerns, Geeta Mehta, Ramiro Iglesias-Bartolome, Pierre A. Coulombe
Noor Al-Hajri, Mar Alujas, Sofia J. Araújo, Guy Tear
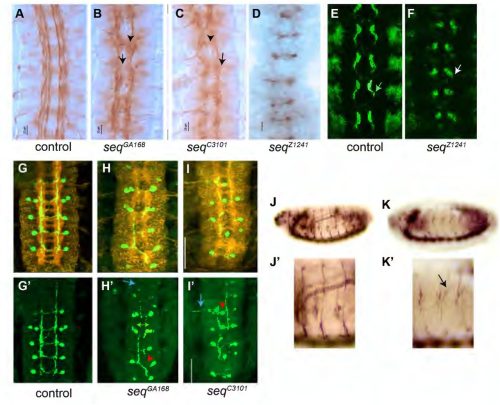
Sox2 Regulates Lateral Line Morphogenesis via Yap/Taz-Mediated Mechanotransduction
Akshai Janardhana Kurup, Aya Mikdache, Patricia Diabangouaya, Gwendoline Gros, Camila Garcia-Baudino, Cristian A. Undurraga, Andres F. Sarrazin, Pedro P. Hernandez
Evgeny G. Ivashkin, Olga I. Taimanova, Anton I. Bogomolov, Elena N. Temereva
A structural transition ensures robust formation of skeletal muscle
Mario A. Mendieta-Serrano, Yiqi Hou, Sophie Theis, Thomas E. Hall, Shannon E. Taylor, Berta Verd, Robert G. Parton, Timothy E. Saunders
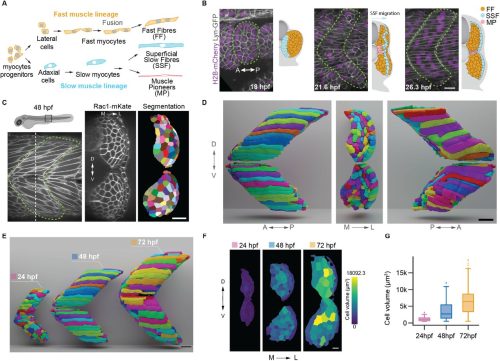
Sunandan Dhar, Serena Thomas, Hui Li Yeo, Timothy E. Saunders, Sudipto Roy
Elizabeth A. Bearce, Samuel G. Bertrand, Samara Williams, Sophie I. Fisher, Adamend L. Freda, Zoe H. Irons, Calvin Chmelir, Daniel T. Grimes
A self-limiting mechanotransduction feedback loop ensures robust organ formation
Yusuke Mori, Paul Robin, Anne Belle Briggs, Daniel S. Levic, Jiacheng Wang, Kira L. Heikes, Michel Bagnat, Edouard Hannezo, Akankshi Munjal
Dynamics of bicoid mRNA localisation and translation dictate morphogen gradient formation
T. Athilingam, E.L. Wilby, P. Bensidoun, A. Trullo, M. Verbrugghe, X. Shi, M. Lagha, T.E. Saunders, T.T. Weil
Roles of K-channel activity in feather bud morphogenesis
Madison Zitting, Zhou Yu, Ting-Xin Jiang, Ping Wu, Randall Widelitz, Cheng-Ming Chuong, Robert Hsiu-Ping Chow
| Genes & genomes
Alka Rani, Melissa Arboleda, Heidi Stuhlmann
Bhawana Maurya, Allan C Spradling
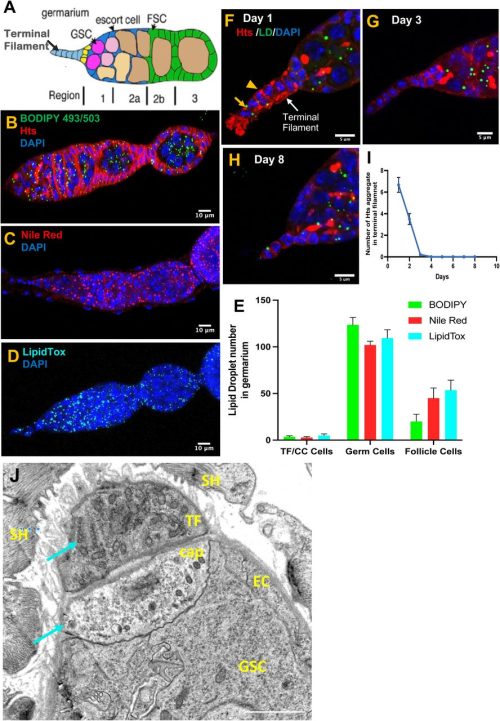
Pauliina Paloviita, Sonja Nykänen, Sanna Vuoristo
Felipe L. Teixeira, Brian Sanderson, Jennifer L. Hackett, Erik A. Lundquist
Liesl G. Strand, Charlotte P. Choi, Savannah McCoy, Emmanuel T. Nsamba, Nicola Silva, Anne M. Villeneuve
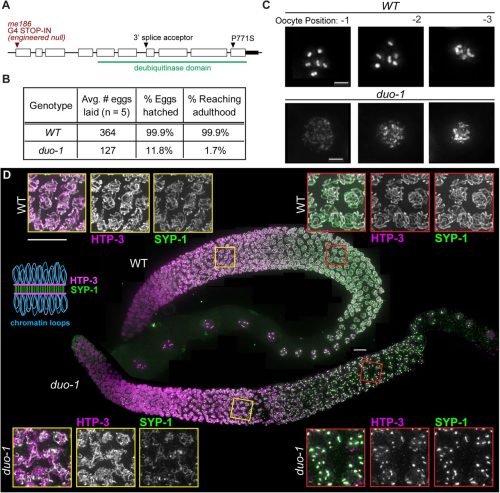
Avijit Mallick, Theresa Ramalho, Yunguang Du, Rui Li, Levi Ali, Sookyung Kim, Pooja Rai, Andreas Bergmann, Lihua Julie Zhu, Cole M. Haynes
Neural crest induction requires SALL4-mediated BAF recruitment to lineage specific enhancers
Martina Demurtas, Samantha M. Barnada, Emma van Domselaar, Zoe H. Mitchell, Laura Deelen, Marco Trizzino
Editing-independent effects of Adar in Drosophila melanogaster
Khadija Hajji, Damiano Amoruso, Barbora Nováková, Nagraj Sambrani, Deying Yang, Anzer Khan, Vojtech Bystry, Alejandro Medaglia-Mata, Domenico Alessandro Silvestris, Ernesto Picardi, Mary A. O’Connell, Liam P. Keegan
Rafael Casado-Navarro, Ana Bermejo-Santos, Rodrigo Torrillas-de la Cal, María Pilar Madrigal, Virgilia Olivé, Li Ying Chen-Chen, Sonia Amorós-Bru, Sandra Jurado, Esther Serrano-Saiz
Single-cell-scale spatial transcriptome of the developing and adult mouse ovary
Anthony S. Martinez, Tyler J. Gibson, Jennifer McKey
Decoding the Spatial and Epigenetic Logic of Early Skin Development
Nigel L. Hammond, Syed Murtuza Baker, Amanda McGovern, Antony Adamson, Andrew D Sharrocks, Magnus Rattray, Jill Dixon, Michael J. Dixon
Nicotinamide opposes ET-1’s adverse effect on uterine decidualization via EDNRB
