What does a Reviews Editor do?
Posted by Saanjbati Adhikari, on 14 May 2026
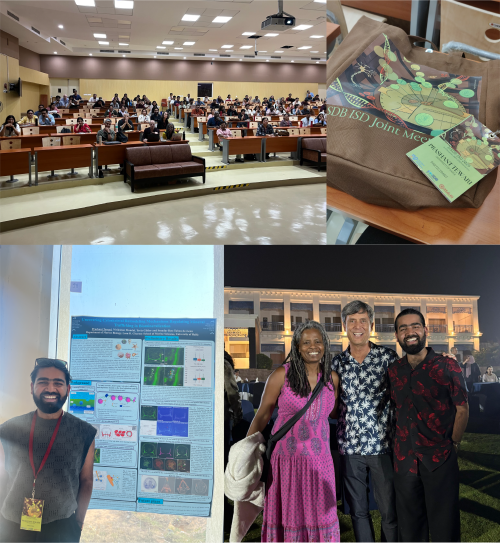
So, recently I attended a developmental biology conference – my first one of 2026, with six more to go! While socialising and networking with a group of truly amazing stem cell researchers, many of them asked, after I introduced myself as a Reviews Editor at Development, “What exactly does a Reviews Editor do?” After answering this question at least five times – across scientists at different career and life stages – I realised it might be time to share with you all what we ‘cool kids’ actually do.
Normally, at The Company of Biologists (Development’s publisher), we follow a hybrid working model, which means that we work from home for half of the week (which is usually 2-3 days a week for me) and the rest in the office. If you haven’t seen a photo of our office building yet, it is a rather beautiful building, combining the charm of a cottage-style exterior (complete with hipped roofs and classic sash windows) with a bright, modern and open-style office inside.
At Development, Ingrid Tsang and I are the Reviews Editors and we mainly handle the journal’s front-section content (so, that includes Reviews, Spotlights, Perspectives, Hypotheses, Primers, interviews – yeah, we have an extensive list!) and we work closely with Alex Eve, Executive Editor of Development.

I normally start my workday between 9:30-10 am, and my first task is always to reply to emails while sipping my morning coffee (a strong flat white if I am home and a long cappuccino when in the office).
After the first half an hour to attending to emails regarding submissions, chasing authors for their submissions, or finishing off a pending task from the previous day (which often involves taking a final read through a decision letter), I move on to the main tasks of the day.
If I am working on a chunky edit – meaning a developmental edit of a Review article – I would usually block off an entire day for it (at least 7–8 hours). This typically happens once a Review-type article (which we commission in-house and invite authors to submit) has gone through peer review. At that stage, we, the in-house Reviews Editors, read through the full manuscript in detail, commenting on scientific accuracy, language and structure, conciseness and accessibility, journal style, article length and references – all while helping authors address the Reviewers’ comments more effectively. I also go through the figures and legends (we take our display items very seriously, as a single figure often speaks a thousand words), commenting on visual appeal, labelling and other finer details. Developmentally editing an article is usually the most rigorous part of the job, at least in my opinion, as it ensures that the final piece is not only of high quality but also forward-looking and engaging for our wide readership. At Development, we pride ourselves on being extremely hands-on when guiding authors and helping them address both our feedback and the Reviewers’ comments, in order to publish the best possible version of their review.
Another crucial part of our job is commissioning. We have our in-house commissioning meetings every two weeks. So, if you catch me the afternoon before, I am usually frantically reading articles on a certain topic of interest, trying to prepare somewhat cohesive pitches to discuss with the rest of the team. We mainly invite authors to write peer-reviewed, review-type content for us. To identify emerging topics in the field, we attend important conferences, chat with researchers across a wide range of developmental biology disciplines, keep an eye on their websites, analyse research trends across primary research articles and participate in extensive 1-1.5-hour long commissioning meetings.
Once we have agreed on a topic and a suitable author, we invite them to write for us. If they accept our invitation, the author will then often involve their students and collaborators as co-authors, and at that point we discuss the potential scope and type of the article. Of course, we also have a thorough in-house pipeline that monitors the status of all articles from invitation through to acceptance.
When we’re in the office, Alex, Ingrid, Andrea (Community Manager of the Node) and I often chat about various aspects of the job throughout the day – both formally and informally – because our work requires teamwork and collaboration. So, Ingrid and I will often discuss scheduling to make sure our publication pipeline stays tight (i.e. that every Issue publishes a few front-section content). If we have just returned from a conference, we chat with Alex and Andrea about emerging research trends and potential blogs/ posts for the community site. We also bounce around ideas for commissioning topics and share feedback on each other’s pitches, amidst a healthy dose of random life chats.
For me, the day usually ends with a quick catch-up on plans for the next day. This is also when I respond to any remaining email replies from the morning. I usually like to do a final run-through of my to-do list, ticking things off and marking any pending tasks (if there are any). And with that, I sign off for the day!
PS: Ingrid has also written a piece on a day in the life of a Reviews Editor – so do give that a read as well!


 (4 votes)
(4 votes)
 (No Ratings Yet)
(No Ratings Yet)