Yuye Wang, Qing Ma, Meitong Chen, Yukako Kayashima, Jiayi Zhou, Balaji Rao, Jessica L Bowser, Xianwen Yi, Nobuyo Maeda-Smithies, Feng Li
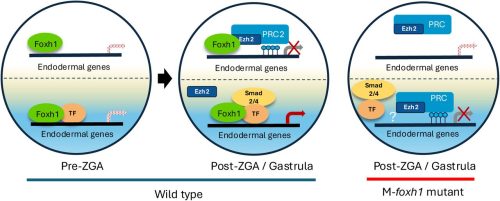
Foxh1 is a locus-specific PRC2 recruiter governing germ layer silencing
Jin Cho, Clark L. Hendrickson, Nathan Mar, Ira L. Blitz, Margaret Fish, Wenqi Wang, Ken W.Y. Cho
Dissecting Gene Regulatory Networks Governing Human Cortical Cell Fate
Jingwen W. Ding, Chang N. Kim, Megan S. Ostrowski, Yashodara Abeykoon, Bryan J. Pavlovic, Jenelle L. Wallace, Tomasz J. Nowakowski, Alex A. Pollen
eIF4E assembly into C. elegans germ granules is essential for its repressive function
Carmen Herrera Sandoval, Christopher Borchers, Boyoon Yang, Becky Boyd, Heather A. Hundley, Scott T. Aoki
Kei Fukuda, Kimiko Inoue, Chikako Shimura, Moe Kitazawa, Michiko Hirose, Shogo Matoba, Atsuo Ogura, Yoichi Shinkai
Ovarian development is driven by early spatiotemporal priming of the coelomic epithelium
Cyril Djari, Chloé Mayère, Maëva Guy, Aitana Perea-Gomez, Paul Barreau, Agathe Rozier, Anthony S. Martinez, Tyler J. Gibson, Cassandre Guérin, Herta Ademi, Dagmar Wilhelm, Jennifer McKey, Marie-Christine Chaboissier, Serge Nef
Sen-Lin Lai, Chris Q. Doe
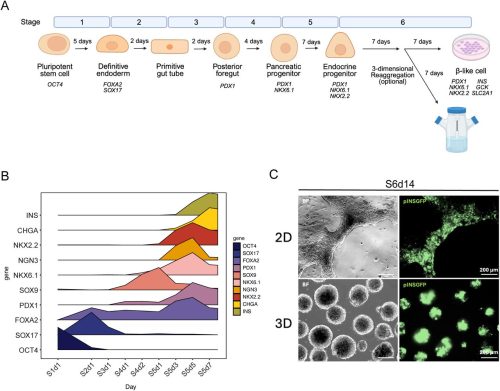
Human NKX2.2 influences islet endocrine cell fate choices through regulation of WNT pathway genes
Christopher Schaaf, Fiona M. Docherty, Madison X. Rodriguez, Patrick Sean McGrath, Christopher J. Hill, Kristen L. Wells, Lori Sussel
Rodrigo O. de Castro, Agustin Carbajal, Katarzyna P. Nowak, Ayelén González Montoro, Monika Kawecka, Luciana Previato, Roberto J. Pezza
SPT5 regulates Pol II pausing and elongation in different ways at early versus late embryonic stages
Alessandro Dulja, Marvin Mayer, Niklas Engel, Arkadiy K. Golov, Mattia Forneris, Yacine Kherdjemil, Songjie Feng, Rebecca R. Viales, Eileen E.M. Furlong
EHMT2 Controls Neural Crest-Derived Craniofacial Development but is Dispensable in Limb Development
Ye Liu, Yaguang Zhao, Minmin Liu, Paul Kim, Ji Liao, Di Lu, Huadie Liu, Piroska E. Szabó, Tao Yang
Andrea S. K. Jones, D. Ford Hannum, Taylor Schissel, Jordan H. Machlin, Vasantha Padmanabhan, Jun Z. Li, Ariella Shikanov
The genetic basis for DNA methylation variation across tissues and development
Jonathan Rosenski, Ofra Sabag, Eitan Marcus, Netanel Loyfer, Yuval Dor, Howard Cedar, Tommy Kaplan
Carmen G. Palii, Steven Tur, Sirui Yan, William J.R. Longabaugh, Romeo Solano, F. Jeffrey Dilworth, Jeffrey A. Ranish, Marjorie Brand
Judhajeet Ray, Evelyn Jagoda, Maya U. Sheth, James Galante, Dulguun Amgalan, Andreas R. Gschwind, Chad J. Munger, Jacob Huang, Glen Munson, Madeleine Murphy, Eugenio Mattei, Timothy Barry, Vasundhara Singh, Aarthee Baskaran, Helen Kang, Eugene Katsevich, Lars M. Steinmetz, Jesse M. Engreitz
Resetting of H3K4me2 during mammalian parental-to-zygote transition
Chong Wang, Yang Li, Yaqian Wang, Yong Shi, Xiangrui Meng, Wenbo Li, Jia Guo, Kaiyue Hu, Hao Chen, Jiawei Xu
Nucleosome stability safeguards cell identity, stress resilience and healthy aging
Hiroshi Tanaka, Brenna S. McCauley, Clara Guida, Xue Lei, Sha Li, Tatiana M. Moreno, K’leigh Guillotte, Zong-Ming Chua, Adrianna Abele, Aashna Lamba, Rouven Arnold, Adarsh Rajesh, Marcos G. Teneche, Laurence Haddadin, Anagha Deshpande, Aniruddha J. Deshpande, Alexandre Colas, Caroline Kumsta, Michael Petrascheck, Rolf Bodmer, Weiwei Dang, Peter D. Adams
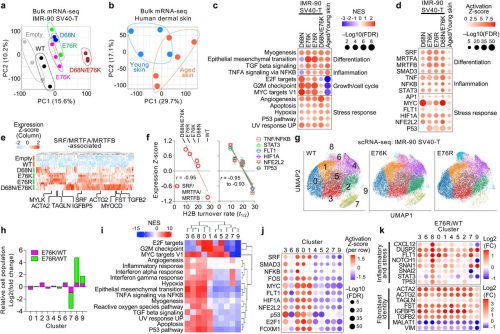
Transcriptional Readthrough at Atf4 Locus Suppresses Rps19bp1 and Impairs Heart Development
Zengming Zhang, Tongbin Wu, Zeyu Chen, Danni Chen, Zhengyu Liang, Christopher Adams, Yu Gu, Mao Ye, Fhujjen Barroga, Sylvia Evans, Xiaohai Zhou, Ju Chen
| Stem cells, regeneration & disease modelling
Erica L. Harris, Rowan D. Taylor, Katarzyna Szymanska, Ailsa M.S. Rose, Jacquelyn Bond, Colin A. Johnson, James A. Poulter
Ye Lynne Kim, Young-Woo Jo, Takwon Yoo, Kyusang Yoo, Ji-Hoon Kim, Myungsun Park, In-Wook Song, Hyun Kim, Yea-Eun Kim, Sang-Hyeon Hann, Jong-Eun Park, Daehyun Baek, Young-Yun Kong
Bernadette Banrezes, Thierry Sainte Beuve, Anne Frambourg, Alice Jouneau
Michael C. Mazzola, Ting Zhao, Anna Kiem, Trine A. Kristiansen, Karin Gustafsson, Lai Ping Wong, Emily Scott-Solomon, Marissa D. Fahlberg, Sarah Forward, Emane Rose Assita, Giulia Schiroli, Maris Handley, Youmna Kfoury, Tsuyoshi Fukushima, Samuel Keyes, Azeem Sharda, Jelena Milosevic, Hiroki Kato, Pavel Ivanov, David B. Sykes, Sheldon J. J. Kwok, Ruslan I Sadreyev, Vijay G. Sankaran, Ya-Chieh Hsu, David T. Scadden
Vcam1 in endothelial and stromal cells regulates hematopoietic stem cell contact with the niche
Octavia Santis Larrain, Alice Alhaj Kadour, Sobhika Agarwala, Wantong Li, Bradley W. Blaser, Michael R. Lasarev, Roxana Alexandridis, Anthony Veltri, Khaliun Enkhbayar, Elliott J. Hagedorn, Owen J. Tamplin
A kidney specific mouse model to study the effects of in vivo induction of Yamanaka factors
Katharina Lemberg, Gijs A.C. Franken, Korbinian M. Riedhammer, Selina Hölzel, Kirollos Yousef, Kraisoon Lomjansook, Gina Kalkar, Caroline M. Kolvenbach, Daniel Marchuk, Elena Zion, Ken Saida, Florian Buerger, Friedhelm Hildebrandt
Declan J. Gainer, Kassandra M. Coyle, Matthew T. Rätsep, Douglas Quilty, Brian Tran, Sofia Skebo, M. Martin VandenBroek, Kimberly L. Laverty, Yupu Deng, Shawyon P. Shirazi, Hugh JM Brady, Jennifer M.S. Sucre, Eric Vivier, Niraj Shrestha, Hing C. Wong, Duncan J Stewart, Nicolle J. Dominik, Mark L. Ormiston
Wentao Han, Hassan Bjeije, Hamza Celik, Michael Rettig, Nancy Issa, Andrew L. Young, Yanan Li, Infencia Xavier Raj, Christine R. Zhang, Aishwarya Krishnan, Tyler M. Parsons, Samantha C. Burkart, Jason Arand, Wei Yang, Jeffrey A. Magee, Grant A. Challen
Michelle Lohbihler, Amos A. Lim, Stéphane Massé, Maggie Kwan, Omar Mourad, Olya Mastikhina, Brandon M. Murareanu, Malak Elbatarny, Renu Sarao, Beiping Qiang, Wahiba Dhahri, Matthew L. Chang, Alice L.Y. Xu, Amine Mazine, Shahryar Khattak, Sara S. Nunes, Kumaraswamy Nanthakumar, Michael A. Laflamme, Stephanie Protze
Cory P. Johnson, Hannah M. Somers, Sophie E. Craig, Heath Fuqua, Lynne Beverly-Staggs, Kailee E. Tanaka, Sydney M. Brown, Charles H. Toulmin, Matthew D. Cox, Joel H. Graber, Melissa S. Maginnis, Hermann Haller
Giorgio Anselmi, Vincent Frontera, Christina Rode, Andrew Jarratt, Naeema T. Mehmood, Matthew Nicholls, Stella Antoniou, Emanuele Azzoni, John Stamatoyannopoulos, Ditsa Levanon, Yoram Groner, Marella F.T.R. de Bruijn
Hypoxia/HIF Signaling Negatively Regulates Bone Marrow Adiposity after Radiation Exposure
Cheyenne A. Jones, Wendi Guo, Kiana A. Gunn, Cahil Potnis, Amaya Sheffield, Colleen Wu
Exploring mechanisms of scar-free skin wound healing in adult zebrafish in comparison to mouse
İsmail Küçükaylak, Francisco Javier Martínez Morcillo, Kai Halwas, Nils Reiche, Manuel Metzger, Petra Comelli, Jürgen Brinckmann, Sabine Eming, Matthias Hammerschmidt
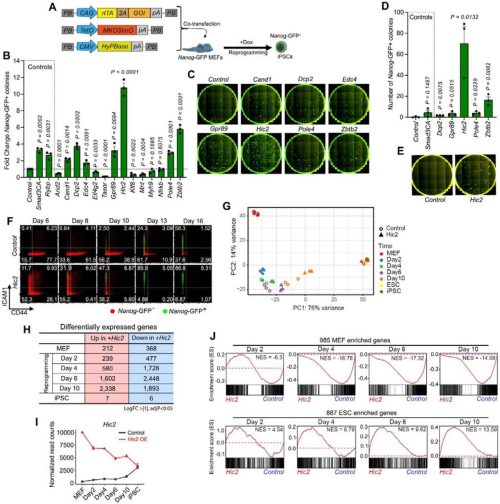
Hic2 fast-tracks iPS cell generation by suppressing KLF4-dependent epidermal detour
Meryam Beniazza, Masahito Yoshihara, Daniel F Kaemena, James Ashmore, Suling Zhao, Michael O’Dwyer, Emil Andersson, Victor Olariu, Shintaro Katayama, Abdenour Soufi, Kosuke Yusa, Keisuke Kaji
Smed-pou4-2 regulates mechanosensory neuron regeneration and function in planarians
Ryan A. McCubbin, Mohammad A. Auwal, Shengzhou Wang, Sarai Alvarez Zepeda, Roman Sasik, Robert W. Zeller, Kelly G. Ross, Ricardo M. Zayas
Lijie Gu, Kun Ho Kim, Xiyue Chen, Stephanie N Oprescu, Yufen Li, Junxiao Ren, Shihuan Kuang, Feng Yue
Elise McCollough Nanista, Landon Elizabeth Poythress, Isabell Reese Skipper, Trevor Haskins, Marieher Felix Cora, Tania Rozario
Microbes and diet reshape the intestine via distinct cellular dynamics
Alessandro Bonfini, Nicolas Buchon
TB Scheidl, JS Yoon, JL Wager, S Yonan, A Lakhdar, S Kailasam, JA Thompson
Cyrina M. Ostgaard, Arjun Rajan, Sophia R. Davidson, Cheng-Yu Lee
Tumors mimic the niche to inhibit neighboring stem cell differentiation
Yang Zhang, Yuejia Wang, Jinqiao Song, Lizhong Yan, Ziguang Wang, Dongze Song, Yudi Zhao, Shaowei Zhao
The circadian regulator PER1 inhibits osteoclastogenesis by activating inflammatory genes
Nobuko Katoku-Kikyo, Elizabeth K. Vu, Samuel Mitchell, Ismael Y. Karkache, Elizabeth W. Bradley, Nobuaki Kikyo
Felicity Ting-Yu Hsu, Rachel Smith-Bolton
Piezo1–Yap1 signalling translates tissue mechanics into regenerative cell states
Fernando Ferreira, Jaime A. Espina, Artemis G. Korovesi, Inês A. Ferreira, Quentin Tirel, Elias H. Barriga
WDR5 and Myc Cooperate to Regulate Formation of Neural Crest Stem Cells
Karlin Compton, Elizabeth Barter, Carole LaBonne
Vincent Giudice, Florence Perold, Yannicke Pijoff, Nathalie Doerflinger, Nicolas Allègre, Claire Chazaud, Irène Aksoy, Pierre Savatier, Pierre-Yves Bourillot
The livebearers platyfish and swordtails partially regenerate their hearts with persistent scarring
Vincent Hisler, Lana Rees, Simon Blanchoud, Heidi E.L. Lischer, Rémy Bruggmann, Anna Jaźwińska
Regional Signaling Controls Stem Cell-Mediated Regeneration in an Invertebrate Chordate
Tal Gordon, Tom Levy, Chester Jiamu Yu, Benyamin Rosental, Lauren Lubeck, Lucia Manni, Irving L. Weissman, Ayelet Voskoboynik
Diogo Nani, Lucas Alvizi, Heloisa Maria de Siqueira Bueno, Ellen Cristina Miranda Lacerda, Maria Rita Passos-Bueno
Jingliang Simon Zhang, Brian Guy, Clayton P. Santiago, Caterina Tiozzo, Meghana Sreenath, Ya-Wen Chen, Seth Blackshaw, Robert J. Johnston Jr.
Accelerated Tempo of Cortical Neurogenesis in Down Syndrome
Jingwen W Ding, Chang N Kim, Marilyn R Steyert, Andrew T Yuan, David Shin, Dimitar Ivanov, Tomasz J Nowakowski, Alex A Pollen
Yuancheng Ryan Lu, James C. Cameron, Yan Hu, Han Shen, Shintaro Shirahama, Alexander Tyshkovskiy, Zhaoyi Chen, Jiahe Ai, Daniel Y. Zhu, Margarete M. Karg, Lindsey A. Chew, George W. Bell, Siddhartha G. Jena, Yue He, Philip Seifert, Daisy Y. Shu, Mohamed A. EI-Brolosy, Qiannuo Lou, Bohan Zhang, Anna M. Puszynska, Xiaojie Qiu, Xiao Tian, Meredith Gregory-Ksander, Vadim N. Gladyshev, David A. Sinclair, Magali Saint-Geniez, Jason D. Buenrostro, Catherine Bowes Rickman, Bruce R. Ksander, Jonathan S. Weissman
Germline stem cell isolation, lineage tracing, and aging in a protochordate
Tom Levy, Chiara Anselmi, Katherine J. Ishizuka, Tal Gordon, Yotam Voskoboynik, Erin McGeever, Angela M. Detweiler, Liron Levin, Karla J. Palmeri, Daniel Dan Liu, Rahul Sinha, Benjamin F. Ohene-Gambill, Tal Raveh, Maurizio Morri, Virginia Vanni, Lucia Manni, Debashis Sahoo, Norma F. Neff, Benyamin Rosental, Irving L. Weissman, Ayelet Voskoboynik
Adrian On-Wah Leung, Chang Li, Xianyin Huang, Zhuming Fan, Yuning Chen, Jiaxing Liu, Hong-Man Lin, Xiaoming Cui, Fujia Zhang, Ruoxi Niu, Kam-Tong Leung, Hoi-Hung Cheung
Julia Schwarzpaul, Clara M. Droell, Afsheen Kumar, Harishny Sarma, Madeleine Gruenauer, Selen Z. Ucar, Julio C. Hechavarría, Andreas G. Chiocchetti, Denise Haslinger
Sonoepigenetic Modification Mechanoprimes Early Osteogenic Commitment in Mesenchymal Stem Cells
Lizebona A. Ambattu, Blanca del Rosal, Carmelo Ferrai, Leslie Y. Yeo
Ioanna Peraki, Ioannis K. Deligiannis, Dimitris Botskaris, Marianna Stagaki, Haroula Kontaki, Elena Deligianni, Giannis Giannoulakis, Matthieu D. Lavigne, Celia P. Martinez-Jimenez, Iannis Talianidis
| Plant development
Yuri S. van Ekelenburg, Ida V. Myking, Cathal Meehan, Morten P. Hauger, Shinichiro Komaki, Keiko Sugimoto, José Gutierrez-Marcos, Paul E. Grini
De novo meristem development in Marchantia requires light and an apical auxin minimum
Eva-Sophie Wallner, Natalie Edelbacher, Liam Dolan
Supergene control of chiral development in mirror-image flowers
Haoran Xue, Marco Saltini, Nicola Illing, Kelly Shepherd, Olivia Page-Macdonald, Oliver Marketos, Caroline Robertson, Anand Shankar, Sarah Süß, Christian Kappel, Saleh Alseekh, Eva E. Deinum, Robert A. Ingle, Michael Lenhard
Peilin Chen, Xiaocai Xu, Cezary Smaczniak, Bénédicte Desvoyes, Crisanto Gutiérrez, Robert Sablowski, Kerstin Kaufmann, Jose M Muino
Transgenerational polarity axis inheritance during Ceratopteris embryogenesis
Sjoerd Woudenberg, Andrew R.G. Plackett, Zhaodong Hao, Hidemasa Suzuki, Luis Alonso Baez, Cecilia Borassi, Thorsten Hamann, Minako Ueda, Jane A. Langdale, Joris Sprakel, Jasper van der Gucht, Dolf Weijers
COCHLEATA controls spatial regulation of cytokinin and auxin during nodule development
Karen Velandia, Muhammad Nouman Sohail, Tiana Scott, Alejandro Correa-Lozano, Alannah Mannix, Eloise Foo
Transfer RNA modifications during rhizome development in Oryza longistaminata
Wenze Li, Guangzhao Yang, Haoyue Xue, Ruichen Ma, Chaoying Zhang, Ruixuan Yao, Yajun Li, Xukai Li, Fengyi Hu, Peng Chen, Zheng Li
Frances K. Clark, Gauthier Weissbart, Xihang Wang, Kate Harline, Chun-Biu Li, Pau Formosa-Jordan, Adrienne H. K. Roeder
Filippo Binci, Giacomo Guarneri, Sofía Cristina Somoza, Filippo Vascon, Arianna Capparotto, Edoardo Di Nuzzo, Giacomo Rago, Laura Cendron, Lorella Navazio, Marco Giovannetti
Progressive oxygenation of developing leaves directs morphogenesis
Gabriele Panicucci, Vinay Shukla, Viktoriia Voloboeva, Leonardo Jo, Kees van Kollenburg, Sara Buti, Laura Dalle Carbonare, Francesco Licausi, Daan A. Weits
Dongeun Go, Bailan Lu, Liang Song
Nadra Tabassum, Justin Goodrich, Gary J. Loake
| Environment, evolution and development
Expanding the fly eye gene regulatory network: from Drosophila to the hoverfly Episyrphus balteatus
T. Navarro, J. Torres, R. Sáez-Moreno, G. Guerrero, I. Almudí, A. Iannini, J. Figueras, K Wotton, S. Aerts, F. Casares
Oogenesis and germinal bed morphology of the brown anole (A. sagrei)
Bonnie K. Kircher, Antonia Weberling, Erin J. Vance, Natalia A. Shylo, Katherine Starr, Zoe B. Griffin, Hannah Wilson, Melainia McClain, Florian Hollfelder, Suzannah A. Williams, Thomas J. Sanger, Richard R. Behringer, Paul A. Trainor
doi:
Axel H Newton, Ella R Farley, Andrew T Major, Jennifer C Hutchison, Ben M Lawrence, Karen E Sears, Marilyn B Renfree, Aiden M C Couzens, Geoff Shaw, Sara Ord, Richard A Schneider, Andrew J Pask
The heterochronic activation of TGF-β signaling drives the diversity of the avian sterna
Seung June Kwon, Zhaonan Zou, Mizuki Honda, Shiro Egawa, Shinya Oki, Yuji Atsuta
Jasmine D. Alqassar, Teomie S. Rivera-Miranda, Joseph J. Hanly, Christopher R. Day, Silvia M. Planas Soto-Navarro, Paul B. Frandsen, Riccardo Papa, Arnaud Martin
Laurel S. Hiebert, Ava Soesbe, Tzung-Fu Hsieh, Qirui Cui, Soojin V. Yi
The evolution of investment in innate-like and diversified T cell receptors across development
Reese A. Martin, Anna E. Savage, Ann T. Tate
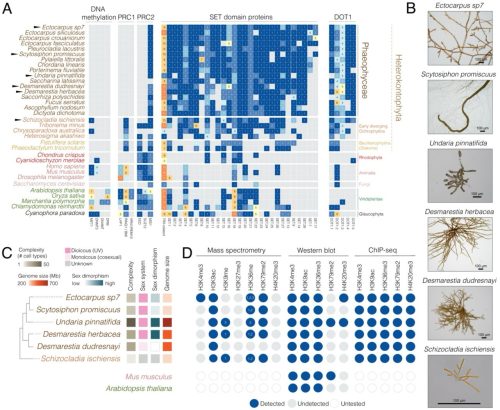
Rewiring of chromatin regulation underlies the evolution of brown algal multicellularity
Jeromine Vigneau, Jaruwatana Sodai Lotharukpong, Pengfei Liu, Remy Luthringer, Bérangère Lombard, Damarys Loew, Fabian B. Haas, Michael Borg, Susana M Coelho
Alison Hall, Manali Rege-Colt, Melissa Pespeni
PIF4-mediated regulation of H2O2 homeostasis controls Arabidopsis seedling thermomorphogenesis
Ritesh Kumar Raipuria, Kumud Saini, Mahesh Kumar Panda, Aditi Dwivedi, Aashish Ranjan
Cell Biology
Separately prestored proteasome components to prevent polyspermy
Liying Wang, Chao Liu, Xing Wang, Yueshuai Guo, Qiuxing Zhou, Yinghong Chen, Huafang Wei, Xiaoming Huang, Shuai Liu, Wei Li, Chun So, Ling Sun, Renjie Jiao, Xuejiang Guo, Qiang Guo, Wei Li
Mitochondrial organization in the developing proximal tubule is controlled by LRRK2
Mohsina Anjum Khan, Kyle Bond, Elyse Grilli, Daniel Cameron, Liyang Zhao, Sunder Sims-Lucas, Andrew P. McMahon, Thomas J. Carroll, Leif Oxburgh
Mitochondrial fission regulates ROS for ventral furrow formation in Drosophila gastrulation
Somya Madan, Sayali Chowdhary, Sakshi Phalke, Harsh Mittal, Manos Mavrakis, Richa Rikhy
Atsushi Saito, Stephanie Tankou, Kazuhiro Ishii, Makiko Sakao-Suzuki, Edwin C. Oh, Hannah Murdoch, Ho Namkung, Sunday Adelakun, Keiko Furukori, Masahiro Fujimuro, Paolo Salomoni, Gerd G. Maul, Gary S. Hayward, Qiyi Tang, Robert H. Yolken, Miles D. Houslay, Nicholas Katsanis, Isao Kosugi, Kun Yang, Atsushi Kamiya, Koko Ishizuka, Akira Sawa
Ji Kent Kwah, Shannon Pfeiffer, Mst Gitika Khanom, Aimee Jaramillo-Lambert
Mitochondria-Lysosome Crosstalk Shapes Metabolic Transition in Neonatal Enterocytes
Gonzalo Herranz, Diego Alonso-Larre, Tamara González, Laura Akintche, Alejandra Ramos-Manzano, Marta Iborra-Pernichi, María Velasco de la Esperanza, Covadonga Díaz-Díaz, Ian G Ganley, Patricia Boya, Sara Cogliati, Nuria Martínez-Martín, Fernando Martín-Belmonte
ANKRD5: a key component of the axoneme required for sperm motility and male fertility
Shuntai Yu, Guoliang Yin, Peng Jin, Weilin Zhang, Yingchao Tian, Xiaotong Xu, Tianyu Shao, Yushan Li, Fei Sun, Yun Zhu, Fengchao Wang
Mouse germline cysts contain a fusome that mediates oocyte development
Madhulika Pathak, Allan C. Spradling
Organ-specific rewiring of mitochondrial integrity through COX7A dictates cellular ploidy control
Archan Chakraborty, Sophia DeLuca, Meera Gangasani, Stephen Rogers, Nenad Bursac, Donald T. Fox
Jyotsna Kawadkar, Ashley Suraj Hermon, Rohit Kumar, Ram Kumar Mishra
Daniel A. Reed, Anna E. Howell, Nadia Kuepper, Alice M. H. Tran, Astrid Magenau, Deborah S. Barkauskas, Max Nobis, Cecilia R. Chambers, Victoria Lee, Lily M. Channon, Jessie Zhu, Shona Ritchie, Janett Stoehr, Kaitlin Wylie, Julia Chen, Denise Attwater, Kate Harvey, Sunny Z. Wu, Kate Saw, Ruth J. Lyons, Anaiis Zaratzian, Michael Tayao, Andrew Da Silva, David Gallego-Ortega, Anthony J. Gill, Thomas R. Cox, Brooke A. Pereira, Kendelle J. Murphy, Jennifer P. Morton, Elgene Lim, Alexander Swarbrick, Sandra O’Toole, Michael S. Samuel, C. Elizabeth Caldon, Alexandra Zanin-Zhorov, Paul Timpson, David Herrmann
Histone methyltransferase DOT1L differentially affects the development of dendritic cell subsets
Rianne G. Bouma, Willem-Jan de Leeuw, Aru Z. Wang, Muddassir Malik, Joeke G.C. Stolwijk, Veronique A.L. Konijn, Anne Mensink, Natalie Proost, Maarten K. Nijen Twilhaar, Tibor van Welsem, Negisa Seyed Toutounchi, Alsya J. Affandi, Jip T. van Dinter, Fred van Leeuwen, Joke M.M. den Haan
Mechanical context defines integrin requirement for maintaining epithelia architecture
Lourdes Rincón-Ortega, Cecilia H. Fernández-Espartero, Isabel M. Palacios, Acaimo González-Reyes, María D. Martín-Bermudo
Yasuko Inaba, Kimika Iwasaki, Aoi Nakamura, Shiro Suetsugu, Yasumasa Bessho
Lamiya Dohadwala, Anupriya Garg, Maithreyi Narasimha
Role of Klhl14 in senescence and epithelial-to-mesenchymal transition via TGF-β modulation
Rufina Maturi, Abel Soto-Gamez, Anne L Jellema-de Bruin, Matteo Esposito, Schelto Kruijff, Gabriella De Vita, Rob P. Coppes
Tooba Khan
Alec McDermott, Melany Juarez, David Tovar-Parra, Capucine Duval, Shunmoogum (Kessen) Patten, Isabelle Plante
Spatio-temporal control of nuclear mechanotransduction during Epithelial-to-Mesenchymal Transition
Ronan Bouzignac, Amandine Palandri, Amal Zine El Aabidine, Thomas Mangeat, Tatiana Merle, Martine Cazales, Antonio Trullo, Christian Rouviere, Virginia Pimmett, Mounia Lagha, Magali Suzanne
Rotational migration in human pancreatic ductal organoids depends on actin and myosin activity
Gengqiang Xie, Chaity Modak, Olalekan H Usman, Raphael WF Tan, Nicole Coca, Gabriela De Jesus, Yue Julia Wang, D. Thirumalai, Xin Li, Jerome Irianto
A dynamic repertoire of wound closure strategies precedes whole-body regeneration
Allison P. Kann, Mansi Srivastava
Akihiro Aikawa, Junya Ito, Mika Toya, Masamitsu Sato
Modelling
Bayesian phylodynamics for developmental biology: incorporating age-dependence
Nicola Mulberry, Julia Pilarski, Jana Dinger, Tanja Stadler
Epigenetic–metabolic axis in the temporal scaling of mammalian cortical neurogenesis across species
Quan Wu, Charlotte Manser, Taeko Suetsugu, Ryo Yoshida, Hideya Sakaguchi, Yoichi Nabeshima, Hiroshi Kiyonari, Ruben Perez-Carrasco, Fumio Matsuzaki
Simulation-Based Inference of Cell Migration Dynamics in Complex Spatial Environments
Jonas Arruda, Emad Alamoudi, Robert Mueller, Marc Vaisband, Ronja Molkenbur, Jack Merrin, Eva Kiermaier, Jan Hasenauer
Tools & Resources
Tongtong Xu, Fujun Cao, Ruihan Zhou, Qin Chen, Jian Zhong, Yulin Wang, Chaoxin Xiao, Banglei Yin, Chong Chen, Chengjian Zhao
FishFeats: streamlined quantification of multimodal labeling at the single-cell level in 3D tissues
Gaëlle Letort, Tanya Foley, Ilona Mignerey, Laure Bally-Cuif, Nicolas Dray
CUT&TIME captures the history of open chromatin in developing neurons
Kiara C. Eldred, Matthew Wooten, Derek H. Janssens, Joshua Hahn, Shane J Neph, Sierra J. Edgerton, Gracious Wyatt-Draher, Stephanie M. Sherman, Jane E. Ranchalis, Andrew B. Stergachis, Thomas A. Reh, Steven Henikoff
SpinePy enables automated 3D spatiotemporal quantification of multicellular in vitro systems
Ryan G. Savill, Alba Villaronga-Luque, Marc Trani Bustos, Yonit Maroudas-Sacks, Julia Batki, Alexander Meissner, Allyson Q. Ryan, Carl D. Modes, Otger Campàs, Jesse V Veenvliet
Poly(A) probe HCR RNA-FISH specifically marks pyriform nurse cells in the brown anole lizard ovary
Zoe B. Griffin, Bonnie K. Kircher, Richard R. Behringer
Prediction of zoonotic virus-host transmissibility using comparative airway organoids
Namwook Kim, Christine S. Kim, Junhee Park, Hyeon Jun Yoon, Ju Ang Kim, Yong Gun Kim, Joonhyung Kim, Injun Song, Donghyuck Ahn, Jihyeon Myeong, Byungmoo Oh, Jaeyoon You, Eunju Hong, Sukin Jeong, Kyungmoo Yea, Sung Won Kim, Ok Sarah Shin, Seung Joon Kim, Minho Lee, Myungin Baek, Youngtae Jeong
Automated Image-Based Profiling of Pluripotent Stem Cell Colonies
Rui Geng, Benjamin L. Kidder
Jose Maria Aguilar-Camacho, Christina Zakas
Rui Geng, Benjamin L. Kidder
Human DCM-time machine unravels cell state changes during primitive gut tube differentiation
Marieke E. van Leeuwen, Beatrice F. Tan, Evelyne Wassenaar, Esther Sleddens-Linkels, Ruben Boers, Cristina Gontan, Joost Gribnau
Identification of optimal fluorophores for use in the Drosophila embryo
Bernardo Chapa-y-Lazo, Thamarailingam Athilingam, Prabhat Tiwari, Prachi Pathak, Shaobo Zhang, Sophie Theis, Timothy E Saunders
Direct, high-throughput linking of single cell imaging and gene expression
Catherine K Xu, Georg Meisl, Nikita Moshkov, Karolis Goda, Alexey Shkarin, Maximilian F Schlögel, Tuomas PJ Knowles, Linas Mazutis, Jochen Guck
Streamlined Montage Cryo-Electron Tomography for Exploring the Ultrastructure of Cells and Tissues
Ryan Hylton, Micaela Boiero Sanders, Adriana Prajica, Gavin Rice, Stefan Raunser
Recapitulating apicobasal tissue polarity in extracellular matrix incorporated airway organoids
Zhuowei Gong, Dhruv Bhattaram, Laura Porritt, Kian Golestan, Amir Barati Farimani, Amy L. Ryan, Daniel J. Weiss, Xi Ren
Bridging the resolution gap in cryo-CLEM by introducing cryo-SXT: cryo-CLXEM
Johannes Groen, Anastasia Gazi, Sergey Kapishnikov, Anne Brelot, Matthijn Vos, Jost Enninga, Eva Pereiro, Anna Sartori-Rupp
Adrien Rihoux, Alexie Gagné, Jean Mezreani, Clémence Gonthier-Cummings, Laura K. Hamilton, Eric Samarut, Martine Tétreault
Jingyu Wang, Danail Stoychev, Mick Phillips, David Miguel Susano Pinto, Richard M. Parton, Nick Hall, Josh Titlow, Ana Rita Faria, Matthew Wincott, Dalia Gala, Andreas Gerondopoulos, Niloufer Irani, Ian Dobbie, Lothar Schermelleh, Martin Booth, Ilan Davis
SPACE: spatially resolved multiomic analysis for high-throughput CRISPR screening in 3D models
Mengwei Hu, Yi Cui, Qianhui Huang, Khoi Chu, Sierra McKinzie, Michael Patrick, Sharanya Iyengar, Maerjianghan Abuduli, Marianne Spatz, Nandita Joshi, Brendan Miller, Shams Vellarikkal, Timothy Riordan, Danny Bitton, Jan Lubojacky, Iya Khalil, Federica Piccioni, Michael Rhodes, Alex Tamburino, Shanshan He, Joseph Beechem, Vanessa Peterson
ArchVelo: Archetypal Velocity Modeling for Single-cell Multi-omic Trajectories
Maria Avdeeva, Sarah Walker, Joris van der Veeken, Alexander Rudensky, Yuri Pritykin
Generation of Valvular Interstitial Cells from Human Pluripotent Stem Cells
Amine Mazine, Alexander A. Mikryukov, Ian Fernandes, Clifford Z. Liu, Soheil Jahangiri, Marcy Martin, Eric K. N. Gähwiler, Melanie Generali, Juliana Gomez, Neda Latifi, Yifei Miao, Yu Liu, Michael A Laflamme, Craig A Simmons, Simon P. Hoerstrup, Maximilian Y. Emmert, Bruce D. Gelb, Mingxia Gu, Gordon M. Keller
Automatic Body Morphometric Analysis of Adult Zebrafish Using MicroCT
Mohammed Kanani, Ronald Young Kwon
Alternating Angle Milling Suppresses Streaking Artifacts in FIB-SEM Imaging
Gleb Shtengel, Wei Qiu, Jesse Aaron, Arthur S. Crowe, Alexey A. Polilov, Katerina Karkali, Christopher K.E. Bleck, Harald H. Hess
Smart Microscopy: Current Implementations and a Roadmap for Interoperability
Lucien Hinderling, Hannah S. Heil, Alfredo Rates, Philipp Seidel, Manuel Gunkel, Benedict Diederich, Thomas Guilbert, Rémy Torro, Otmane Bouchareb, Claire Demeautis, Célia Martin, Scott Brooks, Evangelos Sisamakis, Erwan Grandgirard, Jerome Mutterer, Harrison Oatman, Jared Toettcher, Andrii Rogov, Nelda Antonovaite, Karl Johansson, Johannes K. Ahnlinde, Oscar André, Philip Nordenfelt, Pontus Nordenfelt, Claudia Pfander, Jürgen Reymann, Talley Lambert, Marco R. Cosenza, Jan O. Korbel, Rainer Pepperkok, Lukas C. Kapitein, Olivier Pertz, Nils Norlin, Aliaksandr Halavatyi, Rafael Camacho
Jason John Walsh, Levent Görgü, Emilie Jacob, Victoria Poulain, Laurent Gutierrez, Eleni Mangina, Sónia Negrão
Research practice & education
Mirror-image inversion in commonly used compound microscopes
François Lapraz, Céline Boutres, Baptiste Monterroso, Stéphane Noselli
Angelina Miller, Katja L. Schröder, Karsten E. Braun, Caitlin Steindorf, Richard Ottermanns, Martina Roß-Nickoll, Henner Hollert, Thomas Backhaus
Sue Lim, Ralf Schmälzle, Gary Bente
Raquel Marco-Ferreres, Silvia Ayora, Carmela Cela, Álvaro Sahún-Español, María López-Sanz, Pablo Izquierdo
Natural history museums are missing an opportunity to present research and collections on YouTube
Selina A. Ruzi, Adrian A. Smith, Nicole M. Lee, Nicolás J. Galvez, Timothy A. Dinh
Evaluation of the Utility of a Research Ethics Training Course to Graduate Students
Michael D. Schaller, Peter H. Mathers
Jack Nunn, Håkon da Silva Hyldmo, Lauren McKnight, Heather McCulloch, Jennifer Lavers, Julie Old, Laura Smith, Nicola Grobler, Cheryl Tan Kay Yin, Wing Yan Chan, Candice Raeburn, Nittya S. M. Simard, Adam Kingsley Smith, Sam Van Holsbeeck, Eleanor Drinkwater, Kit Prendergast, Emma Burrows, Christopher L. Lawson


 (No Ratings Yet)
(No Ratings Yet)
 (3 votes)
(3 votes)